Scientific Foundations
Concurrent constraint programming
The class of Concurrent Constraint programming languages (CC) was introduced a decade ago by Vijay Saraswat as a unifying framework for constraint logic programming and concurrent logic
programming. The CC paradigm constitutes a representative abstraction of constraint programming languages, and thus allows a fine grained study of their fundamental properties.
CC generalizes the Constraint Logic Programming framework (CLP) by introducing a synchronization primitive, based on constraint entailment. It is a model of concurrent computation, where
agents communicate through a shared store, represented by a constraint, which expresses some
partial informationon the values of the variables involved in the computation. The variables play the role of transmissible dynamically created communication channels.
One of the big successes of CC has been the simple and elegant reconstruction of finite domain constraint solvers, and the cooperation of several models to solve a single combinatorial
problem. On the other hand, to use CC for programming reactive applications forces one to abandon the hypothesis of monotonic evolution of the constraint store; this is a strong motivation for
new extensions of CC languages.
There are strong completeness theorems relating the execution of a CLP program and its translation in classical logic, which provide smooth reasoning techniques for such programs. However
these theorems are broken by the synchronization operation of CC. Looking for a logical semantics of CC programs in the general paradigm of logic programming,
program = logical formula,
execution = proof search,
leads to a translation in Jean-Yves Girard's linear logic. This allows the recovery of some completeness results about successes and stores; even suspensions may be
characterized with the non-commutative logic of Ruet and Abrusci.
It is thus possible to address important issues for Constraint Programming:
verifying CC programs;
combining CLP and state-based programming;
dealing with local search inside a global constraint solving procedure.
The last two cases rely on a natural extension of CC languages, called Linear Concurrent Constraint languages (LCC), which simply replaces constraint systems built onto classical logic by
constraint systems built onto linear logic. This allows us to represent state changes thanks to the consumption of resources during the synchronization action, modeled by the linear
implication.
Constraint solvers
Our domains of application use quite different constraint systems:
finite domains (bounded natural numbers): primitive constraint of some finite domain membership, numerical, symbolic, higher order and global constraints;
reals: Simplex algorithm for linear constraints and interval methods otherwise;
terms: subtyping constraints and ontologies;
temporal constraints: CTL and LTL formulae, either propositional or with numerical constraints.
The project works on constraint resolution methods and their cooperation. The main focus is the declarativeness of the constraint solver (e.g. implemented by CHR rules), the efficiency of
constraint propagation methods, the design of global constraints and the combination of constraint propagation with local search methods.
Computational systems biology
Systems biology is a cross-disciplinary domain involving biology, computer science, logics, mathematics, and physics to elucidate the high-level functions of the cell from their biochemical
bases at the molecular level.
At the end of the Nineties, research in Bioinformatics evolved, passing from the analysis of the genomic sequence to the analysis of post-genomic interaction networks (expression of RNA and
proteins, protein-protein interactions, etc). The complexity of these networks requires a large research effort to develop symbolic notation and analysis tools for biological processes and
data. In order to scale-up, and get over the complexity walls to reason about biological systems, there is a general feeling that beyond providing tools to biologists, computer science has much
to offer in terms of concepts and methods.
We are interested in the modeling and analysis of complex molecular processes in the cell, at different levels of abstraction, qualitative and quantitative. The most original aspect of our
research can be summarized by the following identifications:
biological model = transition system,
biological property = temporal logic formula,
biological validation = model-checking.
Our main research axis is thus the application of logic programming concepts and circuit or program verification techniques to the analysis of complex biochemical processes in the cell.
Application Domains
Combinatorial optimization problems
The number and economic impact of combinatorial optimization problems found in the industrial world are constantly increasing. They cover:
resource allocation;
placement;
scheduling;
planing;
transport;
etc.
The last forty years have brought many improvements in Operations Research resolution techniques. In this context, Constraint Programming can be seen as providing, on the one hand, local
consistency techniques that can be applied to various numerical or symbolic constraints, and on the other hand, declarative languages. This last point is crucial for quickly developing complex
combinations of algorithms, which is not possible without a language with a high level of abstraction. It allowed for better results, for instance in scheduling problems, than traditional
methods, and is promised to an even better future when thinking about the cooperation of global resolution, local consistency techniques and search methods.
