2023Activity reportProject-TeamMICROCOSME
RNSR: 202124114Z- Research center Inria Centre at Université Grenoble Alpes
- In partnership with:Université de Grenoble Alpes
- Team name: Analysis, engineering, and control of microorganisms
- Domain:Digital Health, Biology and Earth
- Theme:Modeling and Control for Life Sciences
Keywords
Computer Science and Digital Science
- A3.1.1. Modeling, representation
- A3.4.5. Bayesian methods
- A6.1.1. Continuous Modeling (PDE, ODE)
- A6.1.2. Stochastic Modeling
- A6.2.1. Numerical analysis of PDE and ODE
- A6.2.4. Statistical methods
- A6.3.1. Inverse problems
- A6.3.2. Data assimilation
- A6.3.3. Data processing
- A6.4.1. Deterministic control
Other Research Topics and Application Domains
- B1. Life sciences
- B1.1.2. Molecular and cellular biology
- B1.1.4. Genetics and genomics
- B1.1.7. Bioinformatics
- B1.1.8. Mathematical biology
- B1.1.10. Systems and synthetic biology
- B2.2.4. Infectious diseases, Virology
- B4.3.1. Biofuels
1 Team members, visitors, external collaborators
Research Scientists
- Delphine Ropers [Team leader, INRIA, Senior Researcher, Professeur attaché à l'UGA, HDR]
- Eugenio Cinquemani [INRIA, Senior Researcher, HDR]
- Aline Marguet [INRIA, Researcher]
- Hidde de Jong [INRIA, Senior Researcher, HDR]
Faculty Member
- Johannes Geiselmann [UGA, Professor]
Post-Doctoral Fellows
- Arnaud Belcour [INRIA, Post-Doctoral Fellow, from Sep 2023]
- Claudia Fonte Sanchez [UGA, Post-Doctoral Fellow, from May 2023]
PhD Students
- Rand Asswad [INRIA]
- Ignacia Cancino Aguirre [INRIA]
- Thibault Clavier [UGA]
- Charles Medous [UGA]
- Emrys Reginato [INRIA]
- Maaike Sangster [INRIA, until Feb 2023]
Technical Staff
- Soraya Arias [INRIA, Engineer]
- Ludovic Leau-Mercier [INRIA, Engineer]
Interns and Apprentices
- Eugene Ferragu [INRIA, Intern, from Oct 2023]
- Gwendal Le Guennec [ENS DE LYON, Intern, from Jun 2023 until Jul 2023]
Administrative Assistant
- Diane Courtiol [INRIA]
Visiting Scientist
- Antrea Pavlou [UNIV ZURICH]
External Collaborators
- Muriel Cocaign-Bousquet [INRAE, Toulouse Biotechnology Institute, HDR]
- Thibault Etienne [Gencovery, until Mar 2023]
2 Overall objectives
MICROCOSME combines computational and experimental approaches for the analysis, engineering, and control of the growth of microorganisms. Understanding and controlling the dynamics of bacterial growth is vitally important in health, medicine, biotechnology, and food industries, for instance to halt the growth of pathogens or stimulate the growth of probiotics or industrial microorganisms.
We develop multiscale models of growth, where the macroscopic observable, growth of a microbial population or community, depends on various metabolic pathways and regulatory mechanisms operating at microscopic scales within the cells. We use our (deterministic or stochastic) models to interpret experimental data or to infer the underlying growth processes from the data. This requires developing a platform for the automation of experiments, as well as methods and software for model estimation and data analysis. The analysis of microbial growth calls for new methodologies at the interface of microbiology, control theory, applied mathematics, computer science, biophysics, and molecular biology, which also leads to contributions in all of these fields. Our workhorse for the realization of this research program is the bacterium Escherichia coli pictured in Figure 1. Part of the microbiota of the human gut, E. coli is the model organism par excellence in microbiology and a popular platform for bio-based chemical production. We intend to extend approaches developed in-house for this specific microbe to other microorganisms including pathogens.
MICROCOSME has been created on October 1st, 2021. A recomposition and follow-up of former IBIS project-team, MICROCOSME unites researchers from Inria and the Laboratoire Interdisciplinaire de Physique (CNRS UMR 5588) at Université Grenoble Alpes.
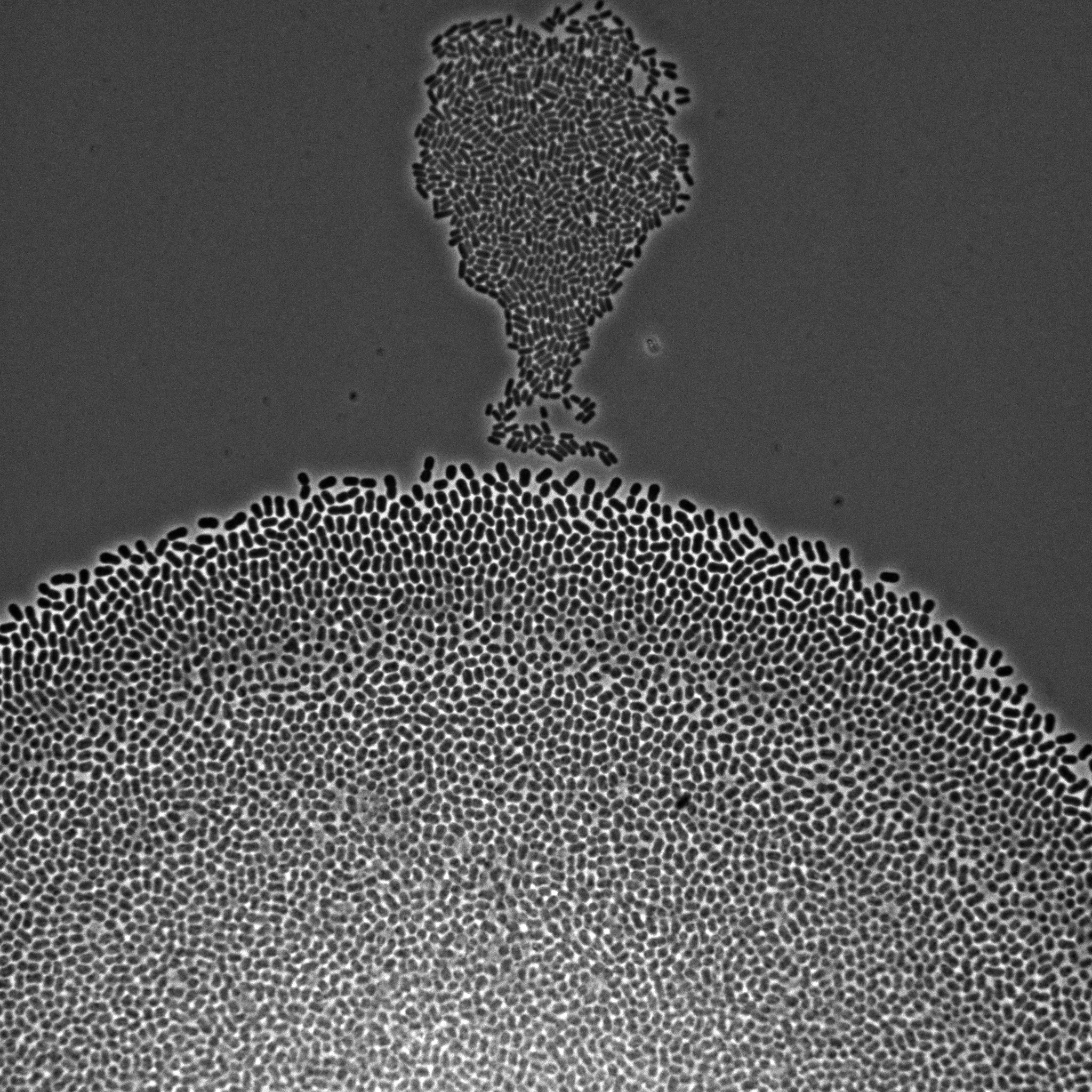
Figure 1: Microscopy image of Escherichia coli bacteria growing on a solid nutrient medium.
