2023Activity reportProject-TeamCRONOS
RNSR: 202224368W- Research center Inria Centre at Université Côte d'Azur
- Team name: Computational modelling of brain dynamical networks
- Domain:Digital Health, Biology and Earth
- Theme:Computational Neuroscience and Medicine
Keywords
Computer Science and Digital Science
- A3.4. Machine learning and statistics
- A6.1. Methods in mathematical modeling
- A9.2. Machine learning
- A9.3. Signal analysis
Other Research Topics and Application Domains
- B1. Life sciences
- B1.2. Neuroscience and cognitive science
- B1.2.1. Understanding and simulation of the brain and the nervous system
- B1.2.2. Cognitive science
- B1.2.3. Computational neurosciences
- B2.2.2. Nervous system and endocrinology
- B2.2.6. Neurodegenerative diseases
- B2.5.1. Sensorimotor disabilities
- B2.6.1. Brain imaging
1 Team members, visitors, external collaborators
Research Scientists
- Théodore Papadopoulo [Team leader, INRIA, Senior Researcher, HDR]
- Rachid Deriche [INRIA, Emeritus, HDR]
- Samuel Deslauriers-Gauthier [INRIA, ISFP]
- Olivier Faugeras [INRIA, Emeritus, from Oct 2023, HDR]
- Romain Veltz [INRIA, Researcher, from Oct 2023]
PhD Students
- Yanis Aeschlimann [UNIV COTE D'AZUR]
- Igor Carrara [UNIV COTE D'AZUR]
- Petru Isan [CHU NICE, from Sep 2023]
Technical Staff
- Laura Gee [INRIA, Engineer, from Dec 2023]
- Eleonore Haupaix-Birgy [INRIA, Engineer, from Apr 2023]
Interns and Apprentices
- Martin Bouchet [INRIA, Intern, from Apr 2023 until Aug 2023]
- Arturo Cabrera Vazquez [UNIV COTE D'AZUR, Intern, from Mar 2023 until May 2023]
- Tijn Logtens [UNIV COTE D'AZUR, Intern, from Mar 2023 until May 2023]
Administrative Assistant
- Claire Senica [INRIA]
External Collaborator
- Petru Isan [CHU NICE, from Mar 2023 until Aug 2023]
2 Overall objectives
The objective of Cronos is to develop models, algorithms, and software to estimate, understand, and quantify the whole brain dynamics. We will achieve this objective by modeling the macroscopic architecture and connectivity of the brain at three complexity levels: source, sensor, and group (see Figure 1). These three levels, detailed in section 3, will be studied through the unifying representation of dynamic networks.
Cronos aims at pushing forward the state-of-the-art in computational brain imaging by developing computational tools to integrate the dynamic and partial information provided by each measurement modality (fMRI, dMRI, EEG, MEG, ...)1 into a consistent global dynamic network model.
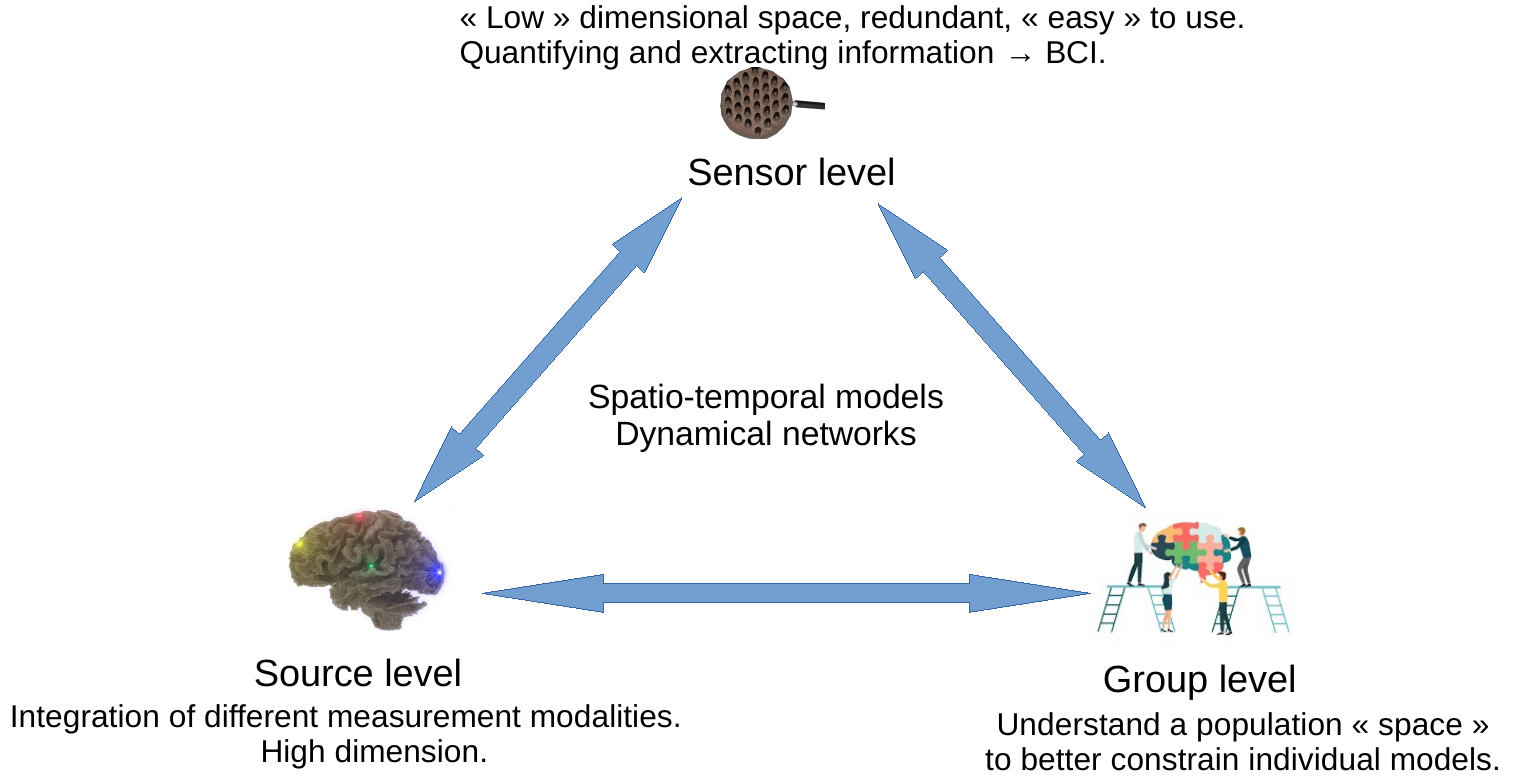
A graphic representation of the 3 Cronos axes (sensor, source and group) and their interactions.
Developing the computational models, algorithms, and software to estimate, understand, and quantify the brain's dynamical networks is a mathematical and computational challenge. Starting from several specific partial models of the brain, the overall goal is to assimilate in a single global numerical model the diversity of observations provided by non-invasive imaging. Indeed, the observations provided by various modalities are based on very different physical and biological principles. They thus reveal very different but complementary aspects of the brain, including its structural organization, electrical activity, and hemodynamic response. In addition, because of the varied nature of the physical principles used for imaging, different modalities also have drastically different spatial and temporal resolutions making their unification challenging. Some examples of sub-objectives we will develop are:
- Go beyond the hypothesis that brain networks are static over a given period of time.
- Study the possibility of estimating brain functional areas simultaneously with the networks.
- Study the impact on the reconstructed networks of the brain area to brain area activity transfer model.
- Develop a complete modeling of time delays involved by long range connections.
- Develop network based regularization methods in estimating brain activity.
- Go from subject specific studies to group level studies and increase the level of abstraction implied by the use of brain networks to open research perspectives in identifying network characteristics and invariances.
- Study the impact of the model imperfections on the obtained results.
Finally, the software implementing our models and algorithms must be accessible to non-technical users. They must therefore provide a level of abstraction, ease of use, and interpretability suitable for neuroscientists, cogniticians, and clinicians.
2.1 Main Research hypotheses
To achieve our goals, we decided to make some hypotheses on the way brain signals and networks are modeled.
