Overall Objectives
Introduction
Constraint Logic Programming supports a great ambition for programming: the one of making of programming essentially a modeling task, with equations, constraints and logical formulas.
Constraint Programming is a field born during the mid 80s from Logic Programming, Linear Programming coming from Operations Research, and Constraint Propagation techniques coming from
Artificial Intelligence. Its foundation is the use of relations on mathematical variables to compute with partial information. The successes of Constraint Programming for solving combinatorial
optimization problems, from pure problems to real problems in industry or commerce, owe much to the bringing of , on the one hand, new local consistency techniques, and, on the other hand,
declarative languages which allow control on the mixing of heterogeneous resolution techniques: numerical, symbolic, deductive and heuristic.
The "Contraintes" group investigates the logical foundations, design, implementation, programming environments and applications of constraint programming languages. The study of Concurrent
Constraint languages is a core aspect of the project as they provide a conceptual framework for analyzing different issues of constraint programming, like constraint resolution techniques,
concurrent modeling, reactive applications, etc.
The main application domains investigated are combinatorial optimization problems and computational systems biology. In bioinformatics, our objective is not to work on structural biology
problems which has been the main trend up to now, but to attack the great challenge of systems biology, namely to model the function, activity and interaction of molecular processes in living
cells, with logic programming concepts and program verification techniques.
Highlight: Model-checking generalized to constraint solving
In
D, and by computing a validity domain for the variables rather than a truth value for the formula. When
Dis a metric space, this allows us to define a continuous degree of satisfaction for a temporal logic formula in a given Kripke structure, opening up the field of model-checking to
optimization.
This work originates from previous work in
[BIOCHAM]on reverse engineering problems coming
from systems biology. The algorithm used in BIOCHAM for solving Linear Time Logic constraints over the reals (LTL(
R)) has been generalized to a fixpoint algorithm for branching time logics, namely for the quantifier-free first-order Computation Tree Logic over some arbitrary domain
D(QFCTL(
D)).
Our result shows that any constraint solver over some domain
Dcan be lifted to a QFCTL(
D) constraint solver over a finite Kripke structure over
D, and that optimization techniques can be used to synthesize or parameterize deterministic as well as non-deterministic systems, in order to satisfy high-level QFCTL(
D) specifications.
Highlight: Robustness measure of temporal logic properties
Robustness is the capacity of a system to maintain a function in the face of perturbations. It is essential for the correct functioning of natural as well as engineered biological systems.
Robustness is generally defined in an
ad-hocproblem-dependent manner, thus hampering the fruitful development of a theory of biological robustness, advocated by Kitano [Mol Syst Biol, 3:137, 2007].
In
[BIOCHAM]2.8 for the automated estimation of the
robustness of a given behavior with respect to a given set of perturbations. The applicability and biological relevance of our approach is demonstrated by testing and improving the robustness
of the timed behavior of a synthetic transcriptional cascade that could be used as a biological timer for synthetic biology applications.
Scientific Foundations
Concurrent constraint programming
The class of Concurrent Constraint programming languages (CC) was introduced a decade ago by Vijay Saraswat as a unifying framework for constraint logic programming and concurrent logic
programming. The CC paradigm constitutes a representative abstraction of constraint programming languages, and thus allows a fine grained study of their fundamental properties.
CC generalizes the Constraint Logic Programming framework (CLP) by introducing a synchronization primitive, based on constraint entailment. It is a model of concurrent computation, where
agents communicate through a shared store, represented by a constraint, which expresses some
partial informationon the values of the variables involved in the computation. The variables play the role of transmissible dynamically created communication channels.
One of the big successes of CC has been the simple and elegant reconstruction of finite domain constraint solvers, and the cooperation of several models to solve a single combinatorial
problem. On the other hand, to use CC for programming reactive applications forces one to abandon the hypothesis of monotonic evolution of the constraint store; this is a strong motivation for
new extensions of CC languages.
There are strong completeness theorems relating the execution of a CLP program and its translation in classical logic, which provide smooth reasoning techniques for such programs. However
these theorems are broken by the synchronization operation of CC. Looking for a logical semantics of CC programs in the general paradigm of logic programming,
program = logical formula,
execution = proof search,
leads to a translation in Jean-Yves Girard's linear logic. This allows the recovery of some completeness results about successes and stores; even suspensions may be
characterized with the non-commutative logic of Ruet and Abrusci.