3 Research program
The research program of MICROCOSME is articulated around four research axes combining theory and experiments, which are illustrated in Figure 2 and detailed below.
3.1 Genome-scale analysis of microbial physiology
The molecular foundations of bacterial growth remain little understood today, because they involve large biochemical networks with physical and regulatory interactions across different levels of cellular organization. We investigate at the genome scale how the dynamics of gene expression and metabolism leads to microbial growth, using a combination of mathematical models and high-throughput data. The challenge is to integrate, in models of thousands of equations, multiple and heterogeneous datasets on the metabolic, transcriptomic, and proteomic level. We typically use constraint-based models to investigate the relations between microbial growth and metabolism, while the effect of growth on mRNA stability is analysed by means of non-linear mixed-effect models.
3.2 Natural and engineered resource allocation strategies in microorganisms
Microorganisms have evolved strategies to allocate their resources to different cellular functions and thus adjust their growth rate to fluctuating environments. We study these natural resource allocation strategies, by viewing cells as self-replicators that can be described using coarse-grained models and analysed by means of optimal and feedback control theory. The models take the form of systems of 5-10 nonlinear ordinary differential equations, with parameters estimated from published data or data obtained from dedicated experiments. Experimental work in the lab allows to validate model predictions on the single-cell and population level and to engineer new strategies for the reallocation of cell resources from growth to bioproduction.
3.3 Variability and robustness of microbial adaptation
The development of experimental techniques and the use of video-microscopy have led to a growing number of high-quality data showing the heterogeneity among cells in a population. We combine these single-cell data with models describing the stochastic dynamics of individual cells, such as birth-death processes, branching processes, and mixed-effect models. The models allow to investigate the origins of heterogeneity and its role in the adaptation of microorganisms to environmental changes, and to leverage population heterogeneity for biotechnological applications. In practice, this requires the extension of modelling approaches by taking into account the specificities of heterogeneity, as well as the development of appropriate methods and software for the inference of models and of biological quantities from quantitative time-course profiles of the microbial response to environmental changes.
3.4 Analysis and control of microbial communities
Heterogeneity also arises within communities consisting of different microbial species. Understanding microbial interactions is a challenging task that goes well beyond the characterization of single species, and offers great opportunities for applications, such as the control of the community for bioproduction. Indeed, suitably constructed microbial consortia carry the potential to outperform single species in the accomplishment of processes of societal interest, such as biofuel synthesis. On the theoretical side, we develop (deterministic or stochastic) models of microbial dynamics similar to those in the three other research axes, which can be used to investigate new control approaches for microbial communities. On the experimental side, the application of control strategies for biotechnological applications requires the engineering of microbial strains and the automation of experiments. To that aim we have been developing a platform for feedback control experiments allowing the real-time monitoring, data processing, evaluation, and application of control laws.
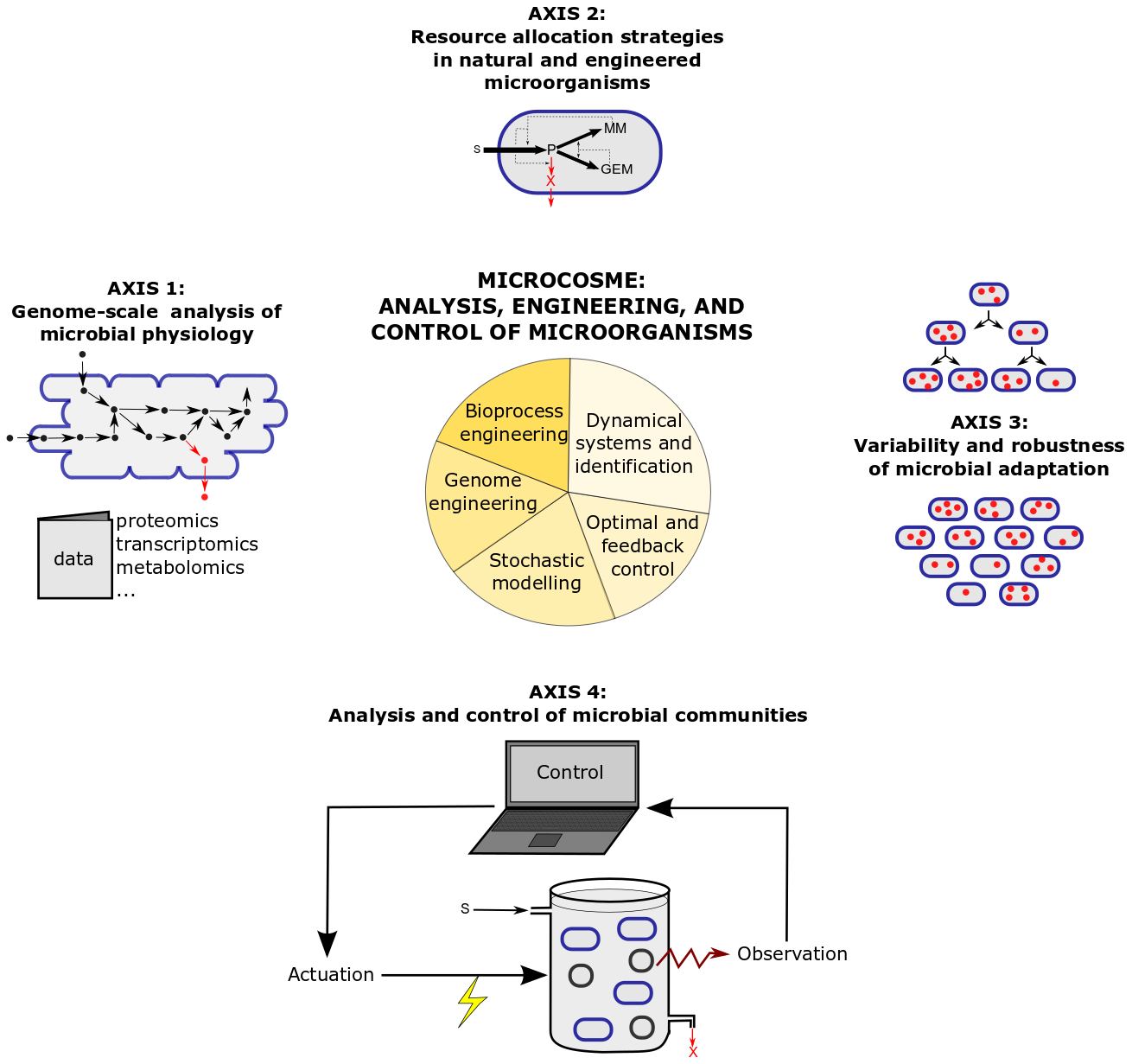
Figure 2: Research axes and methods in the MICROCOSME project-team.