2.1.1 Brain Signal Modeling
To model brain signals, we adopt a signal processing perspective, i.e. consider that at the macroscopic scale we look at signals, general signal modeling approaches are sufficient for the goal of describing brain dynamics. This contrasts with many other teams, which often follow a more constructivist approach, where brain signal models are derived from a computational neuroscience perspective, i.e. emerge from a mathematical modeling of neurons, axons and dendrites or assemblies of those (spiking neurons, neural mass models, ...). Our approach gives more freedom to model signals and allows for descriptions with a very limited number of parameters at the cost of losing some connection with the microscopic reality. Additionally, the reduced number of parameters simplifies the problem of parameter identification. Depending on the type of modeling, we may use rather simple phenomenological descriptions (e.g a region is activated or not) or more or less complicated signal models (sampled signals with no specific temporal models, various multivariate autoregressive models, integro-differential equations, ...). Generally speaking, the more we will be interested in dynamical properties of the signals, the more we will need sophisticated signal models.
2.1.2 Network Modeling
Classical – deterministic – networks only strictly exist at the microscopic level where neurons are connected together through axons. For complex neural systems, such networks are huge, currently unavailable and too complex for a macroscopic view of the brain. At such a scale, networks are of stochastic nature. Both nodes and edges are only defined probabilistically. We allow ourselves to work either directly with these probablistic networks or with deterministic networks obtained by thresholding the probabilistic ones, in which case it is important to pay attention to the amount of approximation and bias introduced by the thresholding operation. Its is important to note that, while deterministic networks have received a lot of attention, the domain of stochastic networks or its tranfer to computational neurosciences is still in its infancy.
3 Research program
As described in section 2 and Figure 1, the research program of Cronos is organized in 3 main axes corresponding to different levels of complexity:
- The Sensor level aims at extracting information directly from sensor data, without necessarily using the underlying brain anatomy. It can be thought of as a projection of the underlying brain networks onto a low dimensional space thus allowing computationally efficient processing and analysis. This provides a first window for observing and estimating brain dynamics. This sensor space is particularly convenient for the estimation of properties that are projection invariants (e.g. number of sources, state changes, ...). It is also the level that is most suitable for real time applications, such as brain computer interfaces.
- The Source level aims at measurement integration into a unified and high dimensional spatio-temporal space, i.e. a computational representation of dynamic brain networks. This is the level that is understood by neuroscientists, cogniticians, and clinicians, and therefore has an essential role to play in the visualization of brain activities.
- The Group level aims at providing tools to constrain the search space of the two previous levels. This is the natural space to develop statistical models and tests of the functioning of the brain. It is the level that allows the study of inter-subject variability of brain activity and the development of data driven priors. In particular, the topic of making statistics over noisy networks is a challenging task for this level.
These three levels closely interact with each other, as depicted by Fig. 1, towards the ultimate goal of non-invasively and continuously localizing brain activity in the form of brain networks.
3.1 Sensor level: the first window on brain dynamics
Sensor level mostly concerns MEG and EEG measurements. These serve as a support not only for experiments aimed at understanding the functioning of the brain when it performs certain tasks, but also for the characterization of certain pathologies (such as epilepsy). As an inexpensive modality, EEG is also widely used for the development of brain-computer interfaces (BCI). EEG and MEG are characterized by a high temporal resolution (up to 1000Hz or more), by a poor spatial resolution (measurable events involve several centimers-square portions of the cortex) and by a rather poor signal-to-noise ratio (SNR)2. For both EEG and MEG, the obtained measurements are linear mixtures of the “true” electrical sources that are at a distance of the sensors. MEG and EEG temporal resolution characteristics allow them to reveal changes in the dynamic of the brain activity. This “sensor space” has two main advantages: 1) its relatively small dimension (from a few to several hundreds sensors) compared to that of the “source space” (tens of thousands of degrees of freedom) and 2) its ease-of-use3. It is thus opportune to extract as much information as possible from this lower dimensional space, which already contains all the dynamical information of the functioning brain available from these modalities. This involves a better understanding of the invariants between source and sensor levels, which can lead to better Brain Computer Interfaces (BCI) classification algorithms. BCI not only clearly is an applicative domain that will benefit from this research, but will also help in validation of algorithms and methods. Our ambition is also to study how BCI-like techniques can be used for constraining the reconstruction of brain networks, for detecting some brain events in real time for clinical applications for creating improved cognitive protocols using real time responses to dynamically adapt stimulations.
This axis relies on three major ideas, which are necessary steps towards the ultimate goal of using MEG or EEG systems to non-invasively and continuously localize brain activity in the form of brain networks (i.e. without averaging multiple trials signals).
3.1.1 Automating the detection of brain state changes from the acquired data
Experimental practice in MEG, EEG, and BCI shows that very valuable information can be obtained directly from the “sensor space”. This encompasses estimating the number 43 or the time courses of sources 38, 37 or the detection of changes in the brain dynamical activity. This type of information can be an indication of a “brain state change”. In particular, automating the detection of changes in brains states would allow the splitting of the incoming functional data into segments in which the brain network can be considered as stationary. During such stationary segments of data, the sources and their locations remains fixed, which offers a means to regularize the solution of the inverse problem of source reconstruction. Furthermore, any improvement on EEG signal classification has direct applications for Brain Computer Interfaces. One approach that we will explore is the use of autoregressive spatio-temporal models or lagged correlation measurements as a means to improve classification of MEG/EEG signals. These autoregressive spatio-temporal models are a possible first step towards using more sophisticated causality modeling.
3.1.2 Better modeling of the spatio-temporal variability of brain signals
Hand in hand with the previous sub-goal, there is the need to better understand the spatio-temporal variability of brain signals. This variability originates from several factors ranging from the intrinsic variability of the brain sources (even in a same subject) to important differences in the spatial organization of the cortex across subjects or to variations in the way sensors are setup by experimenters. Thus, variability occurs not only across subjects, but also across sessions or even trials for a same subject.
The most common approach developed to cope with the noisy MEG and EEG signals is commonly referred to as evoked potentials: the signal is “clocked” on some stimulus or reaction of the subject and the low amplitude signals – almost completely hidden by background activity – are then averaged over multiple repetitions (trials) of the experiment to improve the SNR. But, because of variability, such an averaging distorts the overall activity and hides specific activities such as high frequency components. It is thus advisable to improve models so that they can work in single event mode (i.e without averaging). Relying on multiple trials to obtain better statistical models of individual signals (with techniques such as dictionary learning, autoregressive models, deep-neural networks, etc) would be an important improvement over the current state of the art as it would provide a more objective and systematic way of characterizing single trial data. Events will be extracted separately for each trial without relying on averaging, but with the knowledge of the “group” model. This will allow the study of the variability of brain activity across trials (attention, habituation are e.g. known to change activity). This is a difficult long term challenge with possible short-middle term advances for some specific cases such as epileptic spikes. This research path is grounded by some of our previous work 5, 2. Understanding variability is particularly important for Brain Computer Interfaces as it is often necessary to “learn” a classifier to detect the subject's brain state with a limited dataset (because of time constraint). At the sensor level, spatial patterns of activity are often described by covariance/correlation matrices. Using Riemannian metric over the space of symmetric definite positive matrices is a powerful technique that has been used these last years by the BCI community. Extending and improving these techniques as well as finding proper low dimensional spaces that characterize brain activity are other research paths we will follow.
3.1.3 Adapting to new sensor modalities
During the last few years, tremendous improvements have been made on means to acquire functional brain data. Yet, novelty in this domain continues and new sensors are regularly proposed. These sensors can offer more accurate measurements with improved signal-to-noise ratio (SNR) and/or some ease-of-use improvements. For example, the use of room temperature MEG sensors (such as those developed by Mag4Health) promise improvements in both aspects. EEG dry electrodes hold the promises to simplify the setup of BCI systems, but are difficult to master. MR machines are also more and more powerful with either higher fields (better signal quality and/or better spatial resolution) or, on the contrary, lower fields (easier to use and better contrasts in some cases). It is important to continually adapt processing methods to exploit the specificities of these new sensors as they become available to the community.
3.2 Source level: the integration space
Magnetic Resonance (MR) images of the brain offer different views of its organization and function via diffusion MRI tractography and functional MRI connectivity. When combined with MEG and EEG, we obtain complementary, but highly heterogeneous perspectives on brain networks and their dynamics. The natural space to integrate this complex information is the space of brain sources, allowing a unified model of imaging data. This axis is built over our past research (in the team Athena) in modeling the propagation of the electromagnetic field from brain sources to sensors 6, brain structural connectivity 3 or mapping different brain imaging modalities 4. Our plan for fusing the data originating from different modalities into a single network based model can be described in 3 sub-topics.