The project builds upon its knowledge of CC languages, constraint solvers and their implementation to work in these directions. The LCC paradigm offers at the same time a theoretical
framework for analysis, and a valuable guide for practical language design and implementation. The work on programming environments helps to integrate the Constraint Programming tools into this
application domain.
In-silico cell
In 2002 , we started a Collaborative Research Initiative ARC CPBIO on ``Process Calculi and Biology of Molecular Networks''. By working on well understood biological models, we sought:
to identify in the family of competitive models coming from the Theory of Concurrency
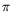 -calculus, Join-calculus and their derivativesand from Logic Programming
Constraint Logic Programming, Concurrent Constraint languages and their extensions to discrete and continuous time, TCC, HCC, the
ingredients of a language for the modular and multi-scale representation of biological processes;
-calculus, Join-calculus and their derivativesand from Logic Programming
Constraint Logic Programming, Concurrent Constraint languages and their extensions to discrete and continuous time, TCC, HCC, the
ingredients of a language for the modular and multi-scale representation of biological processes;
to provide a series of examples of biomolecular processes transcribed in formal languages, and a set of biological questions of interest about these models;
to design and apply to these examples formal computational reasoning tools for the simulation, the analysis and the querying of the models.
This work lead us to the design and implementation of the Biochemical Abstract Machine BIOCHAM that has the unique feature of providing formal languages corresponding to different
qualitative and quantitative levels of abstraction, for, on the one hand, modeling biomolecular interaction diagrams with reaction rules, and on the other hand, modeling the biological
properties of the system in temporal logic. This double formalization of both the model and the biological properties of the system at hand opens several new research avenues on the design and
systematic validation of biological models.
We participate in two European Projects of the 6th PCRD. First the STREP
[APrIL II]where the main objective is to
apply probabilistic inductive logic programming techniques to bioinformatics applications. Our main focus is on the regulatory and metabolic networks in the cell, and the semi-automatic
completion or revision of models from observed temporal properties of the system
[REWERSE], in which we focus, among other
themes, on bioinformatics as a field of application of the new Semantic Web technologies based on rules and constraints. In this context we intend to use the biological knowledge stored in
online ontologies like
[GO]in order to improve the re-use
and composition of models, as well as the semi-automatic correction/completion of models. In this respect, we developped type inference and abstract interpretation techniques to extract
information from reaction models
We coordinate the ARC
[MOCA](2006-2008) on
``MOdularity, Compositionality and Abstraction in gene and protein networks'', and collaborate in this framework with Denis Thieffry, Claudine Chaouyia, Univ. Marseille, Anne Siegel and Ovidiu
Radulescu, IRISA SYMBIOSE and IHES, Ralf Blossey IRI and Vincent Danos CNRS Paris, on the exploration of different techniques and methods allowing semi-automatic composition, decomposition,
simplification and refinement of biological models. More precisely, we study the formal links between logical and numerical models of some parts of the cell cycle and the decomposition in
modules based on control theory intuitions.
This technology developed in the last years is now applied to new biological questions that we investigate in partnerships with biologists in two new projects. First, the EU STREP project
TEMPO on ``temporal genomics for patient tailored chornotherapeutics'', coordinated by Francis Levi INSERM Villejuif, where, in partnership with Jean Clairambault of the BANG project-team, we
develop in BIOCHAM coupled models of the cell cycle, the circadian cycle and the effect of cytotoxic drugs in cancer therapies. Second, the INRA AgroBi project, coordinated by Eric Reiter INRA
Tours, where, in partnership with Frédérique Clément of the SOSSO2 project-team, we develop in BIOCHAM models of FSH signaling networks in mammalian cells.
Software
BIOCHAM
Nathalie
Chabrier-Rivier
François
Fages
Sylvain
Soliman
The Biochemical Abstract Machine
[BIOCHAM]is a modeling and
validation environment for molecular systems biology. BIOCHAM provides precise semantics to biomolecular interaction maps at three abstraction levels:
the boolean semantics of molecules presence and absence,
the continuous semantics of molecular concentrations,
the stochastic semantics of molecule populations.