4 Application domains
The research agenda of MICROCOSME is interdisciplinary in nature, driven by fundamental questions in biology, which we address by a combination of mathematical, computational, and experimental tools. This enables us to develop and share with partners a know-how useful to address challenging problems in health, bioeconomy, and biotechnology.
4.1 Biotechnology and bioeconomy
Bioproduction imposes a strong metabolic burden on microorganisms, detrimental to their growth and the production yield. Our studies of natural resource allocation strategies lead us to explore and engineer various reallocation strategies to improve bioproduction through growth control. For instance, in the past, we have successfully implemented a growth switch in E. coli bacteria, aiming at shuttling resources, away from protein synthesis (key for bacterial growth) to the high-yield production of a metabolite of interest (glycerol) 727. We also develop and test control strategies for synthetic microbial communities, composed of populations of different E. coli strains or in consortia with other species. In the wake of our studies on the relation between growth and metabolism, we develop bioeconomy strategies for the valorisation of vegetal waste into value-added product. In one project, notably, we aim at designing E. coli strains able to efficiently degrade carbon sources derived from pre-treatment of agricultural and forestry residues (Section 9).
4.2 Health
Numerous Mycobacteria species pose serious threats to human and animal health. Mycobacteria tuberculosis strains are also known to withstand several of the antibiotics used to treat the infection. We have started to extend our microbial physiology analyses by means of constraint-based models to understand the molecular control of mycobacterial growth and characterize the relations between metabolism, pathogenicity, and growth phenotype of mycobacterial species. This may lead, in the long term, to the development of new treatments for curing tuberculosis and other mycobacterial infections.
5 Social and environmental responsibility
Several of our research activities have a direct societal impact. Our work on Mycobacteria addresses important questions of public health, while the project on the degradation and valorisation of vegetal waste meet European efforts in Circular Bioeconomy to replace fossil feedstock with renewable resources. Our recently accepted Clean Energy Transition Partnership project will allow us to address the issue of microbial risks associated with underground gas storage, as part of Europe's efforts to develop innovative energy system solutions towards net-zero by 2050.
6 Highlights of the year
In a recent paper published in eLife 14, we developed a coarse-grained resource allocation model to investigate the quantitative relationship between growth rate and growth yield in different strains of the bacterium Escherichia coli. The model, which has been validated using a large number of datasets, shows that there is no simple correlation between growth rate and growth yield, as is usually assumed.
A recent collaboration with BRGM has led MICROCOSME to explore new avenues in environmental microbiology as part of the European HyLife project on microbial risks associated with hydrogen underground storage.
7 New software, platforms, open data
7.1 New software
7.1.1 ODIN+
-
Name:
Platform for advanced monitoring, control and optimisation of bioprocesses
-
Keywords:
Systems Biology, Biotechnology, Automatic control, Monitoring
-
Functional Description:
This application proposes a framework for on-line supervision of bioreactors. It gathers the data sampled from different on-line and off-line sensors. ODIN+ is a distributed platform, enabling remote monitoring as well as remote data acquisition.More originally, it enables researchers and industrials to easily develop and deploy advanced control algorithms, optimisation strategies, together with estimates of state variables or process state. It also contains a process simulator which can be harnessed for experimentation and training purposes. It is modular in order to adapt to any plant and to run most of the algorithms, and it can handle the high level of uncertainties that characterises the biological processes. The architecture is based on Erlang, and communication between modules through a MQTT Broker with Python for running the algorithms. ODIN+ is developed in collaboration with the INRIA MICROCOSME research team.
-
Authors:
Nicolas Niclausse, Nicolas Chleq, Jean-Luc Szpyrka, Pierre Fernique, Thibaud Kloczko, Tamas Muszbek, Amine Lahouel, Olivier Bernard, Eugenio Cinquemani
-
Contact:
Olivier Bernard
7.1.2 GNA
-
Name:
Genetic Network Analyzer
-
Keywords:
Model Checking, Bioinformatics, Gene regulatory networks, Qualitative simulation
-
Scientific Description:
Genetic Network Analyzer (GNA) is the implementation of methods for the qualitative modeling and simulation of gene regulatory networks developed in the IBIS (now MICROCOSME) project-team.
-
Functional Description:
The input of GNA consists of a model of the regulatory network in the form of a system of piecewise-linear differential equations (PLDEs), supplemented by inequality constraints on the parameters and initial conditions. From this information, GNA generates a state transition graph summarizing the qualitative dynamics of the system. In order to analyze large graphs, GNA allows the user to specify properties of the qualitative dynamics of a network in temporal logic, using high-level query templates, and to verify these properties on the state transition graph by means of standard model-checking tools, either locally installed or accessible through a remote web server.
-
Release Contributions:
(1) it supports the editing and visualization of regulatory networks, in an SBGN-compatible format, (2) it semi-automatically generates a prototype model from the network structure, thus accelerating the modeling process, and (3) it allows models to be exported in the SBML Qual standard.
