Overall Objectives
Overall objectives
medical imaging
registration
image segmentation and analysis
neuroimaging
statistical analysis in medical imaging
management of information in medical imaging
3D free-hand ultrasound
image-guided intervention
clinical neurosciences
Since 1970s, medical imaging is a very rapidly growing research domain; the last three decades have shown a rapid evolution of the dimension and quantity of data physicians have to work
with. The next decade will follow this evolution by adding not only new spatio-temporal dimensions to the image data produced and used in a clinical environment but also new scales of analysis
(nano or micro biological and molecular images to macro medical images). Another evolution will also consist in adding new effectors during image-guided interventional procedures (surgery,
interventional radiology...). The classical way of making use of these images, mostly based on human interpretation, becomes less and less feasible. In addition, the societal pressure for a
cost effective use of the equipments on the one hand, and a better traceability and quality insurance of the decision making process on the other hand, makes the development of advanced
computer-assisted medical imaging systems more and more essential. According to this context, our research team is devoted to the development of new processing algorithms in the context of
medical image computing and computer-assisted interventions: image fusion (registration and visualization), image segmentation and analysis, management of image-related information ... In this
very large domain, our work is primarily focused on clinical applications and for the most part on head and brain related diseases.
Research activities of the
VisAGeSteam are concerned with the development of new processing algorithms in the field of medical image computing and computer assisted
interventions: image fusion (registration and visualization), image segmentation and analysis, management of image related information ... Since this is a very large domain, for seek of
efficiency, the application of our work will be primarily focused on clinical aspects and for the most part on head and neck related diseases. Our research efforts mainly concern:
The field of image fusion and image registration (rigid and deformable transformations) with a special emphasis on new challenging registration issues especially when statistical
approaches based on joint histogram cannot be used or when the registration stage has to cope with loss or appearance of material (like in surgery or in tumor imaging for instance).
The field of image segmentation and structure recognition, with a special emphasis on the difficult problems of
i) image restoration for new imaging sequences (new Magnetic Resonance Imaging protocols, 3D ultrasound sequences ...), and
ii) structure segmentation and labelling based on shape and statistical information.
The field of image analysis and statistical modelling with a new focus on Voxel Based Analysis (VBA) and group analysis problems. A special attention will be given also to the
development of advanced frameworks for the construction of probabilistic atlases since this complicated problem is still only partially solved.
The field of information management in neuroimaging following the Neurobase project, for the development of distributed and heterogeneous medical image processing systems
(
[
http://www.irisa.fr/visages/neurobase]).
Concerning the application domains, we emphasize our research efforts on the neuroimaging domain with two up-front priorities: Image Guided Neurosurgery and Image Analysis in Multiple
Sclerosis, while developing new ones especially in the interventional aspects (per-operative imagery, robotics...).
Scientific Foundations
Introduction
The scientific objectives of our team, concern the development of new medical image computing methods, dealing with image fusion (registration and visualization), image segmentation and
analysis, and management of image-related information.
In addition, since these methods are devoted (but not specific) to solve actual medical applications, a constant concern is to build an evaluation framework at each stage of the
methodological development process. Therefore, this topic is present as a transversal concern among the generic developments and the applications.
Registration
Rigid registration
deformable registration
similarity measures
Image registration consists in finding a geometrical transformation in order to match
nsets of images. Our objective is to work both, on rigid registration methods in order to develop new similarity measures for new imaging modalities, and on deformable registration to
address the problem of tissue dissipation.
The registration between two images can be summarized by the expression

where
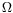
sand
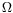
tare respectively the two homologous sets of features respectively extracted from the source and the target images. These sets represent the two images in the registration process. They
can be very different in nature, and can be deduced from a segmentation process (points, contours, crest lines ...) or directly from the image intensities (e.g. the joint histogram).
 is the transformation, (
is the transformation, (
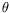
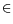
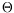 being the set of parameters for this transformation),
being the set of parameters for this transformation),
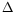 is the cost (or similarity) function, and
is the cost (or similarity) function, and
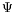 is the optimization method. {
is the optimization method. {
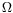 ,
,
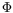 ,
,
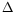 ,
,
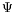 } are the four major decisive factors in a registration procedure, the
set
} are the four major decisive factors in a registration procedure, the
set
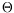 being a priori defined. In addition to new evolutions of these factors, a constant concern is to propose a methodology for validating this registration procedure. We already have been
largely involved in these aspects in the past and will maintain this effort
being a priori defined. In addition to new evolutions of these factors, a constant concern is to propose a methodology for validating this registration procedure. We already have been
largely involved in these aspects in the past and will maintain this effort
In the domain of
rigid registration, our research is more focused on new problems coming from the applications. For instance, the mono and multimodal registration of ultrasound images is still an open
problem. In this context we are working in looking at new similarity measures to better take into account the nature of the echographic signal. Similarly, in the interventional theatre, new
matching procedures are required between for instance video, optical or biological images and the pre-operative images (CT, MRI, SPECT/PET, Angiography ...). Some of these problems can be very
challenging. For a number of new applications, there are no existing solutions to solve these problems (e.g. fusion of biological images with interventional images and images coming from the
planning).