It is thus possible to address important issues for Constraint Programming:
verifying CC programs;
combining CLP and state-based programming;
dealing with local search inside a global constraint solving procedure.
The last two cases rely on a natural extension of CC languages, called Linear Concurrent Constraint languages (LCC), which simply replaces constraint systems built onto classical logic by
constraint systems built onto linear logic. This allows us to represent state changes thanks to the consumption of resources during the synchronization action, modeled by the linear
implication.
Constraint solvers
Our domains of application use quite different constraint systems:
finite domains (bounded natural numbers): primitive constraint of some finite domain membership, numerical, symbolic, higher order and global constraints;
reals: polyhedral libraries and Simplex algorithm for linear constraints and interval methods otherwise;
terms: subtyping constraints and ontologies;
temporal constraints: CTL and LTL formulae, either propositional or with numerical constraints.
The project works on constraint resolution methods and their cooperation. The main focus is the declarativeness of the constraint solver (e.g. implemented by CHR rules), the efficiency of
constraint propagation methods, the design of global constraints and the combination of constraint propagation with heuristic search.
Computational systems biology
Systems biology is a cross-disciplinary domain involving biology, computer science, logics, mathematics, and physics to elucidate the high-level functions of the cell from their biochemical
bases at the molecular level.
At the end of the Nineties, research in Bioinformatics evolved, passing from the analysis of the genomic sequence to the analysis of post-genomic interaction networks (expression of RNA and
proteins, protein-protein interactions, etc). The complexity of these networks requires a large research effort to develop symbolic notation and analysis tools for biological processes and
data. In order to scale-up, and get over the complexity walls to reason about biological systems, there is a general feeling that beyond providing tools to biologists, computer science has much
to offer in terms of concepts and methods.
We are interested in the modeling and analysis of complex molecular processes in the cell, at different levels of abstraction, qualitative and quantitative. The most original aspect of our
research can be summarized by the following identifications :
biological model = state transition system,
biological property = temporal logic formula,
validation = model-checking,
inference = constraint solving.
Our main research axis is thus the application of logic programming concepts and program verification techniques to the analysis of complex biochemical processes in the cell.
Application Domains
Combinatorial optimization problems
The number and economic impact of combinatorial optimization problems found in the industrial world are constantly increasing. They cover:
resource allocation;
placement, bin packing;
scheduling;
planning;
transport;
etc.
The last forty years have brought many improvements in Operations Research resolution techniques. In this context, Constraint Programming can be seen as providing, on the one hand, local
consistency techniques that can be applied to various numerical or symbolic constraints, and on the other hand, declarative languages. This last point is crucial for quickly developing complex
combinations of algorithms, which is not possible without a language with a high level of abstraction. It allowed for better results, for instance in scheduling problems, than traditional
methods, and is promised to an even better future when thinking about the cooperation of global resolution, local consistency techniques and search methods.
The project builds upon its knowledge of CC languages, constraint solvers and their implementation to work in these directions. The LCC paradigm offers at the same time a theoretical
framework for analysis, and a valuable guide for practical language design and implementation. The work on programming environments helps to integrate the Constraint Programming tools into this
application domain.
The European FP6 Strep project
[Net-WMS]that we coordinate, makes us work on pure and
non-pure bin packing problems combining discrete geometry constraints with physical, common sense and packing business rules, in the context of warehouse management systems for the automotive
industry. In this context, we have developed a rule-based modeling language, called
[Rules2CP], to express requirements in a
declarative and flexible manner, and its compiler to efficient constraint programs using a global constraint dedicated to geometrical placement problems in high dimensions, together with
reified finite-domain constraints.
In-silico cell processes
In 2002 , we started a Collaborative Research Initiative ARC CPBIO on “Process Calculi and Biology of Molecular Networks”. By working on well understood biological models, we sought:
to identify in the family of competitive models coming from the Theory of Concurrency and from Logic Programming (Constraint Logic Programming, Concurrent Constraint
languages and their extensions to discrete and continuous time, TCC, HCC), the ingredients of a language for the modular and multi-scale representation of biological processes;
to provide a series of examples of biomolecular processes transcribed in formal languages, and a set of biological questions of interest about these models;
to design and apply to these examples formal computational reasoning tools for the simulation, the analysis and the querying of the models.