- Publications:
-
Contact:
Hidde de Jong
-
Participants:
Hidde de Jong, Delphine Ropers
-
Partner:
UGA
7.1.3 WellInverter
-
Name:
WellInverter
-
Keywords:
Bioinformatics, Statistics, Data visualization, Data modeling
-
Scientific Description:
WellInverter is a web application that implements linear inversion methods for the reconstruction of gene expression profiles from fluorescent or luminescent reporter gene data. WellInverter makes the methods available to a broad audience of biologists and bioinformaticians. In particular, we have put in place a parallel computing architecture with a load balancer to distribute the analysis queries over several back-end servers, redesigned the graphical user interface, and developed a plug-in system for defining high-level routines for parsing data files produced by microplate readers from different manufacturers.
-
Functional Description:
As input, WellInverter reads the primary data file produced by a 96-well microplate reader, containing time-series measurements of the absorbance (optical density) as well as the fluorescence and luminescence intensities in each well (if available). Various modules exist to analyze the data, in particular for detecting outliers, subtracting background, estimating growth rates, promoter activities and protein concentrations, visualizing expression profiles, synchronizing replicate profiles, etc. The computational core of the web application consists of the Python library WellFARE.
- URL:
- Publications:
-
Contact:
Hidde de Jong
-
Participants:
Delphine Ropers, Hidde de Jong, Johannes Geiselmann
-
Partner:
UGA
7.1.4 WellFARE
-
Name:
WellFARE
-
Keywords:
Bioinformatics, Statistics, Data visualization, Data modeling
-
Scientific Description:
WellFARE is a Python library implementing linear inversion methods for the reconstruction of gene expression profiles from fluorescent or luminescent reporter gene data. WellFARE form the computational core of the WellInverter web application.
-
Functional Description:
As input, WellFARE reads the primary data file produced by a 96-well microplate reader, containing time-series measurements of the absorbance (optical density) as well as the fluorescence and luminescence intensities in each well (if available). Various functions exist to analyze the data, in particular for detecting outliers, subtracting background, estimating growth rates, promoter activities and protein concentrations, visualizing expression profiles, synchronizing replicate profiles, etc. WellFARE is the computational core of the web application WellInverter.
- URL:
- Publication:
-
Contact:
Hidde de Jong
-
Participants:
Delphine Ropers, Johannes Geiselmann, Hidde de Jong
-
Partner:
UGA
7.2 New platforms
Participants: Soraya Arias, Eugenio Cinquemani, Johannes Geiselmann, Ludovic Léau-Mercier.
7.2.1 Automated mini-bioreactor platform for (dynamical) monitoring and control of microbial cultures
Advanced dynamical experiments with microbial cultures require regular, complex measurement operations over several days or weeks. Manual execution by human operators is error-prone and exposed to weak reproducibility, besides being a poor utilisation of human resources. Reactive control experiments, in addition, necessitate online calculation of control actions in response to all acquired measurements. MICROCOSME is actively developing an automated platform for automated monitoring and reactive control experiments on microbial cultures. The platform consists of a system of mini-bioreactors connected to nutrient sources and measurement devices via a pump-based fluidic network, and it also supports optogenetic control. Computer-operated by software ODIN+ (Section 7.1) as well as via platform-specific software developments, it enables automated monitoring and online data processing, as already achieved in week-long experiments over several bioreactors 21, and it will be exploited for feedback control experiments as part of the ongoing (Section 9) and future research projects of the group.
8 New results
8.1 Resource allocation models of bacterial growth
Participants: E. Cinquemani, H. de Jong, J. Geiselmann, A. Pavlou, D. Ropers.
Various mathematical approaches have been used in the literature to describe the networks of biochemical reactions involved in microbial growth. With different levels of detail, the resulting models provide an integrated view of these reaction networks, including the transport of nutrients from the environment and their conversion into biomass. Recent work has shown that coarse-grained models of resource allocation can account for a number of empirical regularities relating the macromolecular composition of the cell to the growth rate. A well-known example of such a growth law is the linear relation between the growth rate and the ribosomal protein concentration.
In collaboration with the BIOCORE project-team and Tomas Gedeon, invited professor from Montana State University in our team in 2019, we developed a coarse-grained resource allocation model to account for the observation that different strains of a microorganism growing in the same environment display a wide variety of growth rates and growth yields. We used our model to test the hypothesis that different resource allocation strategies, corresponding to different compositions of the proteome, can account for this rate-yield variability. The model predictions were verified by means of a database of hundreds of published rate-yield and uptake-secretion phenotypes of Escherichia coli strains grown in standard laboratory conditions. We found a very good quantitative agreement between the range of predicted and observed growth rates, growth yields, and glucose uptake and acetate secretion rates. These results support the hypothesis that resource allocation is a major explanatory factor of the observed variability of growth rates and growth yields across different bacterial strains. The model thus provides a fundamental understanding of quantitative bounds on rate and yield in E. coli and other microorganisms. It may also be useful for the rapid screening of strains in metabolic engineering and synthetic biology. An article based on this work has been published in the journal eLife 14.
While these studies focus on steady-state or balanced growth, such conditions are rarely found in natural habitats, where microorganisms are continually challenged by environmental fluctuations. This requires microbial cells to continually adapt their functioning and the allocation of resources to gene expression and different metabolic functions. In the framework of the PhD thesis of Antrea Pavlou, we studied resource allocation on the experimental level using a combination of fluorescent reporter genes, time-lapse fluorescence microscopy and statistical inference techniques. The results of this study are currently being prepared for publication.
8.2 Community text book: Economic Principles in Cell Biology
Participant: H. de Jong.
The Economic Cell Collective is a group of researchers, informally coordinated by Wolfram Liebermeister (INRAE), with an interest in the analysis of the functioning of (microbial) cells from an economic perspective. In particular, the growth of microorganisms is seen as an optimization problem consisting in the allocation of limiting resources to cellular functions so as to maximize growth rate or another fitness criterion. One of the most visible outputs of the collective is a free, on-line textbook distributed via a dedicated website and Zenodo. The first version of the book was released in July 2023; updated versions are released every three months. H. de Jong participated in the writing of two chapters of the book. A first chapter deals with coarse-grained models of cellular growth and how they are able to reproduce well-known empirical growth laws 18. A second chapter deals with the use of dynamic optimization to predict the adaptation of microbial physiology to changes in the environment 19.
8.3 Biotechnological applications of bacterial growth control
Participants: H. de Jong, J. Geiselmann, T. Clavier.