In many contexts, a rigid transformation cannot account for the underlying phenomena. This is for instance true when observing evolving biological and physiological phenomena. Therefore,
deformable registrationmethods (also called non-rigid registration) are needed
Non-rigid registration algorithms benefit from the incorporation of statistical priors. These statistical priors can be expressed locally (for instance through a statistical analysis of
segmented shapes) or globally (by learning statistics about deformation fields directly). Statistical priors (local and global) are useful to capture probable or relevant deformations.
Non-rigid registration methods can be broadly sorted in two classes: geometric methods that rely on the extraction and matching of sparse anatomical structures and photometric methods
that rely on image intensities directly. These two kinds of methods have their advantages and drawbacks. We are working on further cooperative approaches where information of different
nature (global, hybrid and local) could be mixed in an elegant mathematical way.
Finally, our research is focused on a better modeling of the problems, mainly in two directions: firstly the relationship between the observed data (image intensities) and the variables
(registration field) should be better understood. This leads to more adapted similarity measures in specific application contexts (for instance when registering ultrasound images).
Secondly, specific modeling of the deformation field is useful in specific contexts (for instance when matter is disappearing, fluid mechanics models will be more adapted than classical
regularized deformation fields).
Image segmentation and analysis
image restoration
deformable shape models
level sets
3D ultrasound
MRI
This topic is very classical in computer vision. For the concern of medical image computing, we are focusing on the development of new tools devoted to the restoration of corrupted images
coming from the sources and to the segmentation of anatomical structures based on deformable shape models.
Statistical methods for image restoration:New applications of medical imaging systems are parallel to the development or the evolution of new machinery which come with specific artifacts
that are still only partially understood. This is the case for instance with high field MRI, 3D ultrasound imaging or other modalities. With regards to the images to process and analyze, these
artifacts translate into geometric or intensity distortions that drastically affect not only the visual interpretation, but also most of the segmentation or registration algorithms, and the
quantitative measures that follow. A better comprehension of these artifacts necessitates an increased dialogue between the physicists (who make the images), the computer scientists (who
process the images) and the clinicians (who interpret the images). This should lead to define new, specifically-designed algorithms, based on statistical models taking into account the physics
of the acquisition.
Segmentation using deformable shapes:We aim at proposing a generic framework to build probabilistic shape models in a 3D
+
tspace applied to biomedical images with a particular emphasis on the problem of modeling anatomical and functional structures in neuroimaging (functional
delineations, cortical or deep brain structures). Based on our previous contributions in this domain
Statistical analysis in medical imaging
voxel based analysis
probabilistic brain atlas
image classification
group analysis
Nowadays, statistical analysis occupies a central place for the study of brain anatomy and function in medical imaging. It is indeed a question of exploiting huge image data bases, on which
we look to reveal the relevant information: measure the anatomical variability to discover better what deviates from it, to measure the noise to discover an activation, etc., in brief, to
distinguish what is statistically significant of what is not.
Statistical methods for voxel-based analysis:Statistical analysis tools play a key role in the study of the anatomy and functions of the brain. Typically, statisticians aim at extracting
the significant information hidden below the noise and/or the natural variability. Some specific tools exist for the comparison of vector fields or geometrical landmarks. Some others have been
developed for the analysis of functional data (PET, fMRI...). Thus, statistics are generally either spatial, or temporal. There is an increasing need for the development of statistics that
consider time and space simultaneously. Applications include the follow-up of multiple sclerosis in MR images or the tracking of a deformable structure in an ultrasound image sequence.