This work lead us to the design and implementation of the Biochemical Abstract Machine
[BIOCHAM]that has the unique feature of providing
formal languages corresponding to different qualitative and quantitative levels of abstraction for, on the one hand, modeling biomolecular interaction diagrams with reaction rules, and on the
other hand, modeling the biological properties of the system in temporal logic. This double formalization of both the model and the biological properties of the system at hand has opened
several new research avenues on the design and systematic validation of biological models.
In the 6th PCRD STREP project
[APrIL II](2004-2007) the focus was on probabilistic inductive
logic programming for metabolic networks and we developed semi-automatic methods for model completion/revision from temporal logic specification of the system's behavior as observed in
biological experiments. In the Network of Excellence
[REWERSE](2004-2008), the focus was on the application of the
new Semantic Web technologies based on rules and constraints to bioinformatics. In this context, we developped type inference and abstract interpretation techniques to relate biological models
at different levels of abstraction, providing a formal ground for reusing and combining models available on the web. In the ARC
[MOCA](2006-2007) on “MOdularity, Compositionality
and Abstraction in gene and protein networks”, we studied with our partners the formal links between logical and numerical models of some parts of the cell cycle control, and modular
decompositions based on control theory considerations
Currently, we develop these technologies and apply them to new biological questions which we investigate in partnerships with biologists in three projects. First, the EU STREP project
[TEMPO](2006-2009) on “temporal genomics for patient
tailored chronotherapeutics”, coordinated by Francis Lévi INSERM Villejuif, where, in partnership with Jean Clairambault of the BANG project-team, we develop coupled models of the cell cycle,
the circadian cycle and the effect of cytotoxic drugs in cancer therapies using BIOCHAM.
Second, the INRA AgroBi project
[INSIGHT], coordinated by Eric
Reiter INRA Tours, where, in partnership with Frédérique Clément of the SISYPHE project-team, we develop models of FSH and GPCR signaling networks in mammalian cells. This research is now part
of the AE
[REGATE]coordinated by F.
Clément SISYPHE.
Third, the AE COLAGE coordinated by Hughes Berry of the ALCHEMY project-team before his move to Lyon, with François Taddei, Ariel Lindner, INSERM Paris Necker, Hidde de Jong, Delphine
Ropers, IBIS, J.L. Gouzé, and Madalena Chaves, COMORE, where we investigate the possibilities to control and reprogram growth and aging in bacteria
E. coliusing synthetic biology approaches.Along the same line of research, but in a different context, we collaborate with Pascal Hersen and Samuel Bottani, biophysicists at the Matière
and Systèmes complexes lab, CNRS/Paris Diderot University, to design a prototypic platform and develop control software for the real-time control of gene expression in yeast.
Software
BIOCHAM
François
Fages
Aurélien
Rizk
Sylvain
Soliman
The Biochemical Abstract Machine
[BIOCHAM]is a modeling and validation environment
for molecular systems biology
BIOCHAM is compatible with the Systems Biology Markup Language (
[SBML]) and adds precise semantics to biomolecular interaction
maps at three abstraction levels:
the boolean semantics (presence or absence of molecules),
the differential semantics (concentrations of molecules),
the stochastic semantics (discrete numbers of molecules).
Based on this formal framework, BIOCHAM features:
a compositional rule-based language for modeling biochemical systems, allowing patterns for expressing set of rules in a compact form;
numerical as well as boolean simulators (Rosenbrock's method for the differential semantics, Gillespie's algorithm with tau lipping for the stochastic semantics);
a temporal logic language (CTL for qualitative models and QFLTL(R) with numerical constraints for quantitative models) for formalizing biological properties such as
reachability, checkpoints, oscillations or stability, and checking them automatically with model-checking techniques;
automatic search procedures to infer parameter values, initial conditions and even reaction rules from temporal logic properties of the system;
automatic conservation law detection, through constraint-based structural analysis of the underlying Petri-net.
BIOCHAM was used in synergy with Beta WB and GINsim by the winning team of the
Biological Modelling Competitionof the
Formal Methods in Molecular Biologyworkshop held in Dagstuhl in Feb. 2009.