The ability to experimentally control the growth rate is crucial for studying bacterial physiology. It is also of central importance for applications in biotechnology, where often the goal is to limit or even arrest growth. Growth-arrested cells with a functional metabolism open the possibility to channel resources into the production of a desired metabolite, instead of wasting nutrients on biomass production. In recent years we obtained a foundational result for growth control in bacteria 7, in that we engineered an E. coli strain where the transcription of a key component of the gene expression machinery, RNA polymerase, is under the control of an inducible promoter. By changing the inducer concentration in the medium, we can adjust the RNA polymerase concentration and thereby switch bacterial growth between zero and the maximal growth rate supported by the medium. The publication also presented a biotechnological application of the synthetic growth switch in which both the wild-type E. coli strain and our modified strain were endowed with the capacity to produce glycerol when growing on glucose. Cells in which growth has been switched off continue to be metabolically active and harness the energy gain to produce glycerol at a twofold higher yield than in cells with natural control of RNA polymerase expression, putting the yield very close to the theoretical maximum.
In the framework of the PhD thesis of Thibault Clavier, we addressed one of the limits of the growth switch that is inherent to the construction of synthetic networks in living microorganisms, namely that the latter evolve over time under the pressure of natural selection. In the case of the growth switch, the selection pressure is particularly high, since any spontaneously arising mutations disabling the inhibition of the expression of the RNA polymerase subunits will cause the population of growth-arrested cells to be taken over by cells that have resumed growth (at the expense of metabolite production). We improved the genetic stability of the growth switch by means of a redundant control mechanism of RNA polymerase expression, reducing the escape frequency to less than one in cells. We deposited a patent of this invention. An article describing the results obtained with the extended growth switch has been submitted. Moreover, this work is the subject of the start-up project Switch2Prod, led by Thibault Clavier.
8.4 Synthetic microbial communities for bioproduction processes: modelling, analysis and real-time monitoring
Participants: S. Arias, R. Asswad, E. Boucher, E. Cinquemani, T. Clavier, H. de Jong, J. Geiselmann, L. Léau-Mercier, M. Sangster.
Modelling, analysis and control of microbial community dynamics is a fast-developing subject with great potential implications in the understanding of natural processes and the enhancement of biotechnological processes. Within the now-ended project IPL COSY, we picked up the challenge to design and investigate the dynamics of synthetically engineered microbial communities with a consortium of Inria partners, and to test control strategies in vivo.
In MICROCOSME, in particular, we have addressed the design of a bacterial community of two E. coli strains, mimicking mutualistic relationships found in nature, and with the potential to outperform a producer strain working in isolation in the production of a heterologous protein. We developed an ODE model of the key growth phenotypes of the community and their interactions, calibrated the model on literature data, and analysed the model for an in-depth understanding of the conditions supporting coexistence and of the tradeoffs encountered in this production process 10. With the work of Maaike Sangster, who defended her Ph.D. in May 2023, we have bioengineered one version of the consortium, characterized individual species in batch as well as explored the consortium growth in continuous-flow experiments on an automated platform. Somewhat unexpected results have been obtained experimentally, which led to advancements on the modelling and analysis of E. coli overflow metabolism in the context of microbial consortia, as included in the thesis 21, and gave us new indications for modifications of the consortium that should enable co-existence. The finalization of this work by a joint effort of several team members is expected to lead to a paper submission in the near future. As a subsequent challenge, feedback control of the community for stabilization of biomasses to desirable bioproduction regimes remains in the aims of this research direction.
In parallel, we have pushed further the investigation of the synthetic community design for bioproduction optimization. In collaboration with BIOCORE, we considered a generalized version of the synthetic consortium proposed in 10 where both species of the consortium can synthesize a product of interest. In addition to a theoretical analysis of equilibria and coexistence regimes, we showed that the generalized design can indeed outperform our original consortium design based on a single producer species, thus shifting the interest from the commonly evoked concept of division of labor (different tasks for different species) to distribution of labor (same task across different species). This work is now published in 16.
The ANR Ctrl-AB puts related concepts to work for the synergistic growth of E. coli and microalgae. As per project goals, vitamin synthesis by modified E. coli strains represents a leverage to control vitamin-dependent growth of suitable algal strains, with the potential to robustify and maximize algal lipid synthesis. On the experimental side, we advanced on the design and synthesis of the bacterial strain in the consortium. Two alternative strategies for controlling bacterial vitamin synthesis via optogenetics are being explored and, together with the lab advancements on optogenetic systems and experimental equipment, will be tested next. On the methodological side, in collaboration with BIOCORE, modelling, analysis and control problems are being addressed with the Ph.D. thesis of Rand Asswad, started in 2022. Toward model-based feedback control of microbial communities, we focused so far on the problem of real-time state estimation of microbial growth in a bioreactor. In connection with a simple algal-bacterial growth model established within Ctrl-AB, we developed, theoretically analyzed and demonstrated in simulation estimation methods for a single species that can be generalized to co-cultures. The work resulted so far in a conference submission, currently under review. It will be extended to co-cultures and integrated into the upcoming work on control of algal-bacterial growth and interaction dynamics in a bioreactor.
8.5 Modelling of carbon and mRNA metabolism in microorganisms
Participants: A. Belcour, I. Cancino-Aguirre, E. Cinquemani, M. Cocaign-Bousquet, H. de Jong, D. Ropers.
The ability to rapidly respond to changing nutrient availability is crucial for microorganisms to survive in many environments. The first step in adjusting metabolism is to reorganise gene expression. It involves fine-tuning of both transcription and mRNA stability by dedicated regulatory interactions. While transcriptional regulation has been extensively studied, the role of mRNA stability during a metabolic switch is poorly understood and often overlooked, as shown by our recent compilation of literature evidence in E. coli, in collaboration with the team of Muriel Cocaign-Bousquet at the Toulouse Biotechnology Institute 28. We also addressed this question in 12, where we combined genome-wide transcriptome and mRNA decay analyses to investigate the role of mRNA stability in the response of E. coli to nutrient changes. We demonstrated that while transcription of most genes is down-regulated when glucose is exhausted, concomitant mRNA stabilization of many mRNAs occurs and leads to the upregulation of genes involved in responses to nutrient changes and stresses. The observation of a global stabilization of cellular mRNAs during adaptation to carbon source depletion raises questions about the regulatory mechanisms at work. Are these regulatory mechanisms sufficient to explain the systematic adjustment of mRNA half-lives?