Probabilistic atlases:One of the major problems in medical image analysis is to assist the clinician to interpret and exploit the high dimensionality of the images especially when he/she
needs to confront his/her interpretation with "classical" cases (previous or reference cases). A solution to deal with this problem is to go through the use of an atlas which can represent a
relevant
a prioriknowledge. Probabilistic atlases have been studied to tackle this problem but most of the time they rely on global references which are not always relevant or precise enough, to
solve some very complex problems like the interpretation of inter-individual variations of brain anatomy and functions. Based on our previous work proposing a cooperation between global and
local references to build such probabilistic atlases
Classification and group analysis:One of the major problems in quantitative image analysis is to be able to perform clustering based on descriptors extracted from images. This can be
done either by using supervised or unsupervised algorithms. Our objectives is to develop statistical analysis methods in order to discriminate groups of data for clinical and medical research
purposes (e.g. pathologic
vs. normal feature, male
vs.female, right-handed
vs.left-handed, etc.), these data may come from descriptors extracted by using image analysis procedures (e.g. shapes, measurements, volumes, etc.).
Management of information in medical imaging
mediation
wrapper
workflows
ontology
web services
There is a strong need of a better sharing and a broader re-use of medical data and knowledge in the neuroimaging field. One of the most difficult problems is to represent this information
in such a way that the structure and semantics are shared between the cognitive agents involved (i.e. programs and humans). This issue is not new, but the recent evolution of computer and
networking technology (most notably, the Internet) increases information and processing tools sharing possibilities, and therefore makes this issue prevailing. The notion of ``semantic web''
denotes a major change in the way computer applications will share information semantics in the future, with a great impact on available infrastructures and tools. In coherence with the rest of
our research topics, we are focussing on brain imaging. This deals with accessing, referring to, and using knowledge in the field of brain imaging, whatever the kind of knowledge - either
general knowledge (e.g. models of anatomical structures, ``know-how'' knowledge such as image processing tools), or related to individuals (such as a database of healthy subjects' images). This
covers both information of a numerical nature (i.e. derived from measurements such as images or 3D surfaces depicting anatomical features), of a symbolic nature (such as salient properties,
names - referring to common knowledge - and relationships between entities), as well as processing tools available in a shared environment. Two major aspects are considered: (1) representing
anatomical or anatomo-functional data and knowledge and (2) sharing neuroimaging data and processing tools.
Application Domains
Neuroimaging
neuroimaging
clinical neuroscience
image-guided surgery
multiple sclerosis
preoperative imaging
3D ultrasound
multispectral MRI
brain atlas
One research objective in neuroimaging is the construction of anatomical and functional cerebral maps under normal and pathological conditions.
Many researches are currently performed to find correlations between anatomical structures, essentially sulci and gyri, where neuronal activation takes place, and cerebral functions, as
assessed by recordings obtained by the means of various neuroimaging modalities, such as PET (Positron Emission Tomography), fMRI (Functional Magnetic Resonance Imaging), EEG
(Electro-EncephaloGraphy) and MEG (Magneto-EncephaloGraphy). Then, a central problem inherent to the formation of such maps is to put together recordings obtained from different modalities and
from different subjects. This mapping can be greatly facilitated by the use of MR anatomical brain scans with high spatial resolution that allows a proper visualization of fine anatomical
structures (sulci and gyri). Recent improvements in image processing techniques, such as segmentation, registration, delineation of the cortical ribbon, modeling of anatomical structures and
multi-modality fusion, make possible this ambitious goal in neuroimaging. This problem is very rich in terms of applications since both clinical and neuroscience applications share similar
problems. Since this domain is very generic by nature, our major contributions are directed towards clinical needs even though our work can address some specific aspects related to the
neuroscience domain.
Multiple sclerosis:Over the past years, a discrepancy became apparent between clinical Multiple sclerosis (MS) classification describing on the one hand MS according to four different
disease courses and, on the other hand, the description of two different disease stages (an early inflammatory and a subsequently neurodegenerative phase). It is to be expected that
neuroimaging will play a critical role to define
in vivothose four different MS lesion patterns. An
in vivodistinction between the four MS lesion patterns, and also between early and late stages of MS will have an important impact in the future for a better understanding of the natural
history of MS and even more for the appropriate selection and monitoring of drug treatment in MS patients. Since MRI has a low specificity for defining in more detail the pathological changes
which could discriminate between the different lesion types, but a high sensitivity to detect focal and also widespread, diffuse pathology of the normal appearing white and grey matter, our
major objective within this application domain is to define new neuroimaging markers for tracking the evolution of the pathology from high dimensional data (e.g. nD+t MRI). In addition, in
order to complement MR neuroimaging data, we ambition to perform also cell labeling neuroimaging (e.g. MRI or PET) and to compare MR and PET data using standard and experimental MR contrast
agents and radiolabeled PET tracers for activated microglia (e.g. USPIO or PK 11195). The goal is to define and develop, for routine purposes, cell specific and also quantitative imaging
markers for the improved
in vivocharacterization of MS pathology.