3.2.1 Source modeling: anatomical, temporal, and numerical constraints
Brain sources refer to regions of interests whose properties link brain activity to the observed measurements. For example, in EEG, brain sources can be modeled as dipoles representing the superposition of many synchronous and parallel neurons. The combination of the electric fields generated by these dipoles gives rise to the potential differences measured at the surface of the scalp in EEG. The magnetic fields generated by these same dipoles give rise to MEG signals. Regardless of the modality, the task of recovering brain sources from imaging data is an inverse problem. This inverse problem is ill–posed, either because of the limited number of sensors (EEG and MEG), because of poor signal to noise ratio (EEG, MEG, fMRI), or because of the limited spatial (dMRI, EEG, MEG) or temporal resolution (fMRI, dMRI). To recover a unique and stable solution, simple mathematical criteria (minimum norm, sparsity) are used to constrain the source space. However, these regularization approaches do not correspond to any specific anatomical or physiological properties of the brain and are therefore quite arbitrary. The challenge we address here is to increase the amount of subject specific anatomical and physiological constraints taken into acount for the recovery of brain activity from non–invasive imaging. More specifically, we will investigate how brain networks, and in particular their associated delays, can be used to constrain the inverse problem. Previous work has already shown the potential of temporal regularization based on brain connectivity 3, but the topic remains largely open to identifying new models grounded in physiological data. Another difficulty is the dynamic aspect of brain networks: we will first assume that a stationary brain state has been identified (e.g. using methods from section 3.1). Eventually (as a long term objective), transitions between stationary brain state models could be directly modeled in the networks themselves and estimated from the data. The validation of the proposed models is both challenging and essential and will be explored with our clinical partners (see section 9). Finally, non-traditional imaging modalities will also be considered. For example, accurate modeling of electromyography, in collaboration with Neurodec, can provide important timing information of sources.
3.2.2 Unified network models explaining various brain measurements
In the previous section, brain activity is estimated from one modality using information from the others as constraints or priors. This obviously favors one modality over the others and propagates errors of processing pipelines via the constraints. This modus operandi, while suboptimal, is justified historically: analysis pipelines have been developed independently for each modality by different communities. Given the complementary nature of the different modalities, it seems relevant to instead construct one global pipeline encompassing all brain measurement modalities into a single unified framework. With brain networks arising as a central concept in the neuroscientific community, we propose to make it the basis of such a global model. Doing so will enable the recovery of brain network dynamics from non–invasive multi–modal data, an important open problem in neuroscience. To achieve this objective, we propose to devise forward models describing the link between brain networks and each of the various imaging modalities. Specifically, work is needed to evaluate different formalisms describing brain networks that differ in the way nodes and their interactions are defined. For example, we will investigate the relationship between electrical (EEG/MEG), architectural (diffusion MRI) and metabolic (functional MRI) activities. Separated models – either electric or hemodynamic – have been devised for some time, but coupling them is an important challenge all the more that the role of some neural constituents (such as glia) is currently not well understood. Such models have already been – at least partly as in 35 – devised. The main challenge is to define one sufficiently simple and well spatialized (in particular to incorporate connectivity) in order to be useful for our purpose. We will also consider electromyography in this context, extending brain networks to the peripherial nervous system.
3.2.3 Global inverse problem using all measurements altogether
Models established in the previous section can be used in a global inverse problem involving all the sources of measurements to recover the specific brain network involved in a task. In all cases, we need to solve an inverse problem over a complex network model with a sparse prior as we need to find "simple" networks that can explain our data. Depending on the way networks are modeled, some difficulties may arise. For example, the model defined in 41, 40 leads to combinatorial explosion when used for source localization. Finding appropriate models that facilitate the resolution of the inverse problem may require some iterations between this sub-task and the previous one.
3.3 Group level: understanding variability to constrain network properties
Given the high intra- or inter-subject variability of brain activity, it is particularly interesting to be able to characterize the part of the activity that remains invariant (over time for one subject or across different subjects). This will allow a better understanding of brain processes as well as a better understanding of their variability across time or subjects. It also opens the possibility to constrain network models to reduce their complexity and improve their identifiability. The following description mostly refers to group statistics at the source level, but similar techniques are also relevant for the sensor level axis (see section 3.1). This research axis is a longer term research effort as it builds upon previous axes.
3.3.1 Matching individual subject models
Reconstructions obtained in section 3.2 will certainly differ even for different measurement sessions obtained for a same subject. An important problem to solve is the matching of instances of such reconstructions (whether they consist in brain networks or spatio-temporal autoregressive models). In general, the relative positions of functional brain areas are fairly well known. However, timing variations (lag, duration) in these models make the matching problem rather complicated. Finding and using some invariants of the models is one path to find a good compromise that would allow enough flexibility in the matching process while avoiding the combinatorial complexity that e.g. graph matching problems usually exhibit. Another – long term – possibility would be to directly solve “group problems”, but this would complexify even more the modeling problem and might have the same drawbacks as averaging if not done properly (i.e. hiding some activity or the intrinsic variability of the studied phenomenon).
3.3.2 Statistics over brain networks
Once the matching of single subject models has been solved, statistics will be necessary to assess the significance of the various model elements (e.g. in the case of dynamic network models, brain areas and their connections). Such statistics can also be used as a basis to derive “group statistical models” describing a family of task-related models that can in turn be used to constrain models used in section 3.2 and, through a forward model to reduce search space at sensor level 3.1. The BCI community is still divided on the actual benefits in terms of accuracy on going to source space for classifying brain signals. The statistical tools developed in this sub-axis may help to address this problem.
4 Application domains
4.1 Clinical applications
Cronos research has a strong clinical potential impact for brain diseases like epilepsy, brain cancer surgery, phantom pains, traumatic brain injuries, ...We work closely with several hospital teams and research groups (Pasteur hospital in Nice, La Timone hospital in Marseille, CRNL – Centre de Recherche en Neuroscience in Lyon –) towards exporting our research in their contexts. Example of applications are:
- Better understanding brain stimulation used in awake brain surgery and its relation with brain anatomy and in particular fibers as measured by dMRI.
- Real time detection of epileptic spikes from new generation MEG data and the visualization of their associated brain sources in real time.
- Use of BCI (see section 4.2) for helping disabled people to communicate (e.g. for patients suffering from Amyotrophic Lateral Sclerosis) or for helping epileptic patients to learn how to control the occurence of seizures.
- Understand the somatotopy of the thalamus and its relation with the efficiency of the deep brain stimulation therapy.
4.2 Brain Computer Interfaces
Brain Computer Interface is a closely related domain to both the `Signal Processing` and `Network Modeling` aspects. It typically extracts from the signal a 'brain state' that is a correlate of the `brain network'. While traditional MEG/EEG studies have been extensively exploited by the BCI community, the opposite nourishing of the former field by BCI has been much less explored. There is a continuing dispute in the BCI community on the advantages of going to source space or not (see e.g. 42). By studying the invariants between source and sensor space, we hope to not only provide clues on the above dispute but also to open the opportunity of using BCI-like techniques to ease the more traditional brain signal processing. In some sense, it is such invariants that BCI exploits in doing classification on sensor data. BCI also has the advantage of providing a more quantitative way (in term of classification quality) to evaluate methods. A strong BCI concern is also on the resources (computer power, number and quality of the sensors, ...) needed to achieve a given classification accuracy. This point of view is also complementary to that of more traditional brain signal modeling and can have a significant impact in terms of cost and ease of use in the clinical context.
5 Highlights of the year
5.1 Awards
- Rachid Deriche has been elected as a member of Academia Europaea in the section Informatics [May. 2023].
- Rachid Deriche has obtained the position of Inria Emeritus Research Director [Jan. 2023].
- The article 15 has been selected by the “Société des Neurosciences” to be in the "Highlights 2023" of the works of the community.