In a follow-up study, we developed a kinetic model of mRNA degradation that allows to propose hypotheses on the regulatory mechanisms at work to adjust mRNA stability to environmental conditions 4. From a practical point of view, this amounts to infer model parameters from high-throughput biological datasets. In the framework of the ANR project RIBECO (Section 9), we have developed a nonlinear mixed-effects modelling approach for parameter estimation of large-scale mechanistic models from time-series transcriptomics data, which enables biological interpretation of microarray and RNA-Seq gene expression profiles. When integrated in a model describing the degradation kinetics of 4254 mRNAs in E. coli cells, the data allowed to identify a new post-transcriptional regulatory mechanism and its targets. The modelling results are being prepared for publication, and their experimental demonstration will be at the heart of the newly accepted ANR RECOM project, in which the Toulouse Biotechnology Institute and MICROCOSME are partners.
Genome-scale reconstructions of cellular metabolism, such as those used in 12, are often available in public databases for well-studied microorganisms, but this is not the case for poorly studied species. As part of I. Cancino-Aguirre's PhD thesis co-advised by D. Ropers and H. de Jong, Arnaud Belcour's postdoc, and the associate-team GERM (Section 9), we aim to develop such reconstructions from genome sequences for the Mycobacterium genus. They will then be combined with growth kinetics data to analyse how differences in carbon metabolism account for the variability in growth rates of mycobacterial species, which include both dangerous pathogens and non-pathogenic bacteria. Inferring metabolic function from sequenced genomes is a difficult problem, but even more so when dealing with microbial communities in natural environments. In a new collaboration with BRGM and the European project HyLife (Section 9), Arnaud Belcour, Hidde de Jong and Delphine Ropers are beginning to address this issue, by using marker gene sequence data to characterise the metabolic activity of underground microbial communities, which may influence the quality of gas storage in underground reservoirs.
8.6 Inference of parameters on lineage trees
Participants: E. Cinquemani, C. Fonte Sanchez, A. Marguet, E. Reginato.
Recent technological developments have made it possible to obtain single-cell measurements of gene expression and, in some cases, the associated lineage information. However, most of the existing methods for the identification of mathematical models of gene expression do not account for the fact that cells undergo divisions and are related to one another through parental relationships. Most methods developed for single-cell data make the simplifying assumptions that cells in a population are independent, thus ignoring cell lineages. The development of statistical tools taking into account the correlations between individual cells is needed in particular to enable the investigation of inheritance of traits in bacterial populations.
With the near-ending Ph.D. project of E. Reginato, we have advanced in the study of tree-structured single-cell gene expression models with mother-daughter inheritance that we had started in a previous publication 9. Contrary to this previous work, where model inference methods from single-cell gene expression data were developed assuming knowledge of the lineage of the observed cells, the thesis project has been focused on the case where lineage information is not available. We notably explored how well inheritance model parameters can be inferred depending on absence or presence of dynamics in the mean and variance data. In relation with certain literature datasets obtained by videomicroscopy, we developed statistically exact maximum-likelihood estimation methods leveraging correlation of empirical means along generations, and approximate methods also exploiting variance data that we showed to perform well even in absence of transient mean dynamics. Results constitute the object of a conference submission currently under review. Demonstration on the reference datasets is being finalized.
While the above modelling approach to mother-daughter inheritance is of statistical nature, it can be related with mechanisms into play at cell division. For this, within the same Ph.D., we started looking at the regulation of the repartition of multicopy resistance plasmids at cell division and its impact on population growth in selective media. We have developed and compared several individual-based models, with first results on the interplay between plasmid repartition statistics and stochastic cell division modelling in the understanding of the emerging population dynamics. This will be completed and analysed using real data in a future publication. Related to the above studies, Claudia Fonte Sanchez's postdoc in project AnaComBa is looking at individual-based models of population growth and gene expression, and investigating the relation between growth-rate variability across cells and gene expression distribution in population-snapshot data. Theoretical analysis is ongoing and will be subject to publication and subsequent integration with experimental investigations.
8.7 Mathematical analysis of structured branching populations
Participants: C. Fonte-Sanchez, Ch. Medous, A. Marguet.
The investigation of cellular populations at a single-cell level has already led to the discovery of important phenomena, such as the occurrence of different phenotypes in an isogenic population. Nowadays, several experimental techniques, such as microscopy combined with the use of microfluidic devices, enable one to take investigation further by providing time-profiles of the dynamics of individual cells over entire lineage trees. The development of models that take into account the genealogy is an important step in the study of inheritance in bacterial population. In particular, their mathematical analysis is essential for the efficient analysis of single cell data.
Structured branching processes allows for the study of populations, where the lifecycle of each cell is governed by a given characteristic or trait, such as the internal concentration of proteins. The dependence on this characteristic of cellular mechanisms, like division or ageing, has been explored by Aline Marguet via the mathematical analysis of those processes. In collaboration with Charline Smadi (INRAE Grenoble), Aline Marguet investigated the long-time behavior of a parasite infection in a cell population. In this work, accepted for publication in Stochastic Processes and their Applications15, the dynamics of the cell population is modelled using a structured branching process, where the cell cycle depends on the dynamics of the parasites contained in the cell. The results obtained focus on the asymptotic behaviour of the intracellular quantity of parasites. Based on criteria that depend on the comparison between the growth rate of the cell population and the parameters associated with the parasite dynamics, containment or explosion of the infection is established. The strategy of proof relies on the study of auxiliary processes that describe the level of infection of a typical cell in the population, using coupling techniques.
The fate of the infection appears to be very sensitive to the law of parasite sharing between the two daughter cells as they divide. In particular, the effect of various sharing strategies in the case of a constant division rate has been considered in a follow-up work. A perfect symmetric sharing of the parasites between the two daughter cells has been proven to be the worst strategy for the survival of the cell population. Stochastic and deterministic strategies have also been compared, highlighting that variability favors survival. This paper is under revision for Journal of Mathematical Biology25.
Spinal processes and many-to-one formulas have proved very useful for the study of complex structured branching processes, as they allow to reduce the problem to the study of a simpler lineage process. In the context of the project AnaComBa, for the study of microbial communities, such tools appear to be needed for structured branching processes with interactions. In a recently submitted paper 26, Charles Medous developed a spinal construction and established a Girsanov-type result for branching processes describing structured, interacting populations in continuous time, where the dynamics of each individual can be influenced by the entire population. He also derived a modified continuous-time version of the Kesten-Stigum theorem that incorporates interactions, and proposed an alternative simulation approach for stochastic size-dependent populations using appropriate spine constructions.