BIOCHAM is fully implemented in GNU-Prolog and interfaced to the symbolic model checker
[NuSMV]and to the continuous optimization tool
[CMAES].
Rules2CP
François
Fages
Julien
Martin
Thierry
Martinez
[Rules2CP]is a rule-based modelling language for
constraint programming. Rules2CP is under development since 2006, and distributed as open-source since 2009.
Unlike other modelling languages, Rules2CP adopts a single knowledge representation paradigm based on rules without recursion, and a restricted set of data structures based on records and
enumerated lists given with iterators. We show that this is sufficient to model constraint satisfaction problems, together with search strategies where search trees are expressed by logical
formulae, and heuristic choice criteria are defined with preference orderings by pattern-matching on rules' left-hand sides.
The expressiveness of Rules2CP is illustrated with a complete library for packing problems, called PKML, which, in addition to pure bin packing and bin design problems, can deal with common
sense rules about weights, stability, as well as specific packing business rules.
CHRat
Thierry
Martinez
[CHRat]is a modular version of the well
known Constraint Handling Rules language CHR, called for CHRat for CHR with
askand
tell. Inspired by the LLCC framework, this extension of CHR makes it possible to reuse CHRat components both in rules and guards in other CHRat components, and define hierarchies of
constraint solvers. CHRat is a bootstrapped preprocessor for CHR implemented in Prolog.
CLPGUI
François
Fages
[CLPGUI]is a generic graphical user
interface written in Java for constraint logic programming. It is available for GNU-Prolog and SICStus Prolog. CLPGUI has been developed both for teaching purposes and for debugging complex
programs. The graphical user interface is composed of several windows: one main console and several dynamic 2D and 3D viewers of the search tree and of finite domain variables. With CLPGUI it
is possible to execute incrementally any goal, backtrack or recompute any state represented as a node in the search tree. The level of granularity for displaying the search tree is defined by
annotations in the CLP program.
CLPGUI has been mainly developped in 2001 and is distributed as third-party software on GNU-Prolog and SICStus Prolog web sites. In 2009, CLPGUI has been interfaced to Rules2CP/PKML and used
in FP6 Strep Net-WMS with a non-released version.
New Results
Linear Logic Concurrent Constraint Programming and Constraint Handling Rules
François
Fages
Thierry
Martinez
Sylvain
Soliman
We are developing SiLCC an imperative and concurrent constraint programming language based on a single paradigm: the one of Vijay Saraswat's concurrent constraint programming extended with
constraint systems based on Jean-Yves Girard's Linear Logic. From a constraint programming point of view, the unique combination of constraint programming with imperative features opens many
new possibilities, among which:
the capability of programming constraint solvers in the language, making them extensible by the user,
making a fully bootstrapped implementation of a constraint programming language (for the first time since Prolog)
combining constraint reasoning with state change;
embedding program declarations, modules and closures as agents;
proving program correctness using Linear Logic.
The Constraint Handling Rules (CHR) language of Thom Frühwirth shares many similarities with SiLCC. CHR (and its modular version
[CHRat]
In
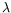 -lifting in functional languages. In conjunction with the equivalence between CHR and CSR with respect to naive operational semantics, these results lead to semantics-preserving
translations from full LCC to CHR and conversely. Immediate consequences of this work include new proofs for CHR linear logic and phase semantics, relying on corresponding results for LCC, plus
an encoding of the
-lifting in functional languages. In conjunction with the equivalence between CHR and CSR with respect to naive operational semantics, these results lead to semantics-preserving
translations from full LCC to CHR and conversely. Immediate consequences of this work include new proofs for CHR linear logic and phase semantics, relying on corresponding results for LCC, plus
an encoding of the
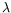 -calculus in CHR.
-calculus in CHR.