Modeling of anatomical and anatomo-functional neurological patterns:The major objective within this application domain is to build anatomical and functional brain atlases in the context
of functional mapping for pre-surgical planning and for the study of neurodegenerative brain diseases (Multiple sclerosis, Epilepsy, Parkinson or even Alzheimer). This is a very competitive
research domain; our contribution is based on our previous works in this field
An additional objective within this application domain is to find new descriptors to study the brain anatomy and/or function (e.g. variation of brain perfusion, evolution in shape and size
of an anatomical structure in relation with pathology or functional patterns, computation of asymmetries ...). This is also a very critical research domain, especially for many
neurodegenerative brain diseases (Epilepsy or Alzheimer for instance).
Epilepsy:The principle of epilepsy surgery is to remove the Epileptic Zone (EZ) (area of the brain where epileptic seizures are originating). The anatomical determination of this EZ is
individualized, and surgery will be therefore individually tailored. To delineate this EZ, different sources of information are used and a congruence of several explorations is needed. Some are
static, such as MRI and PET, and some may reflect the spatio-temporal dynamics of the seizures. Integration of multimodal information about brain perfusion (ictal and interictal SPECT),
metabolism (PET-F18FDG), anatomy (MRI, DTI), as well as direct recording of electrical activity (MEG/EEG) may improve significantly on its own the way epileptic patients are explored and
treated. Although none of these modalities added a significant contribution to this area, SPECT/PET and MEG/EEG could help localizing the EZ in temporal lobe epilepsy and limit the use of depth
electrodes recordings, especially when focussing more specifically on the particular role of sub-cortical structures (such as thalamus, caudate nucleus, pallidum, etc.). From this standpoint,
our goal is to tackle several of these questions? such as:
What is the role of sub-cortical structures in temporal and frontal epilepsy? How do these observations correlate with depth electrodes recordings? How could this knowledge impact on
treatment decisions (chemically on basal ganglia, or surgically with deep brain stimulations ?
What is the optimum use of the various imaging techniques available, in order to examine the various parts of an epileptic network, before performing a decision)?
Image guided intervention
Image-guided neurosurgical procedures rely on complex preoperative planning and intraoperative environment. This includes various multimodal examinations: anatomical, vascular, functional
explorations for brain surgery and an increasing number of computer-assisted systems taking place in the Operating Room (OR). Hereto, using an image-guided surgery system, a rigid fusion
between the patient's head and the preoperative data is determined. With an optical tracking system and Light Emitting Diodes (LED), it is possible to track the patient's head, the microscope
and the surgical instruments in real time. The preoperative data can then be merged with the surgical field of view displayed in the microscope. This fusion is called ``augmented reality''.
Unfortunately, the assumption of a rigid registration between the patient's head and the preoperative images only holds at the beginning of the procedure. This is because soft tissues tend
to deform during the intervention. This is a common problem in many image-guided interventions, the particular case of neurosurgical procedures can be considered as a representative case. Brain
shift is one manifestation of this problem but other tissue deformations can occur and must be taken into account for a more realistic predictive work.
Within this application domain, we aim at developing systems using surgical guidance tools and real-time imagery in the interventional theatre. This imagery can come from video (using
augmented reality procedures), echography or even interventional MRI, biological images or thermal imagery in the future.
Per-operative imaging in neurosurgery:Our major objective within this application domain is to correct for brain deformations that occur during surgery. Neuronavigation systems make it
now possible to superimpose preoperative images with the surgical field under the assumption of a rigid transformation. Nevertheless, non-rigid brain deformations, as well as brain resection,
drastically limit the efficiency of such systems. The major objective here is to estimate brain deformations using 3D ultrasound and video information.
Modeling of surgical gesture expertise:Our objective is to show how the formalization of the medical expertise could improve both the planning and the surgery itself. One way is to rely
on previously defined generic model describing surgical procedures. From a data base of surgical cases described by the generic model and from a limited set of parameters related to the patient
(i.e. extrinsic parameters), the closest surgical case can be retrieved in order to assist the surgical planning. Similarly, global surgical scenarii representing main categories of surgical
procedures could be classified according to extrinsic parameters (coming from the current case) and retrieved from the database. New experiences based on this procedure could then feed the
surgical modelling. Another issue would be to use the knowledge extracted from the data base to pre-fetch the image processing procedures (to speed up or tune processing workflows
parameters).