6 New software, platforms, open data
6.1 New software
6.1.1 BCI-VIZAPP
-
Name:
Real time EEG applications
-
Keywords:
Health, Brain-Computer Interface, GUI (Graphical User Interface)
-
Functional Description:
BCI-VIZAPP is a software suite for designing real-time EEG applications such as BCIs or neurofeedback applications. It has been been developed to build a virtual keyboard for typing text and a photodiode monitoring application for checking timing issues, but can now be also used in other tasks such as EEG monitoring. Originally, it was designed to delegate signal acquisition and processing to OpenViBE but has recently been extended to get some of these capabilities. This allows for more integrated and robust applications but also opens up new algorithmic opportunities, such as real time parameter modification, more controlled interfaces, ...
-
News of the Year:
Bci-Vizapp has been enriched with numerous features, notably to read and save certain files created with OpenViBE or with MNE-python, thus allowing an easier communication with them. Bci-Vizapp is also the software base used (or which we aim to use it in the long term) for different contracts (Demagus, EPIFEED, EPINFB, Techicopa and ConnectTC) and has integrated different elements to support them.
-
Contact:
Théodore Papadopoulo
-
Participants:
Nathanaël Foy, Romain Lacroix, Maureen Clerc Gallagher, Théodore Papadopoulo, Yang Ji, Come Le Breton
6.1.2 BifurcationKit
-
Name:
Automatic computation of numerical bifurcation diagrams
-
Keywords:
Bifurcation, GPU
-
Functional Description:
This Julia package aims at performing automatic bifurcation analysis of possibly large dimensional equations function of a real parameter by taking advantage of iterative methods, dense / sparse formulation and specific hardware (e.g. GPU).
It incorporates continuation algorithms (PALC, deflated continuation, ...) based on a Newton-Krylov method to correct the predictor step and a Matrix-Free/Dense/Sparse eigensolver is used to compute stability and bifurcation points.
The package can also seek for periodic orbits of Cauchy problems. It is by now one of the few software programs which provide shooting methods and methods based on finite differences or collocation to compute periodic orbits.
The current package focuses on large-scale, multi-hardware nonlinear problems and is able to use both Matrix and Matrix Free methods on GPU.
-
News of the Year:
The package was complemented with methods for computing curves of Neimark-Sacker or period-doubling bifurcations in two parameters. The corresponding normal forms have also been implemented. Additionally, a package HclinicBifurcationKit.jl has been added to the github organisation bifurcationkit. It allows computations of homoclinic orbits.
- URL:
- Publication:
-
Contact:
Romain Veltz
-
Participant:
Romain Veltz
6.2 Open data
Temporal Brain Networks Dataset
Participants: Samuel Deslauriers-Gauthier, Aurora Rossi [COATI, Inria], Emanuele Natale [COATI, Inria].
One of the first step in our study of null models for dynamic brain networks 16, 33, 24 was to process a significant amount of the data from the Human Connectome Project (HCP). The result of this work is the dynamic brain networks of 1047 subjects at various resolutions (number of nodes). Recognizing that few teams have the expertise and computational resources to reproduce this dataset, we decided to make it publicly accessible 32. It is our hope that this dataset will not only allow other researchers in network neuroscience to reproduce our results, but also allow researchers in other fields (e.g. graph theory) access to real world networks. This dataset is available in an easy-to-read format from entrepot.recherche.data.gouv.fr.
7 New results
7.1 Sensor Level: Brain Signal Modeling
Augmented Covariance Approaches for BCI
Participants: Igor Carrara, Théodore Papadopoulo.
EEG signals are complex and difficult to characterize, in particular because of their variability, whether in the same subject through the repetition of the same experiment or a fortiori when one wants to carry out multi-subject analyses. They therefore require the use of specific and adapted signal processing methods. In a recent approach 34, sources were modeled as an autoregressive model which explains a portion of a signal. This approach works at the level of the source space (i.e. the cortex), which requires modeling of the head and makes it quite expensive. However, EEG measurements can be considered as a linear mixture of sources and therefore it is possible to estimate an autoregressive model directly at the measurement level. The objectives of this line of work is to explore the possibility of exploiting EEG/MEG autoregressive models to extract as much information as possible without requiring the complex head modeling required for source reconstruction.
A first result in this line of research is the use of Augmented Covariance Matrices (ACMs) for BCI classification. These ACMs appear naturally in the Yule-Walker equations that were derived to recover autoregressive models from data. ACMs being symmetric positive definite matrices, it is natural to apply Riemannian Geometry based classification approaches on these objects, as those are the current state-of-the-art for such matrix types. Using these ACMs noticeably improves classification performance on several BCI benchmarks both in the within-session or in the cross-session evaluation protocols. This is due to the fact that ACMs incorporate not only spatial (the classical covariance matrix) but also temporal information. As such, it contains information on the nonlinear components of the signal through the embedding procedure, which allows the leveraging of dynamical systems algorithms. This comes at the cost of introducing two hyper-parameters: the order of the autoregressive model and a delay that controls the time resolution at which the signal is considered, which have to be estimated. This work is described in the preprint 18, which is currently under evaluation.
The improvements related above are made with a grid search hyper-parameter selection, which can be quite costly. As an ACM can be seen as the standard spatial covariance matrix of an extended signal created from the original one using a multidimensional embedding, there is an obvious link with the delay embedding theorem proposed by Takens 45 for the study of dynamical systems. This view provides an alternate path for estimating the hyper-parameters using methods proposed for dynamical systems such as the MDOP (Maximizing Derivatives On Projection) algorithm 44. This method is evaluated in the above preprint.
Finally, ACM approaches also allow better classification of "reduced signals" with typically only a few EEG channels 31.
Augmented methods with SPDNet
Participants: Igor Carrara, Bruno Aristimunha [Universidade Federal do ABC, Santo André, Brazil and Université Paris-Saclay, Paris, France], Marie-Constance Corsi [ARAMIS, Inria, Paris Brain Institute, France], Raphael Yokoingawa De Camargo [Universidade Federal do ABC, Santo André, Brazil], Sylvain Chevallier [Université Paris-Saclay, Paris, France], Théodore Papadopoulo.
This conference poster 29 studies whether the combination of phase space reconstruction and deep-learning methods can enhance the performance of BCI decoding for motor imagery (MI-BCI) in a low number of electrodes situation. This work relies on the combination of the Augmented Covariance method (ACM) explained in the previous paragraph, but implemented in the deep learning framework of SPDNet. The results show that the combination of ACM and SPDNet improves the classification performance of MI-BCI in a low number of electrodes regimes over the deep learning state-of-the-art. Moreover the Augmented SPDNet is explainable (using GadCam++ 36), with a low number of trainable parameters and a reduced carbon footprint.
In another work 28, we propose a contrastive learning approach to learn discriminative representations of EEG signals in the context of motor imagery (MI) BCI tasks. Specifically, we investigate the use of SPDNet with different EEG signal representations based on symmetric positive definite (SPD) matrix: covariance matrix, augmented covariance matrix, and coherence matrix. The main advantage of contrastive learning is that it can learn representations that are invariant to irrelevant variations, such as differences in electrode placement, noise, and individual differences in brain activity. By learning to contrast similar and dissimilar examples, contrastive learning can discover patterns that are shared across subjects and generalize well to new data. We validate our approach using the MOABB framework (Mother of All BCI Benchmarks), focusing on Cross Subject evaluation. The study is in its initial stages and further research is required to fully explore the potential of contrastive learning and SPDNet in the context of EEG-BCI.
Pseudo-online framework for BCI evaluation
Participants: Igor Carrara, Théodore Papadopoulo.
BCI (Brain-Computer Interface) technology operates in three modes: online, offline, and pseudo-online. In the online mode, real-time EEG data is constantly analyzed. In offline mode, the signal is acquired and processed afterwards. The pseudo-online mode processes collected data as if they were received in real-time, but without the real-time constraints. The main difference is that the offline mode often analyzes the whole data, while the online and pseudo-online modes only analyze data in short time windows, which is more flexible and realistic for real-life applications. Consequently, offline processing tends to be more accurate, but biases the comparison of classification methods towards methods requiring more dataset. In this work, we extended the current MOABB framework 39, that used to operate in offline mode only, with pseudo-online capabilities using overlapping sliding windows. This requires the introduction of an idle state event in the dataset to account for non task related signal portions. We analyzed the state-of-the-art algorithms of the last 15 years over several Motor Imagery (MI) datasets composed by several subjects, showing the differences between the two approaches (offline, and pseudo-online) and show that the adoption of the pseudo-online strategy modifies the hierarchy of classification methods. The publications related to this work are 30, 19. The second reference is a preprint of an accepted article that will appear in 2024.