The study of the asymptotic behavior of general semigroups is important for several aspects of branching processes, especially to prove the efficiency of statistical procedures. In this context, Claudia Fonte Sanchez, in collaboration with Pierre Gabriel (Université de Tours) et Stéphane Mischler (Université Paris-Dauphine) revisited the Krein-Rutman theory for semigroups of positive operators and provided some very general, efficient and practical results with constructive estimates on the existence of a solution to the first eigentriplet problem, the geometry of the principal eigenvalue problem, and the asymptotic stability of the first eigenvector with possible constructive rate of convergence 24.
9 Partnerships and cooperations
9.1 International initiatives
9.1.1 Associate Teams in the framework of an Inria International Lab or in the framework of an Inria International Program
GERM
Participants: A. Belcour, I. Cancino-Aguirre, H. de Jong, D. Ropers.
- Title: Growth-rate control in mycobacteria: Computational exploration of metabolic strategies
-
Partner Institution(s)
- Francis Crick Institute (United Kingdom)
-
Date/Duration:
2022-2024
-
Additionnal info/keywords:
Numerous Mycobacterium species pose serious threats to human and animal health. Genome-scale mathematical models of Mycobacterium metabolism are promising avenues to uncover bottlenecks explaining the growth-rate variability and pathogenicity observed across the genus. We employ these models to integrate and analyze diverse types of experimental data, including measurements of doubling times and metabolite concentrations. The results will allow us to formulate hypotheses on the molecular mechanisms responsible for growth-rate variability and pathogenicity observed across mycobacteria. The hypotheses will be tested by targeted experiments.
9.1.2 Informal international partners
H. de Jong and D. Ropers collaborate with T. Gedeon, former invited researcher in our former team IBIS, on research allocation strategies in microorganisms. The collaboration has already resulted in a paper published this year in eLife 14.
9.1.3 Visits to international teams
Research stays abroad
I. Cancino Aguirre
- Visited institution: Francis Crick Institute
- Country: United Kingdom
-
Dates:
20/08/2023 - 23/09/2023
-
Context of the visit:
visit to the Carvalho's laboratory in the context of the associate-team GERM
-
Mobility program/type of mobility:
research stay
9.2 European initiatives
9.2.1 Horizon Europe
| Project name | HyLife: Optimal control of microbial cells by natural and synthetic strategies |
| Coordinator | N. Dopffel (NORCE, Norway) |
| MICROCOSME participants | A. Belcour, H. de Jong, D. Ropers |
| Type | Clean Energy Transition Co-funded Partnership (2023-2026) |
| Web page | Link to project description |
9.3 National initiatives
| Project name | MAXIMIC: Optimal control of microbial cells by natural and synthetic strategies |
| Coordinator | H. de Jong |
| MICROCOSME participants | E. Cinquemani, T. Clavier, J. Geiselmann, H. de Jong, A. Pavlou, D. Ropers |
| Type | ANR project (2017-2023) |
| Web page | Link to project description |
| Project name | RIBECO (RIBonucleotide ECOnomy): Engineering RNA life cycle to optimize economy of microbial energy |
| Coordinator | M. Cocaign-Bousquet |
| MICROCOSME participants | E. Cinquemani, M. Cocaign-Bousquet, D. Ropers |
| Type | ANR project (2018-2023) |
| Web page | Link to project description |
| Project name | Ctrl-AB : Optimization and control of the productivity of an algal-bacterial consortium |
| Coordinator | J.-L. Gouzé |
| MICROCOSME participants | R. Asswad, S. Arias, E. Boucher, E. Cinquemani, Th. Clavier, H. de Jong, J. Geiselmann, L. Léau-Mercier, A. Marguet, M. Sangster |
| Type | ANR project (2020-2025) |
| Project name | PlugNBio: A plug-and-play platform for reproducible microbial culture control experiments |
| Coordinator | E. Cinquemani |
| MICROCOSME participants | S. Arias, E. Boucher, E. Cinquemani, J. Geiselmann, L. Léau-Mercier |
| Type | Inria ADT (2022-2024) |
The following two projects have just been accepted and will start in spring 2024.
| Project name | ARBOREAL: Branching resource allocation processes for the analysis and inference of phenotypic growth variability |
| Coordinator | A. Marguet |
| MICROCOSME participants | E. Cinquemani, J. Geiselmann, H. de Jong, A. Marguet |
| Type | ANR project (2024-2029) |
| Web page |
| Project name | RECOM: Competition of RNAs for RNase E, a mechanism regulating their degradation and the energy and carbon metabolism in the cell |
| Coordinator | M. Cocaign-Bousquet |
| MICROCOSME participants | E. Cinquemani, M. Cocaign-Bousquet, D. Ropers |
| Type | ANR project (2023-2027) |
| Web page |
In addition to the above projects, A. Marguet contributes to the ANR JCJC NOLO of Bertrand Cloez (INRAE Montpellier) on non-local branching processes.
9.4 Regional initiatives
| Project name | ATLAS: Analysis of brain energy metabolism in the context of Parkinson’s disease |
| Coordinator | F. Fauvelle (Grenoble Institute of Neurosciences) |
| MICROCOSME participants | D. Ropers |
| Type | IXXI/BioSyl project (2022 – 2024) |
| Web page | Link to project description |
| Project name | AnaComBa: Analyse de Communautés Bactériennes : modélisation stochastique |
| Coordinator | A.Marguet & L. Coquille |
| MICROCOSME participants | E. Cinquemani, C. Fonte-Sanchez, A.Marguet, C.Medous |
| Type | Equipe-Action du LABEX Persyval (2021 – 2024) |
10 Dissemination
Participants: R. Asswad, I. Cancino Aguirre, E. Cinquemani, Th. Clavier, M. Cocaign-Bousquet, H. de Jong, J. Geiselmann, A. Marguet, Ch. Medous, D. Ropers.