Robotics for 3D echography:This project is conducted jointly with the Lagadic project-team. The goal is to use active vision concepts in order to control the trajectory of a robot based
on the contents of echographic images and video frames (taken from the acquisition theatre). Possible applications are the acquisition of echographic data between two remote sites (the patient
is away from the referent clinician) or the monitoring of interventional procedure like biopsy or selective catheterisms.
3D free-hand ultrasound: Our major objective within this application domain is to develop efficient and automatic procedures to allow the clinician to use conventional echography to
acquire 3D ultrasound and to propose calibrated quantification tools for quantitative analysis and fusion procedures. This will be used to extend the scope of view of an examination.
Software
Introduction
Our objectives concerning the software development and dissemination are directed to the set-up of a software platform at the University Hospital in order to deploy new research advances and
to validate them in the clinical context with our local partners. We intend to disseminate our results via a free software distribution. Complying with both objectives requires software
engineering resources, which could be partially covered in the short term by a current PRIR application "PlogICI", but a longer term alternative already needs to be foreseen.
VistaL
VistaL is a software platform of 3D and 3D+t image analysis allowing the development of generic algorithms used in different contexts (rigid and non-rigid registration, segmentation,
statistical modelling, calibration of free-hand 3D ultrasound system and so on). This software platform is composed of generic C++ template classes (Image3D, Image4D, Lattice and so on) and a
set of 3D/3D+t image processing libraries. VistaL is a multi-operating system environment (Windows, Linux/Unix...). VistaL APP registration number is: IDDN.FR.001.200014.S.P.2000.000.21000.
Romeo
Romeo (
RObust
Multigrid
Elastic registration based on
Optical flow) is a non-rigid registration algorithm based on optical-flow. Romeo is developed using Vistal (C++ template classes described above). Romeo estimates a regularized
deformation field between two volumes in a robust way: two robust estimators are used for both the data term (optical flow) and the regularization term (smoothness of the field). An efficient
multiresolution and multigrid minimization scheme is implemented so as to estimate large deformations, to increase the accuracy and to speed up the algorithm
Juliet
Juliet (
Joint
Use of
Landmarks and
Intensity for
Elastic regis
Tration) is a non-rigid registration algorithm that is built on the Romeo software. Juliet makes it possible to incorporate sparse constraints deduced from the matching of anatomical
structures such as cortical sulci for instance. A sparse deformation field is introduced as a soft constraint in the minimization to drive the registration process. A robust estimator is used
so as to limit segmentation errors and false matching
Tulipe
TULIPE (
Three dimensional
ULtrasound reconstructioin
Incorporating Prob
Etrajectory) was developed using Vistal and is a registered at APP under IDDN.FR.001.120034.S.A.2006.000.21000. 3D freehand ultrasound is a technique based on the acquisition of B-scans,
which can be parallel or not, whose position in 3D space is known by a 3D localizer (optic or magnetic) attached to the probe. From these irregularly distributed B-scans and their positions, a
regular 3D lattice volume can be reconstructed. This reconstruction step is needed to apply conventional 3D computer vision algorithms like volumetric registration and segmentation, but is
still an acute problem with regards to computation time and reconstruction quality. Tulipe explicitly takes into account the 3D probe trajectory. In the classical distance weighted
interpolation, the interpolation kernel is composed of the orthogonal projections of the current point on the closest B-scans. In Tulipe, the interpolation kernel is composed of intersections
between the probe trajectory (passing through the current point) and the closest B-scans
![]()
![]()
![]()
![]() is the transformation, (
is the transformation, (
![]()
![]()
![]() being the set of parameters for this transformation),
being the set of parameters for this transformation),
![]() is the cost (or similarity) function, and
is the cost (or similarity) function, and
![]() is the optimization method. {
is the optimization method. {
![]() ,
,
![]() ,
,
![]() ,
,
![]() } are the four major decisive factors in a registration procedure, the
set
} are the four major decisive factors in a registration procedure, the
set
![]() being a priori defined. In addition to new evolutions of these factors, a constant concern is to propose a methodology for validating this registration procedure. We already have been
largely involved in these aspects in the past and will maintain this effort
,
.
being a priori defined. In addition to new evolutions of these factors, a constant concern is to propose a methodology for validating this registration procedure. We already have been
largely involved in these aspects in the past and will maintain this effort
,
.