Familiarity and mind wandering in neural tracking of music
Participants: Joan Belo, Maureen Clerc [Inria], Daniele Schön [ILCB, Aix-Marseille University].
One way to investigate the cortical tracking of continuous auditory stimuli is to use the stimulus reconstruction approach. However, the cognitive and behavioral factors impacting this cortical representation remain largely overlooked. Two possible candidates are familiarity with the stimulus and the ability to resist internal distractions. To explore the possible impacts of these two factors on the cortical representation of natural music stimuli, forty-one participants listened to monodic natural music stimuli while we recorded their neural activity. Using the stimulus reconstruction approach and linear mixed models, we found that familiarity positively impacted the reconstruction accuracy of music stimuli and that this effect of familiarity was modulated by mind wandering. This work is published in 10.
7.2 Source Level: Brain Dynamic Network Modeling
Null models of dynamic brain networks
Participants: Samuel Deslauriers-Gauthier, Aurora Rossi [COATI, Inria], Emanuele Natale [COATI, Inria].
Null models are crucial for determining the degree of significance when testing hypotheses about brain dynamics modeled as a temporal complex network. The comparison between the hypotheses being tested on empirical data and on the null model enables us to assess the extent to which an apparently remarkable feature of the former can be attributed to randomness. In 16, 33, 24, we consider networks generated from resting-state functional magnetic resonance imaging (fMRI) signals from the Human Connectome Project using standard processing techniques. The network nodes correspond to brain regions while the edges represent the presence of a correlation greater than a certain value between the signals of two regions. Over time, edges may appear and disappear, indicating that the associated correlation may jump above and below a certain value. While null models for static networks have been studied extensively, there is a lack of attention to temporal networks. Therefore, our investigation focuses on the study of temporal null models. In doing so, we focus on the ability of the null models to reproduce the temporal small-worldness present in the empirical data, a property associated with the efficient exchange of information at local and global scales over time.
The networks used in this work are publicly available 32.
Challenge on the estimation of structural connectivity from dMRI
Participants: Samuel Deslauriers-Gauthier, Rachid Deriche, and many other researchers from the dMRI community.
Estimating structural connectivity from diffusion-weighted magnetic resonance imaging is a challenging task, partly due to the presence of false-positive connections and the misestimation of connection weights. Building on previous efforts, the MICCAI-CDMRI Diffusion-Simulated Connectivity (DiSCo) challenge was carried out to evaluate state-of-the-art connectivity methods using novel large-scale numerical phantoms 13. The diffusion signal for the phantoms was obtained from Monte Carlo simulations. The results of the challenge suggest that methods selected by the 14 teams participating in the challenge can provide high correlations between estimated and ground-truth connectivity weights, in complex numerical environments. Additionally, the methods used by the participating teams were able to accurately identify the binary connectivity of the numerical dataset. However, specific false positive and false negative connections were consistently estimated across all methods. Although the challenge dataset does not capture the complexity of a real brain, it provided unique data with known macrostructure and microstructure ground-truth properties to facilitate the development of connectivity estimation methods.
Brain surgery: Free-water correction DTI-based tractography in brain
Participants: Fabien Almairac [Pasteur Hospital, UR2CA, Nice], Drew Parker [University of Pennyslvania], Lydiane Mondot [UR2CA, Nice], Petru Isan [Pasteur Hospital, UR2CA, Nice], Marie Onno [Pasteur Hospital, Nice], Théodore Papadopoulo, Patryk Filipiak [NYU Langone Health], Denys Fontaine [Pasteur Hospital, UR2CA, Nice], Ragini Verma [University of Pennyslvania].
Peritumoral edema prevents fiber tracking from diffusion tensor imaging (DTI). A free-water correction may overcome this drawback, as illustrated in the case of a patient undergoing awake surgery for brain metastasis. The anatomical plausibility and accuracy of tractography with and without free-water correction were assessed with functional mapping and axono-cortical evoked-potentials (ACEPs) as reference methods. The results suggest a potential synergy between corrected DTI-based tractography and ACEPs to reliably identify and preserve white matter tracts during brain tumor surgery. This work has been published in 9.
Brain surgery: identifying subcortical connectivity
Participants: Fabien Almairac [Pasteur Hospital, UR2CA, Nice], Petru Isan [Pasteur Hospital, UR2CA, Nice], Marie Onno [Pasteur Hospital, Nice], Théodore Papadopoulo, Lydiane Mondot [UR2CA, Nice], Stéphane Chanalet [UR2CA, Nice], Chalotte Fernandez [Pasteur Hospital, Nice], Maureen Clerc [Inria], Rachid Deriche, Denys Fontaine [Pasteur Hospital, UR2CA, Nice], Patryk Filipiak [NYU Langone Health].
Bipolar direct electrical stimulation (DES) of an awake patient is the reference technique for identifying brain structures to achieve maximal safe tumor resection. Unfortunately, DES cannot be performed in all cases. Alternative surgical tools are, therefore, needed to aid identification of subcortical connectivity during brain tumor removal. In this pilot study, we sought to (i) evaluate the combined use of evoked potential (EP) and tractography for identification of white matter (WM) tracts under the functional control of DES, and (ii) provide clues to the electrophysiological effects of bipolar stimulation on neural pathways. We included 12 patients (mean age of 38.4 years) who had had a dMRI-based tractography and a functional brain mapping under awake craniotomy for brain tumor removal. Electrophysiological recordings of subcortical evoked potentials (SCEPs) were acquired during bipolar low frequency (2 Hz) stimulation of the WM functional sites identified during brain mapping. SCEPs were successfully triggered in 11 out of 12 patients. The median length of the stimulated fibers was 43.24 ± 19.55 mm, belonging to tracts of median lengths of 89.84 ± 24.65 mm. The electrophysiological (delay, amplitude, and speed of propagation) and structural (number and lengths of streamlines, and mean fractional anisotropy) measures were correlated. In our experimental conditions, SCEPs were essentially limited to a subpart of the bundles, suggesting a selectivity of action of the DES on the brain networks. Correlations between functional, structural, and electrophysiological measures portend the combined use of EPs and tractography as a potential intraoperative tool to achieve maximum safe resection in brain tumor surgery. This work is published in 8.
Connectivity in the rat brain
Participants: Samuel Deslauriers-Gauthier, Joana Gomes-Ribeiro [Instituto de Medicina Molecular, Portugal], João Martins [Instituto de Ciências Nucleares Aplicadas à Saúde and University of Coimbra, Portual], Teresa Summavielle [Instituto de Investigação e Inovação em Saúde, Portugal], Joana E. Coelho [Instituto de Medicina Molecular, Portugal], Miguel Remondes [Instituto de Medicina Molecular, Portugal], Miguel Castelo Branco [Instituto de Ciências Nucleares Aplicadas à Sa'de and University of Coimbra, Portual], Luísa V. Lopes [Instituto de Medicina Molecular, Portugal].
Addiction to psychoactive substances is a maladaptive learned behavior. Contexts surrounding drug use integrate this aberrant mnemonic process and hold strong relapse triggering ability. In 21, we asked where context and salience might be concurrently represented in the brain, and found circuitry hubs specific to morphine-related contextual information storage. Starting with a classical rodent morphine-conditioned place preference (CPP) apparatus, we developed a CPP protocol that allows stimuli presentation inside a magnetic resonance imaging scanner, to allow the investigation of whole brain activity during retrieval of drug-context paired associations, as well as resting state functional connectivity under the effect of morphine conditioning. Using fMRI, we found context-specific responses to stimulus onset in multiple brain regions, namely limbic, sensory, and striatal. Furthermore, we found increased functional connectivity of lateral septum with regions within and beyond a proposed limbic network, and of the lateral habenula with hippocampal CA1 region, in response to repeated pairings of drug and context. Subsequent exposure to either morphine or saline-conditioned contexts led to significant, context-specific, functional interconnectivity among amygdala, lateral habenula, and lateral septum. Resting-state connectivity of the lateral habenula and amygdala, and that during saline-paired context presentation significantly predicted inter-individual CPP score differences. In sum, our findings show that drug- and saline-paired contexts form distinct memory traces in overlapping functional brain microcircuits, and intrinsic connectivity of habenula, septum, and amygdala likely influences the maladaptive contextual learning in response to opioids. We identify functional mechanisms involved in the acquisition and retrieval of drug-related memories that might be behind the relapse-triggering ability of opioid-associated sensory/contextual cues.
