Section: New Results
RNA
|
RNA design through random generation
Extensive experiments revealed a drift of existing software towards sequences with a high G+C -content. Relying on our random generation methods, we showed how to control this distributional bias in sequences using a multidimensional Boltzmann sampling [30] , [22] . We also explored the combination of random generation (global sampling) and local search into a novel category of glocal approaches, yielding promising results.
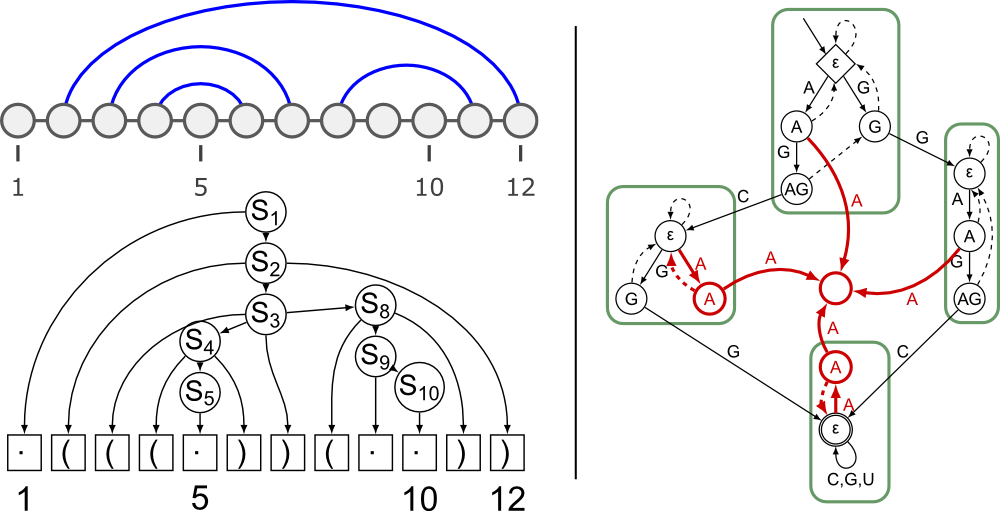
Finally, we explored language-theoretic constructs, namely products of finite-state automata and context-free languages, to force or forbid the presence of identified functional motifs within designed sequences [33] .
Towards 3D modeling of large molecules
Ab initio research benefited from our works on research and classification of RNA structural motifs [63] . Significant progress towards the ab initio prediction of the 3D structure of large RNAs were achieved. This problem is beyond the scope of current approaches and we proposed a promising coarse-grained approach based on game theory [13] that scales up to several hundreds of bases.
Fast-fourier transform for riboswitch
In the field of RNA computational biology, many algorithms use dynamic programming to partition the folding landscape according to a set of structural parameters. More precisely, the goal is to compute the number (resp. cumulated Boltzmann weight) of secondary structures having occurrences of some structural parameter , where may denote the distance to a reference structure, the number of # helices, base-pairs...The resulting algorithms, although polynomial in theory, are usually unusable in practice, particularly due to their unreasonable complexities (typically time/memory for parameters) and the intrinsic difficulties one encounters while trying to distribute their computation over multiple processors (highly connected dependency graph).
In collaboration with P. Clote's group (Boston College), we have described generic algorithmic principles to dramatically decrease these complexities, and make this class of algorithms practical. The main idea is to capture the partitioned space within a large polynomial, which can typically be efficiently evaluated (typically in ) as soon as the parameters are additive. One can then perform (possibly in parallel) independent evaluations of the polynomial, and use the Discrete Fourier Transform to recover the coefficients in time. Applying these principles to the RNAbor algorithm, whose complexities were in , we obtained an novel (parallelizable in time/memory on processors), we obtained a novel algorithm to detect bistable thermodynamic structures, such as riboswitches, which we presented at Recomb '13 [32] .