Based on this formal framework, BIOCHAM offers:
a compositional rule-based language for modeling biochemical systems, allowing patterns and kinetic expressions when numerical data are available;
numerical and boolean simulators;
a temporal logic language (CTL for qualitative models and LTL with numerical constraints for quantitative models) for formalizing biological properties such as reachability, checkpoints,
oscillations or stability, and checking them automatically with model-checking techniques;
automatic search procedures to infer interaction rules and parameter values from known temporal properties of the system.
BIOCHAM is fully implemented in GNU-Prolog and interfaced to the state-of-the-art symbolic model checker
[NuSMV]for evaluating boolean CTL
queries in large models over several hundreds of variables. BIOCHAM models can be imported from, and exported to, the standard Systems Biology Markup Language
[SBML].
TCLP
Emmanuel
Coquery
[TCLP]is a
prescriptive type system for Constraint Logic Programming, currently: ISO-Prolog, GNU-Prolog, SICStus Prolog and the constraint programming libraries of SICStus Prolog. TCLP can also type check
constraint solvers written in the
Constraint Handling Rules(CHR) language.
The flexibility of the TCLP type system is obtained by the combination of three kinds of polymorphism: parametric polymorphism (e.g.
list(A)), subtyping (e.g.
list(A)
<term) and overloading (e.g.
-:num
*num
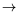 numand
-:A
*B
numand
-:A
*B
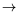 pair(A,B)). No type declaration are required, thanks to a type inference algorithm for predicates and to a default
termtype for function symbols.
pair(A,B)). No type declaration are required, thanks to a type inference algorithm for predicates and to a default
termtype for function symbols.
GNU-Prolog
Daniel
Diaz
Rémy
Haemmerlé
[GNU Prolog]is a free Prolog compiler
with constraint solving over finite domains developed by Daniel Diaz. GNU Prolog accepts Prolog extended with primitives for constraint programming and produces native binaries (like gcc does
from a C source). The Prolog part conforms to the ISO standard for Prolog with many practical extensions (global variables, OS interface, sockets,...). GNU Prolog also includes an efficient
constraint solver over Finite Domains (FD), giving the user the combined power of constraint programming and the declarativity of logic programming.
An experimental version, called GNU-Prolog-RH, is developed as an extension introducing concurrency, attributed variables, and constraint solving over the reals. This version greatly extends
the expressive and modeling power of GNU-Prolog. It is also used to prototype the design and implementation of our new SiLCC language.
CLPGUI
François
Fages
[CLPGUI]is a generic
graphical user interface written in Java for constraint logic programming. It is available for GNU-Prolog and SICStus Prolog. CLPGUI has been developed both for teaching purposes and for
debugging complex programs. The graphical user interface is composed of several windows: one main console and several dynamic 2D and 3D viewers of the search tree and of finite domain
variables. With CLPGUI it is possible to execute incrementally any goal, backtrack or recompute any state represented as a node in the search tree. The level of granularity of the search tree
is defined by annotations in the CLP program.
Tra4CP
Pierre
Deransart
Following the former OADymPPaC project, we have set up with J.-D Fekete a new open source project called
[Tra4CP](Traces for Constraint
Programming). The main objective of this open project is to build a repository of constraint resolution traces and analysis tools.
The purpose is to pursue dissemination the results of the former
[OADymPPaC]project, to
facilitate contacts and working exchanges between researchers in the field of complex problem solving using constraint programming, and mainly to promote new analysis methods. Following this
idea, the project allowed to integrate recent developments related to the use of the generic trace format Gentra4CP for constraint programming to produce trace analysis based on music (work of
Jeremie Vautard
[Tra4CP]). The creation of a working group on constraint problem XML repsesentation and its integration in the Gentra4CP is also considered.