7.3 Group Level
Alignment of brain networks
Participants: Yanis Aeschlimann, Samuel Deslauriers-Gauthier, Théodore Papadopoulo, Anna Calissano [Inria EPIONE], Xavier Pennec [Inria EPIONE].
Every brain is unique, having its structural and functional organization shaped by both genetic and environmental factors over the course of its development. Brain image studies tend to produce results by averaging across a group of subjects, under a common assumption that it is possible to subdivide the cortex into homogeneous areas while maintaining a correspondence across subjects. This project questions such assumption: can the structural and functional properties of a specific region of an atlas be assumed to be the same across subjects? In 27, 11, this question is addressed by looking at the network representation of the brain, with nodes corresponding to brain regions and edges to their structural relationships. We perform graph matching on a set of control patients and on parcellations of different granularity to understand which is the connectivity misalignment between regions. The graph matching is unsupervised and reveals interesting insight on local misalignment of brain regions across subjects.
Structural connectivity is one view of brain networks provided by diffusion MRI, but these networks can also be observed from a functional point of view via functional MRI. In 26 and 17, we propose to simultaneously exploit structural and functional information in the alignment process. This allows us to explore multiple perspective of brain networks. Our results show the permutations induced by one type of connectivity (structural, functional) are not always supported by the other connectivity network, but when considering combined alignment, it is possible to find permutations of regions which are supported by both connectome modalities, leading to an increased similarity of functional and structural connectivity across subjects.
7.4 Other Results
Realistic simulation of Electromyogram (EMG) data
Participants: Samuel Deslauriers-Gauthier, Kostiantyn Maksymenko [Neurodec, Sophia Antipolis, France], Alexander Kenneth Clarke [Imperial College London, United Kingdom], Irene Mendez Guerra [Imperial College London, United Kingdom], Dario Farina [Imperial College London, United Kingdom].
In 14, we introduce the concept of a Myoelectric Digital Twin: a highly realistic and fast computational model tailored for the training of deep learning algorithms. It enables the simulation of arbitrary large and perfectly annotated datasets of realistic electromyographic signals, allowing new approaches to muscular signal decoding, accelerating the development of human-machine interfaces.
While the myoelectric digital twin is highly accurate, it is computationally expensive and thus is generally limited to modeling static systems such as isometrically contracting limbs. As a solution to this problem, we propose a transfer learning approach, in which a conditional generative model is trained to mimic the output of an advanced numerical model. To this end, we present BioMime in 22, a conditional generative neural network trained adversarially to generate motor unit activation potential waveforms under a wide variety of volume conductor parameters. We demonstrate the ability of such a model to predictively interpolate between a much smaller number of numerical model's outputs with a high accuracy. Consequently, the computational load is dramatically reduced, which allows the rapid simulation of EMG signals during truly dynamic and naturalistic movements.
Together, the myoelectric digital twin and BioMime provide a fast and accurate simulation of electromyography. However, the input to these systems is abstract: a percentage of voluntary contraction per muscle. A more intuitive and natural input would be a movement description, from which muscle contractions could be extracted. To this end, we propose in 23 NeuroMotion, an open-source simulator that provides a full-spectrum synthesis of EMG signals during voluntary movements. NeuroMotion is comprised of three modules. The first module is an upper-limb musculoskeletal model with OpenSim API to estimate the muscle fibre lengths and muscle activations during movements. The second module is BioMime, a deep neural network-based EMG generator that receives nonstationary physiological parameter inputs, such as muscle fibre lengths, and efficiently outputs motor unit action potentials (MUAPs). The third module is a motor unit pool model that transforms the muscle activations into discharge timings of motor units. The discharge timings are convolved with the output of BioMime to simulate EMG signals during the movement. NeuroMotion is the first full-spectrum EMG generative model to simulate human forearm electrophysiology during voluntary hand, wrist, and forearm movements.
Analysis of large size networks of Hopfield neurones
Participants: Olivier Faugeras, Etienne Tanré [LJAD, Nice, France].
We revisit the problem of characterizing the thermodynamic limit of a fully connected network of Hopfield-like neurons. Our contributions are that we provide a) a complete description of the mean-field equations as a set of stochastic differential equations depending on a mean and covariance functions, b) a provably convergent method for estimating these functions, and c) numerical results of this estimation as well as examples of the resulting dynamics. The mathematical tools are the theory of Large Deviations, Itô stochastic calculus, and the theory of Volterra equations.
An abstract has been submitted to the 2024 International Conference on Mathematical Neuroscience, deposited onto HAL and a manuscript is almost ready for submission to a Journal.
Pros and cons of mean field representations of large size networks of spiking neurons
Participants: Olivier Faugeras, Romain Veltz.
We have laid out a roadmap for classifying the behaviors of large ensembles of networks of spiking neurons through the analysis of the corresponding Nonlinear Fokker-Planck equation. Rather than using this equation to simulate the network equations, we apply advanced methods for analyzing the bifurcations of its solutions. This analysis can then be used to predict the behaviors of the original network. We have used the example of a fully connected network of N Fitzhugh-Nagumo neurons with electrical and chemical synapses to convey the interest of our approach.
An abstract has been submitted to the 2024 International Conference on Mathematical Neuroscience and a manuscript is almost ready for submission to a Journal.
Modeling of pain processing in spinal cord
Participants: Romain Veltz, Emmanuel Deval [IPMC].
In a recent publication 12, we finalized a work concerning the modeling of processing of pain information by deep projection neurons and more specifically the implication of ASIC1a channels. This involved experiments in rats with single neuron modeling done at the Institute of Molecular and Cellular Pharmacology (IPMC).
Modeling of excitatory synapse in rat hippocampus
Participants: Romain Veltz, Hélène Marie [IPMC].
In the publication 15, a stochastic model of hippocampal and excitatory synaptic plasticity has been finalized. It is the current state of the art model for such synapse. More precisely, we provide a multi-timescale model of hippocampal synaptic plasticity induction that includes electrical dynamics, calcium, CaMKII and calcineurin, and accurate representation of intrinsic noise sources. Using a single set of model parameters, we demonstrate the robustness of this plasticity rule by reproducing nine published ex vivo experiments covering various spike-timing and frequency-dependent plasticity induction protocols, animal ages, and experimental conditions. This is an on-going collaboration with H. Marie.
8 Bilateral contracts and grants with industry
8.1 Bilateral Grants with Industry
Demagus: BPI Grant April 2022–September 2024
Participants: Laura Gee, Éléonore Haupaix-Birgy, Côme Le Breton, Théodore Papadopoulo, Julien Wintz.
This grant has been obtained with the startup Mag4Health (other partners are CNRS and INSERM). This company develops a new MEG machine working with optically pumped magnetometers (magnetic sensors), which potentially means lower costs and better measurements. Cronos is in charge of developing a real time interface for signal visualization, source reconstruction and epileptic spikes detection. The initial funding was for 2 years, but an extension of 6 months has been obtained.
9 Partnerships and cooperations
9.1 International initiatives
9.1.1 Inria associate team not involved in an IIL or an international program
MUSCULAR
Participants: Samuel Deslauriers-Gauthier, Théodore Papadopoulo, Yanis Aeschlimann, Igor Carrara, Dario Farina [Imperial College London, United Kingdom], Shihan Ma [Imperial College London, United Kingdom], Bruno Grandi Sgambato [Imperial College London, United Kingdom].
Surface electromyography (EMG) is the non-invasive recording of electrical activity produced by skeletal muscle contractions. Surface electroencephalography (EEG) is the recording of electrical activity produced by the activity of neural assemblies in the brain. In both modalities, there is a strong interest in recovering the underlying physiological sources, either the motor unit spike trains or the activity of cortical neural assemblies, that explain the recordings. In addition, the underlying physics of EMG and EEG which relates sources to measurements are identical and governed by Maxwell’s equations. Despite their obvious similarities, the methodological landscapes of EMG and EEG have very little intersection and there is little bandwidth between the two scientific communities.