Rule-based Modeling of Constraint Satisfaction Problems
François
Fages
Curtis
Fonger
Julien
Martin
Thierry
Martinez
Surinderjeet
Singh
[Rules2CP]
The expressiveness and effectiveness of Rules2CP have been illustrated through the development of the Packing Knowledge Modeling Language PKML as a library on top of Rules2CP, dedicated to
bin packing problems. PKML language is used in the FP6 Strep
[Net-WMS]to study and solve higher-dimensional bin packing
problems and placement constraints, including common sense, physical and industrial requirements expressed by rules
Furthermore, in
Modelling Search in Rules2CP
François
Fages
Julien
Martin
Thierry
Martinez
Surinderjeet
Singh
One originality of
[Rules2CP]as a modeling language is that it
allows us to express search tree and labeling ordering heuristics declaratively by pattern matching on rule left-hand sides' derivation
In order to deal with dynamic ordering criteria, a new compilation scheme for Rules2CP is introduced in
Symmetry Detection and Breaking in Constraint Satisfaction Problems
François
Fages
Valentin
Gatien-Baron
In
Trace Development Methodology
Pierre
Deransart
Rafaël
Oliveira
We worked on a general theory of trace design based on the observation of the way trace files are accumulated as knowledge bases and elaborated in differents fields of activity like software
engeneering, rule based systems and resolution, learning in context, or personal experience storing systems. The state of this work is regularly updated on
One of the important aspect of this research is to give a proper semantics to a trace, called “observational semantics” (OS). We followed two ways: the use of the simple fluent calculus as
formalism for the OS, and the software component modeling approach to combine traces of components
vwith several extensions, in the framework called CHROME-REF
Petri Net Representation of Biological Networks
François
Fages
Faten
Nabli
Sylvain
Soliman
Denis
Thieffry
In
[BIOCHAM]modelling environment.
Other uses of Petri Nets structural properties computation were investigated in
These developments are focused on the applications of the
[CALAMAR]ANR project, where large
models of the E2F/RB pathway and of other cancer-related processes are studied. This project also dwelves, on the formal side, on relationships between different formalisms, whether
BIOCHAM-like reaction-centered, or regulation-centered like logical models
Model Reductions by Graph Transformations
François
Fages
Steven
Gay
Sylvain
Soliman
Denis
Thieffry
Biologists use diagrams to represent interactions between molecular species, and on the computer, diagrammatic notations are also more and more employed in interactive maps. These diagrams
are fundamentally of two types: reaction graphs and positive/negative influence graphs. The analysis of circuits in the influence graphs has been introduced by René Thomas in the late 70's with
necessary conditions for oscillations (homeostasie) and multistability (cell differentiation).
In
In
Analysis of Interlocked Positive Feedback Loops
François
Fages
Sylvain
Soliman
The two element mutual activation and inhibitory positive feedback loops are a common motifs that occur in many biological systems in both isolated and interlocked form, as for example,in
the cell division cycle and thymus differentiation in eukaryotes.
In
Temporal Logic Constraint Solving
François
Fages
Aurélien
Rizk
Temporal logics and model-checking are at the core of
[BIOCHAM]to express biological properties of
complex biochemical systems and automatically verify their satisfaction in both qualitative and quantitative models. In particular, linear time logic constraints (QFLTL(
 )) are used to formalize the global behavior of the system known from biological experiments, and infer the unknown parameter values of the model.
)) are used to formalize the global behavior of the system known from biological experiments, and infer the unknown parameter values of the model.
In
D. We have shown that the QFCTL constraint satisfiability problem is decidable in finite Kripke structures over an arbitrary computation domain with a decidable language of constraints,
i.e. that any constraint solver can be lifted to a temporal logic constraint solver over finite Kripke structures. We have presented a generic QFCTL constraint solver which computes validity
domains for the free variables of a formula, in quadratic time in the number of states. We show that when
Dis a metric space, this allows us to define a continuous degree of satisfaction for a temporal logic formula in a given Kripke structure, opening up the field of model-checking to
optimization, in a very general setting.
Parameter Search w.r.t. Temporal Logic Properties
Grégory
Batt
François
Fages
Sylvain
Pradalier
Sylvain
Soliman
Aurélien
Rizk
In
[BIOCHAM], our method for solving QFLTL(R)
constraints allows us to define a continuous degree of satisfaction of LTL(R) formulae in a given trace, and use it as a fitness function in continuous optimization methods
namely the Covariance Matrix Adaptation Evolutionary Strategy
[CMAES]of Nikolaus Hansen
from the TAO project-team.in order to find unknown parameter values in a model satisfying a set of biological properties formalized in temporal logic
The study of similar metrics and methods for stochastic processes is under investigation.