New Results
Implementation of SiLCC
François
Fages
Rémy
Haemmerlé
Adiga
Satish
Sylvain
Soliman
We are developing SiLCC an imperative and concurrent constraint programming language based on a single paradigm: the one of Vijay Saraswat's concurrent constraint programming extended with
constraint systems based on Jean-Yves Girard's Linear Logic. In the late 90's we developed the theory of this extension and we have now begun its implementation.
From a constraint programming point of view, the unique combination of constraint programming with imperative features opens many new possibilities, among which:
the capability of programming constraint solvers in the language, making them extensible by the user,
making a fully bootstrapped implementation of a constraint programming language (for the first time since Prolog)
combining constraint reasoning with state change;
proving program correctness using Linear Logic.
Our current implementation of SiLCC uses
[
GNU-Prolog-RH]as temporary kernel language, on top of which a module system and the first bootstrapping libraries have been developed. The objective is to define a small kernel language as
an instance of LCC over a simple constraint domain of labelled graphs, on top of which the complete SiLCC language will be built by bootstrapping. Bootstrapping is a fundamental step for
getting over the current limits of today constraint programming tools, concerning their extensibility, robustness, and teaching. The main step realized this year has been the definition of the
module system.
Module System for SiLCC
François
Fages
Rémy
Haemmerlé
Sylvain
Soliman
Module systems are an essential feature of programming languages as they facilitate the re-use of existing code and the development of general purpose libraries. However, there has been no
consensual module system for Prolog, hence no strong development of libraries, in sharp contrast to what exists in Java for instance. One difficulty comes from the
callpredicate which interferes with the protection of the code, an essential task of a module system. In
The module system of SiLCC goes however beyond the one we propose for Prolog-like languages. There are indeed two somewhat contradictory ways of looking at modules in a given programming
language. On the one hand, module systems are largely independent of the particulars of programming languages, and several examples of module systems have indeed been adapted to different
programming languages. On the other hand, the module constructs often interfere with the programming constructs, and may be redundant with other scope mechanisms of programming languages, such
as closures for instance. There is therefore a need to unify the programming concepts and constructs that are similar, and retain a minimum number of essential constructs to avoid arbitrary
programming choices. In
Type systems
Emmanuel
Coquery
Pierre
Deransart
François
Fages
Sylvain
Soliman
The
Constraint Handling Rules(CHR) are an untyped rewrite rule language, initially designed for implementing constraint solvers. In order to benefit from type programming for the development
of larger programs, we have designed a generic type system for CHR
In
In the framework of computational systems biology, and in particular for the BIOCHAM modeling environment, we developed type inference and abstract interpretation techniques to either type
check reaction models or infer types from reaction models
Constraint programming environments
Pierre
Deransart
Semantics based debugging, static and dynamic typing, and more generally validation of CC programs are also studied in the project.
We started to study theoretical questions issued from the former OADymPPaC project (RNTL ended in 2004) by trying to clarify the notions of trace exchanged between observed and observing
processees and their semantics called "observational semantics". We wanted also to better formalize the interactions between these processes in order to be able to optimize their communications
in particular by formalizing the introduction of Tracer Driver and Analyser Manager respectively added to the oberserved and observing processes. Finally the question of extending the domain of
application of the approach to other domains than constraints solvers is considered by introducing the concept of "generic obervational semantics" and "generic trace scheme".
This work is the continuation of the work started with Mireille Ducassé and Ludovic langevine on these topics during the OADymPPaC project. First developments have been published in
Global constraint of scheduling with concentration constraints
François
Fages
Andras
Kovacs
Julien
Martin
Jean-Christophe
Reussner
Aurélie
Strobbe
Global constraints provide a way to introduce redundant constraints from which tighter propagation mechanisms may be infered, for instance by re-using a wide area of algorithms from graph
theory. In
In collaboration with TOTAL, we study an optimization problem of crude oil blending in refineries. The problem is to find a plan of transfer and mixing of crude oil, from the boats to the
distillation units through storage and mixing tanks, under various constraints of product concentrations, continuous distillation, schedule, and optimization criteria. This NP-hard problem is
the combination of a planning problem (for choosing the transfer tasks and quantities) with a scheduling problem.