Recognizing the need for an increase in exchange and collaboration between the EEG and EMG communities, we proposed the associated team MUSCULAR between Cronos and Dario Farina's lab at Imperial College. The nature and objective of the MUSCULAR associated team is to increase the bandwidth between the EEG and EMG communities and promote a mutually beneficial exchange of methodology and knowledge. The Cronos team has a longstanding history in forward modeling and inverse problem in electroencephalography and magnetoencephalography. The Imperial College London team has extensive expertise in the processing and analysis of EMG signals for neurorehabilitation and the control of movements. By collaborating via the MUSCULAR associated team, the two research labs will rapidly move forward both the EMG and the EEG communities.
Because of our early work on the hyper-realistic simulation of EMG, published in Nature Communications in March 2023 14, the associated team was able to move quickly and has proposed two new methods to model EMG signals. The first contribution is the development of BioMime,conditional generative neural network trained adversarially to generate motor unit activation potential waveforms 22. The second contribution is the development of NeuroMotion, an open-source simulator that provides a full-spectrum synthesis of EMG signals 23. These contributions are further detailed in section 7.4.
9.2 European initiatives
9.2.1 H2020 projects
HBP SGA3
Participants: Romain Veltz.
-
Coordinator Name :
Victor Jirsa
-
Partner Institutions:
117 partner institutions in 19 countries across Europe
-
Date/Duration:
ended in September 2023
- Web site
Within the SGA3 component of HBP, we develop mean field models of stochastic and spiking neural networks with heterogenous connections.
9.3 National initiatives
ANR ChaMaNe, Mathematical Challenges in Neurosciences
Participants: Romain Veltz.
- Coordinator Name :
-
Partner Institutions:
- Partner 1: Biologie Computationnelle et Quantitative (CQB), France
- Partner 2: LJAD, Université Nice, France
-
Date/Duration:
2020-2025
- Web site
This project aims at a mathematical study, on the one hand, of the intrinsic dynamics of a neuron and their consequences, and on the other hand, of the qualitative dynamics of large neural networks with respect to the intrinsic behavior of the individual neurons, the interactions between them, memory effects, spatial structure, etc.
FHU InnovPain 2
Participants: Théodore Papadopoulo.
-
Coordinator Name :
Denys Fontaine (Pasteur Hospital)
-
Date/Duration:
2023–2027
- Web site
The FHU INOVPAIN is a Hospital-University Federation focused on chronic refractory pain and innovative therapeutic solutions. It involves 6 hospital facilities, 13 academic research teams and 12 platforms and core facilities.
9.4 Regional initiatives
Project NeuroKoopman - AAP neuromod GP
Participants: Samuel Deslauriers-Gauthier, Théodore Papadopoulo, Romain Veltz.
-
Coordinator Name :
Patricia Reynaud-Bouret (UniCA).
-
Partner Institutions:
- Partner 1: Neuromod
- Partner 2: IIT of Genova
- Partner 3: Ecole Polytechnique Paris
-
Date/Duration:
2024
This project aims at exploring Koopman's theory for characterizing dynamical system in the context of brain data. This grant is obtained to get first results and use them for applying to a bigger one (ANR or European grant).
Cooperation with clinical partners
The team Cronos also has some long-term cooperations with several clinical partners:
-
Awake brain surgery:
-
Institution:
Pasteur Hospital in Nice
-
Collaborators:
F. Almairac, P. Isan
-
Study name:
ConnectTC
-
Institution:
-
Using dry electrodes for the P300 speller BCI:
-
Institution:
Pasteur Hospital in Nice
-
Collaborators:
M.-H. Soriani, M. Bruno, E. Guichard
-
Study name:
TechIcopa
-
Institution:
-
Neurorehabilitation for epileptic patients:
-
Institution:
La Timone Hospital in Marseille
-
Collaborators:
F. Bartolomei, C. Bénar, R. Guex, ...
-
Study names:
EPIFEED and NFBEPI
-
Institution:
10 Dissemination
Participants: Rachid Deriche, Samuel Deslauriers-Gauthier, Olivier Faugeras, Théodore Papadopoulo, Romain Veltz.
10.1 Promoting scientific activities
10.1.1 Scientific events: organisation
Member of the conference program committees
- O. Faugeras is a member of the Scientific Committee of the International Conference on Mathematical Neuroscience, Dublin, June 2024
- O. Faugeras is a member of the Scientific Committee Random processes in the brain: From experimental data to Math and back, IHP, Paris, February-April
Reviewer
- T. Papadopoulo served several international and national conferences (ISBI, GRETSI, CORTICO days)4.
- S. Deslauriers-Gauthier served several international and national conferences (ISBI, IABM)4 and an international workshop (MICCAI-CDMRI)4.
- R. Deriche served several international conferences (ISBI, MICCAI)4 and an international workshop (MICCAI-CDMRI)4.
10.1.2 Journal
Member of the editorial boards
- T. Papadopoulo serves as Associate Editor in “Frontiers: Brain Imaging Methods” and as Review Editor for “Frontiers: Artificial Intelligence in Radiology”.
- R. Veltz is in the Editorial Board of the journal Mathematical Neuroscience and Applications.
- S. Deslauriers-Gauthier serves as Guest Associate Editor in Frontiers: Brain Imaging Methods and as Review Editor for Frontiers: Frontiers in Computational Neuroscience.
- R. Deriche is member of the Editorial Boards of the Journal of Neural Engineering and of the Medical Image Analysis Journal. He is also an editorial board member at Springer for the book series entitled Computational Imaging and Vision.
- O. Faugeras is the Editor in Chief of the journal Mathematical Neuroscience and Applications.
Reviewer - reviewing activities
- R. Deriche serves several international journals (NeuroImage, IEEE Transactions on Medical Imaging, Magnetic Resonance in Medicine, Medical Image Analysis Journal, Journal of Neural Engineering, ...).
- S. Deslauriers-Gauthier served several international journals (Medical Image Analysis, Nature Machine Intelligence, Network Neuroscience).
- O. Faugeras does reviews for several international Journals including Science, SIAM Journal on Applied Mathematics, SIAM Journal on Applied Dynamical Systems, Journal of Mathematical Biology, Journal of Mathematical Neuroscience and Applications.
- T. Papadopoulo served several international journals (Journal of Neural Engineering, Physics & Medicine in Biology, Inverse Problems, IEEE Transactions on Biomedical Engineering, Frontiers in Neurosciences, Frontiers in Human Neurosciences, Brain Topography, Computers in Biology and Medicine, IEEE Transactions on Medical Imaging).
- R. Veltz served several international journals (Chaos, Theory in Biosciences, SIAM for Mathematical Analysis, Physica D, Nonlinear Dynamics, Kyungpook Mathematical Journal).
10.1.3 Invited talks
- R. Deriche, From Computational Imaging to Brain Connectomics: A Geometrical Perspective, keynote speech at the Workshop of Informatics Section at Academia Europaea - Munich - Germany - Oct 2023.
- O. Faugeras, Two examples of thermodynamic limits in neuroscience, One World Dynamics seminar, Dec 5th, 2023.
- O. Faugeras, Mathematical Neuroscience: A personal perspective, Structural learning by the Brain, IHP workshop.
- O. Faugeras, Des mathématiques au chevet des neurones et des astrocytes, Semaine du cerveau, Paris.
- O. Faugeras, Two examples of thermodynamic limits in neuroscience, A Random Walk in the Land of Stochastic Analysis and Numerical Probability, CIRM.
- T. Papadopoulo, Improving BCI classification accuracy using tools from autoregressive models, ISAE-SUPAERO.
- T. Papadopoulo and I. Carrara, Toward everyday BCI: Augmented Covariance Method in a reduced dataset setting, 10th International BCI Meeting.
- R. Veltz, Some results concerning a mean field of a network of 2d spiking neurons, Oxford.
- R. Veltz, Theoretical / numerical study of modulated traveling pulses in inhibition stabilized networks, NFW Paris.
- R. Veltz, Study of a mean field of a network of 2d spiking neurons. One world dynamics seminar.
- R. Veltz, Theoretical study of a mean-field model of two-dimensional nonlinear integrate-and-fire neurons, ANR Chamane.
- R. Veltz, Theoretical / numerical study of modulated traveling pulses in inhibition stabilized networks, Workshop: Networks of spiking neurons, IHP Paris.
10.1.4 Leadership within the scientific community
- O. Faugeras is a member of the French Academy of Sciences and of the French Academy of Technologies.