Robustness Analysis w.r.t. Temporal Logic Properties
Grégory
Batt
François
Fages
Sylvain
Soliman
Aurélien
Rizk
In
[BIOCHAM]v2.8, for computing the continuous degree
of satisfaction of QFLTL(R) formulae. The applicability and biological relevance of our approach is demonstrated by testing and improving the robustness of the timed behavior of a synthetic
transcriptional cascade that could be used as a biological timer for synthetic biology applications. These novel methods are evaluated on models of the cell cycle and of the MAPK signalling
cascade. In , they are used to analyze a gene activation cascade in synthetic biology.
Automated selection of PADE models by symbolic model checking
Grégory
Batt
Qualitative piecewise-affine differential equation (PADE) models offer an attractive formalism for representing genetic regulatory networks in absence of precise information on parameter
values. Predictions on systems behaviors can be obtained in the form of a discrete state transition system simply by using parameter relative order. In this work, we encode in a symbolic way
the transition relation of the graph and use symbolic model checkers to automatically select the set of models whose parameter order is consistent with a set of observations. Because in PADE
systems the dynamics in a state may be defined with respect to the dynamics in neighbouring states, the main difficulty was to obtain a purely state-based encoding of the transition relation,
similar to those used by model checkers. This approach is applied to a synthetic gene network built in
E. colifor
in vivobenchmarking of reverse-engineering and modeling approaches (IRMA). This work is done in collaboration with Hidde de Jong and Michel Page in the IBIS research group (INRIA
Rhône-Alpes/UPMF).
Modeling the Cell Cycle
Elisabetta
De Maria
François
Fages
Aurélien
Rizk
Sylvain
Soliman
Recent advances in cancer chronotherapy techniques support the evidence that there exist some links between the cell cycle and the circadian clock genes
[TEMPO]in collaboration with the BANG project-team.
In
[BIOCHAM]for parameter search, can be used to
couple dynamical models. This approach is illustrated by a coupled model composed of:
a four phases model of the mammalian cell cycle by Novak and Tyson,
a circadian clock model by Leloup and Goldbeter,
a DNA damage repair model by Ciliberto et al.,
a model of irinotecan metabolism by Dimitrio and Ballesta,
a simple model of drug administration control.
This coupled model allows us to minimize the toxicity of irinotecan on healthy cells, using BIOCHAM's parameter search method applied on the drug administration control law.
In
One important perspective that is common to all this modeling work around the cell cycle is the promise of patient-tailored therapeutics
Modeling G protein-Coupled Signal Transduction
François
Fages
Steven
Gay
Domitille
Heitzler
Aurélien
Rizk
Sylvain
Soliman
In collaboration with Eric Reiter (UMR CNRS-INRA 6175) and Frédérique Clément (SISYPHE) in the framework of the Initiative Action
[REGATE], we analyse the
connectivity and dynamics of the FSH signalling network in its target cells, and embedding the network within a multi-scale representation, from the molecular up to the organic level.
We study the relative contributions of different pathways to the cell response to FSH signal, in order to determine how each pathway controls downstream cascades and which mechanisms are
involved in the transition between different cellular states.
A systems biology approach is followed using
[BIOCHAM]'s features for rule-based modeling with
temporal logic constraints used for searching unknown kinetic parameter values. In
![]() -lifting in functional languages. In conjunction with the equivalence between CHR and CSR with respect to naive operational semantics, these results lead to semantics-preserving
translations from full LCC to CHR and conversely. Immediate consequences of this work include new proofs for CHR linear logic and phase semantics, relying on corresponding results for LCC, plus
an encoding of the
-lifting in functional languages. In conjunction with the equivalence between CHR and CSR with respect to naive operational semantics, these results lead to semantics-preserving
translations from full LCC to CHR and conversely. Immediate consequences of this work include new proofs for CHR linear logic and phase semantics, relying on corresponding results for LCC, plus
an encoding of the
![]() -calculus in CHR.
-calculus in CHR.![]() )) are used to formalize the global behavior of the system known from biological experiments, and infer the unknown parameter values of the model.
)) are used to formalize the global behavior of the system known from biological experiments, and infer the unknown parameter values of the model.