Previous approaches based on Mixed-Integer Linear Programming (MILP) rely on a discretization of time that may give infeasible or suboptimal solutions. On the other hand, a constraint
programming approach is appealing for its capability to treat efficiently the scheduling part of the problem without time discretization, We have developed such a solution with ILOG tools, by
introducing a global constraint of scheduling with concentration constraints, and using heuristics for the planning part of the problem. A filtering algorithm for this global constraint is
currently developed as a generalization of the multi-resource cumulative constraint.
Because of the importance of the heuristics part on real-size data, we have also considered a constraint-based local search method for this problem
Similar methods are also investigated in collaboration with Evelyne Lutton (COMPLEX Team) for multi-user distributed resource allocation problems and for the implementation of the CONSENSUS
system.
Continuous dynamics in BIOCHAM
Laurence
Calzone
François
Fages
Sriram
Krishnamachari
Sylvain
Soliman
[BIOCHAM](the BIOCHemical
Abstract Machine) is a software environment for modeling biochemical systems
In the differential semantics, the time series obtained from numerical integration allows us to use model-checking techniques for LTL queries with numerical constraints
Current work involves:
using this model-checking features for learning parameter values, as explained in section
identifying precisely which fragment of first-order LTL is wanted;
formalizing, as an abstraction, the relationship between the continuous and boolean semantics;
defining more precisely the relationship with the stochastic semantics;
using bifurcation theory to gain information about possible parameter values for certain behaviors of the system, and elucidating how to connect LTL with ``types of bifurcations''.
Inferring reaction rules in BIOCHAM
Laurence
Calzone
Nathalie
Chabrier-Rivier
François
Fages
Sylvain
Soliman
The formalization of biomolecular interactions in BIOCHAM syntax, and the specification of the observed behavior of the system in temporal logic (CTL or LTL) make it possible to develop
machine learning algorithms to automatically correct or complete existing models. It is worth noting that structural learning of reaction rules from temporal logic properties is quite new, both
from the machine learning perspective and from the systems biology perspective.
In the framework of the APrIL II STREP, we first applied state-of-the-art inductive logic programming tools to simple reachability properties. The objective was to complete a boolean BIOCHAM
model to satisfy a CTL specification. We developed a more powerful search algorithm for learning new reaction rules from general temporal logic properties. This algorithm evolved into a theory
revision algorithm, taking into account that ACTL and ECTL formulae (i.e. those where the only path operator is respectively
Aand
E) can guide the search for the addition or deletion of rules. In the quantitative case, the same strategy can be applied to learn kinetic parameter values. Once the
biological properties of a system under various initial conditions, are described both qualitatively and quantitatively with LTL formulae and constraints, an enumerative algorithm can be
applied to estimate parameter values by model-checking
These algorithms are currently improved in BIOCHAM by applying various optimizations, such as :
compressing the simulation trace used by the LTL model checker with constraints for searching parameter values;
using the type information contained in reaction models
improving the data structures of BIOCHAM code, in the spirit of its migration from Prolog to SiLCC;
exploring inductive logic programming techniques, through a translation of Biocham models to Prolog and Stochastic Logic Programs code.
Coupled model of the cell cycle and the circadian cycle in mammalian cells
Laurence
Calzone
Sriram
Krishnamachari
Sylvain
Soliman
Recent advances in cancer chronotherapy techniques support the evidence that there exist some links between the cell and the circadian cycles. Both cycles have been successfully modeled,
however, as of today, there are no precise models describing the coupling of the two cycles. One purpose of a coupled model is to better understand how to efficiently target malignant cells
depending on the phase of the day. This is at the heart of our participation in the EU STREP project TEMPO.
Our model
The BIOCHAM feature for learning parameter values under period constraints was introduced for this study. This allowed us to compute the domain of entrainment of the cell cycle by the
circadian cycle. This domain mainly depends on the strength of the WEE1 kinase, regulated by circadian genes, on the activity of MPF. We now intend to incorporate other links, based on evidence
from the literature: through the protein c-MYC for instance; and to make the generic cell cycle model more complete in order to see the effect of these other links.
![]() -calculus, Join-calculus and their derivatives
-calculus, Join-calculus and their derivatives