- R. Veltz and J.B. Caillau, organization of a half day conference Julia at Inria, 50 attendees, January 2023.
10.1.5 Scientific expertise
- R. Deriche served several national and international institutions in reviewing applications : 3IA UniCA Chairs, ANR and ERC Grants.
- T. Papadopoulo served in reviewing applications for the Neuromod institute of Université Côte d'Azur.
- T. Papadopoulo served in reviewing for several internal programs of Université Côte d'Azur (see membership in academic council below).
- T. Papadopoulo served as an internal proof-reader for a PEPR proposal.
- T. Papadopoulo served in reviewing a proposal for an INRIA associate team.
- R. Veltz served in reviewing for the PhD grant program in Neuromod.
10.1.6 Research administration
- T. Papadopoulo is elected member in the academic council of University of Côte d'Azur (UniCA).
- T. Papadopoulo is the head of the Technological Development Committee of Inria Sophia Antipolis Méditerranée.
- T. Papadopoulo is a member of the Neuromod scientific council and represents Neuromod in the EUR Healthy (EUR: Universitary Research School) and in the Maison de la Simulation of UniCA.
10.2 Teaching - Supervision - Juries
10.2.1 Teaching
- Master: T. Papadopoulo, Inverse problems for brain functional imaging, 24 ETD, M2, Mathématiques, Vision et Apprentissage, ENS Cachan, France.
- Master: T. Papadopoulo and S. Deslauriers-Gauthier Functional Brain Imaging, each 15 ETD, M1,M2 in the MSc Mod4NeuCog of Université Côte d'Azur.
- Master: T. Papadopoulo and S. Deslauriers-Gauthier, Application of machine learning to MRI, electophysiology & brain computer interfaces, each 10 ETD, M1,M2 in the MSc Data Science and Artificial Intelligence of Université Côte d'Azur.
- PSL Graduate Program: T. Papadopoulo, Brain-Computer interfaces, 3 ETD, Advanced Course in Computational Neuroscience, Ecole Normale Supérieure, Paris, France.
- Master: R. Veltz, Mathematical methods for neuroscience, 24 ETD, M2, Mathématiques, Vision et Apprentissage, ENS Cachan, France.
- Automn School, EITN Paris, France. R. Veltz has been invited for a lecture on mean fields of neural networks.
10.2.2 Supervision
- PhD in progress: I. Carrara, "Auto-regressive models for MEG/EEG processing", started in Oct. 2021. Supervisor: Théodore Papadopoulo.
- PhD in progress: Y. Aeschlimann, "Brain networks from simultaneous modelling of functional MRI and diffusion MRI", started in Oct. 2022. Supervisors: Samuel Deslauriers-Gauthier, Théodore Papadopoulo.
- PhD in progress: P. Isan, "Cerebral eletrophysiological exploitation of in vivo evoked potentials to guide the resection of adult brain tumors", started in Oct. 2023. Supervisor: F. Almairac. Co-Supervisors: T. Papadopoulo, S. Deslauriers-Gauthier.
- Postdoc ended in December 2023: N. Vattuone "Study subject with the title Dynamics of spiking neural networks with connectivity represented by random graphs". Student co-supervised by R. Veltz and E. Tanré (Inria) since December 2021.
- Postdoc: O. Senkevich "Study subject with the title Fluctuations of synaptic connections’ strengths due to the randomness in synaptic protein counts caused by the stochasticity of gene expression and protein transport in neurons.", Student co-supervised by R. Veltz and C. O’Donnell (University of Ulster) since June 2022.
10.2.3 Juries
- R. Deriche participated in the Ph.D. jury of T. Durantel at University of Rennes on Dec. 15th, 2023.
- T. Papadopoulo participated in the Ph.D. jury of X. XU at University of Toulouse on Jan. 27th, 2023 as a reviewer.
- T. Papadopoulo participated in the Ph.D. jury of V. d. R. Hernández Catañón at Université of Lorraine on Feb. 8th, 2023.
- T. Papadopoulo participated in the Ph.D. thesis committees of G. Girier and V. Strizhkova for Université Côte d'Azur.
- T. Papadopoulo and S. Deslauriers-Gauthier participated in the Msc Neuromod first year training jury, 2023.
- R. Veltz participated in the jury of N. Vadot's master 2 defense at EPFL on June 2023. N. Vadot was supervised by Wulfram Gerstner.
- R. Veltz participated in the Ph.D. jury of S. Ebert at Université of Cote d'Azur on Dec. 13th, 2023 as an examiner.
10.3 Popularization
10.3.1 Articles and contents
- T. Papadopoulo has been interviewed for an article published in the magazine Science&Vie.
10.3.2 Interventions
- R. Veltz participated at "Semaine du numérique" at Terra Numerica in May 2023.
- R. Veltz participated at "Robotique MathC2+" in June 2023.
11 Scientific production
11.1 Major publications
- 1 articleStructural connectivity to reconstruct brain activation and effective connectivity between brain regions.Journal of Neural Engineering173June 2020, 035006HALDOI
- 2 articleConsensus Matching Pursuit for multi-trial EEG signals.Journal of Neuroscience Methods1801May 2009, 161-170HALDOIback to text
- 3 articleWhite Matter Information Flow Mapping from Diffusion MRI and EEG.NeuroImageJuly 2019HALDOIback to textback to text
- 4 articleA Unified Framework for Multimodal Structure-function Mapping Based on Eigenmodes.Medical Image AnalysisAugust 2020, 22HALDOIback to text
- 5 articleAdaptive Waveform Learning: A Framework for Modeling Variability in Neurophysiological Signals.IEEE Transactions on Signal Processing65August 2017, 4324 - 4338HALDOIback to text
- 6 articleA common formalism for the Integral formulations of the forward EEG problem.IEEE Transactions on Medical Imaging241January 2005, 12-28HALDOIback to text
- 7 articleA Myoelectric Digital Twin for Fast and Realistic Modelling in Deep Learning.Nature Communications141March 2023, 1600HALDOI
11.2 Publications of the year
International journals
International peer-reviewed conferences
Reports & preprints
Other scientific publications
11.3 Cited publications
- 34 articleStructural connectivity to reconstruct brain activation and effective connectivity between brain regions.Journal of Neural Engineering173June 2020, 035006HALDOIback to text
- 35 articleRelationship Between Flow and Metabolism in BOLD Signals: Insights from Biophysical Models.Brain Topography2412011, 40--53URL: http://dx.doi.org/10.1007/s10548-010-0166-6DOIback to text
- 36 articleGrad-CAM++: Generalized Gradient-Based Visual Explanations for Deep Convolutional Networks.2018 IEEE Winter Conference on Applications of Computer Vision (WACV)2017, 839-847URL: https://api.semanticscholar.org/CorpusID:13678776back to text
- 37 inproceedingsMultivariate Convolutional Sparse Coding for Electromagnetic Brain Signals.Advances in Neural Information Processing Systems31Curran Associates, Inc.2018, URL: https://proceedings.neurips.cc/paper/2018/file/64f1f27bf1b4ec22924fd0acb550c235-Paper.pdfback to text
- 38 articleAdaptive Waveform Learning: A Framework for Modeling Variability in Neurophysiological Signals.IEEE Transactions on Signal Processing6516April 2017, 4324--4338HALDOIback to text
- 39 articleMOABB: trustworthy algorithm benchmarking for BCIs.Journal of neural engineering1562018, 066011back to text
- 40 inproceedingsConnectivity-informed M/EEG inverse problem.GRAIL 2020 - MICCAI Workshop on GRaphs in biomedicAl Image anaLysisLima, PeruOctober 2020HALback to text
- 41 inproceedingsIncorporating transmission delays supported by diffusion MRI in MEG source reconstruction.ISBI 2021 - IEEE International Symposium on Biomedical ImagingNice, FranceApril 2021HALback to text
- 42 articleAs above, so below? Towards understanding inverse models in BCI.Journal of Neural Engineering151December 2017back to text
- 43 articleSource localization using recursively applied and projected (RAP) MUSIC.IEEE Transactions on Signal Processing472February 1988, 332 -- 340DOIback to text
- 44 articleOptimal state-space reconstruction using derivatives on projected manifold.Physical Review E8722013, 022905back to text
- 45 incollectionDetecting strange attractors in turbulence.Dynamical systems and turbulence, Warwick 1980Springer1981, 366–381back to text