10.1 Promoting scientific activities
10.1.1 Scientific events: organisation
Member of organizing committees
| MICROCOSME members | Conference, workshop, school | Date |
| Ignacia Cancino Aguirre | Inria PhD seminar, Montbonnot | 2023 |
| Hidde de Jong | Summer school on Economic Principles in Cell Physiology, Paris | Jul 2023 |
| A. Marguet | Conférence du GDR Branchement, Toulouse | Nov 2023 |
| A. Marguet | GDT Math-Bio, Lyon-Grenoble | 2023 |
10.1.2 Scientific events: selection
Chair of conference program committees
| MICROCOSME member | Conference, workshop, school | Role |
| Eugenio Cinquemani | European Control Conference (ECC 2023, ECC2024) | Associate editor |
| Eugenio Cinquemani | Special Issue of the 19th International Conference on Computational Methods in Systems Biology. BMC Bioinformatics Supplements, 24(1) 20 | Supplement editor (with Loïc Paulevé) |
Member of conference program committees
| MICROCOSME member | Conference, workshop, program |
| Eugenio Cinquemani | ECC 2023 |
10.1.3 Journal
Member of editorial boards
| MICROCOSME member | Journal |
| Hidde de Jong | Journal of Mathematical Biology |
| Hidde de Jong | Biosystems (reviews editor) |
| Johannes Geiselmann | Frontiers in Microbiology (review editor) |
10.1.4 Invited talks and seminars
Ignacia Cancino Aguirre
| Title | Event and location | Date |
| Analysis of mycobacterial metabolic variability using genome-scale metabolic models | Advanced School on Quantitative Principles in Microbial Physiology: from Single Cells to Cell Communities, ICTP Trieste, Italy | Oct 2023 |
Eugenio Cinquemani
| Title | Event and location | Date |
| Power spectral analysis for the optimal design of gene reporter systems | GdT MathBio seminars, Grenoble, France | Nov 2023 |
Thibault Clavier
| Back and forth between growth and biosynthesis : a bacterial metabolism switch system to improve bioproduction | 8th Bioproduction Congress (BIOPC2023), Lyon, France | Oct 2023 |
Hidde de Jong
| Title | Event and location | Date |
| Enhanced production of heterologous proteins in synthetic microbial consortia | Invited talk at summer school Design and Control of Microbial Communities, Leuven, Belgium | Sep 2023 |
| Modelling integrated networks of metabolism and gene expression | Blackboard session at summer school Design and Control of Microbial Communities, Leuven, Belgium | Sep 2023 |
Aline Marguet
| Title | Event and location | Date |
| Spread of parasites affecting death and division rates in a cell population | Informs Applied Probability Society Conference 2023, Nancy, France | Jun 2023 |
| Parasite infection in a cell population | "Biology meets math: building a better understanding of host-parasite dynamics in the wild", Montpellier | Sep 2023 |
| Modélisation de la prolifération de parasites dans une population de cellules | Journées Math-Bio-Santé 2023, Marne-la-Vallée, France | Nov 2023 |
| Parasite infection in a cell population | Workshop EveREvol, Grenoble, France | Dec 2023 |
Charles Medous
| Title | Event and location | Date |
| Epine pour des populations branchantes avec interactions et application à un processus de Yule | Ecole d'été de la chaire MMB, Aussois, France | Jun 2023 |
| Spinal constructions for continuous-space branching processes | Journées Jeunes Probabilistes et Statisticiens, Oléron, France | Oct 2023 |
| Mathematical modeling of interacting communities | Invited talk at Séminaire de l'unité Evolution, Ecologie et Paléontologie, Lille, France | Dec 2023 |
| Construction d'épines pour des populations stochastiques d'individus en interactions | Invited talk at Journée du Groupe de travail Math-Bio Sud-Est, Lyon | Dec 2023 |
Delphine Ropers
| Title | Event and location | Date |
| A short introduction to bioinformatics | Invited talk at Séminaire du département d'informatique de l'ENS Lyon, Le Pleynet, France | Jan 2023 |
| Mathematical modelling of energy metabolism | Invited talk, Grenoble Institute of Neurosciences, Grenoble, France | Jan 2023 |
| Doing a PhD, good practice and pitfalls to avoid | Matinée des doctorants, Centre Inria de l'Université Grenoble Alpes | Oct 2023 |
10.1.5 Scientific evaluation and expertise
| MICROCOSME member | Organism | Role |
| Johannes Geiselmann | UMR5240 CNRS-UCBL-INSA-BayerCropScience | Member scientific council |
| Hidde de Jong | Microbiology and Food Chain Department, INRAE | Member scientific council |
| Hidde de Jong | Univ Grenoble Alpes | Member scientific council of Pôle MSTIC |
| Hidde de Jong | Univ Grenoble Alpes | Member of Vice-Présidence Recherche et Innovation élargie |
| Hidde de Jong | Univ Grenoble Alpes | Member of Collège des Ecoles Doctorales |
| Hidde de Jong | Univ Grenoble Alpes | Member advisory board of LabEX PERSYVAL 3 |
| Delphine Ropers | National Science Centre, Poland | Member of Expert Panel LS2 |
10.1.6 Research administration
| MICROCOSME member | Committee | Role |
| Ignacia Cancino Aguirre | Inria - Univ. Grenoble Alpes | Member Comité du centre |
| Eugenio Cinquemani | Inria - Univ. Grenoble Alpes | Member Comité des Emplois Scientifiques (CES) |
| Eugenio Cinquemani | Inria - Univ. Grenoble Alpes | Member Comité des Utilisateurs des Moyens Informatiques (CUMI) |
| Eugenio Cinquemani | Inria - Univ. Grenoble Alpes | Member Comité Développement Technologique (CDT) |
| Hidde de Jong | Inria - Univ Grenoble Alpes | Member Direction |
| Hidde de Jong | Inria - Univ Grenoble Alpes | President Comité des Equipes Projets (CEP) |
| Hidde de Jong | Inria - Univ Grenoble Alpes | Member Comité des Emplois Scientifiques (CES) |
| Hidde de Jong | Inria - Univ Grenoble Alpes | Member Comité Développement Technologique (CDT) |
| Hidde de Jong | Inria - Univ Grenoble Alpes | Member Comité des Etudes Doctorales |
| Hidde de Jong | Inria - Univ. Grenoble Alpes | President scientific council (COS) |
| Hidde de Jong | Inria | Member Commission d'évaluation (CE) |
| Aline Marguet | Inria - Univ. Grenoble Alpes | Member Comité du centre |
| Aline Marguet | Inria - Univ. Grenoble Alpes | Member Comité des études doctorales |
| Delphine Ropers | Inria - Univ. Grenoble Alpes | Référente chercheurs |
10.1.7 Recruitment committees
| MICROCOSME member | Organism | Recruitment |
| Hidde de Jong | Inria national | CRHC (jury de promotion) |
| Hidde de Jong | Inria | DR2 (jury d'admissibilité) |
| Hidde de Jong | Inria - Univ Grenoble Alpes | ISFP (jury d'admission) |
| Delphine Ropers | Inria - Univ Grenoble Alpes | CRCN/IFSP (jury d'admissibilité) |
| Delphine Ropers | INSA Lyon | Selection committee Professor |
| Delphine Ropers | Université Toulouse III | Selection committee Professor |
| Delphine Ropers | Inria - Université Côte d'Azur | CRCN/IFSP 2024 (Présidente du jury d'admissibilité) |
10.2 Teaching - Supervision - Juries
10.2.1 Teaching
D. Ropers received the title of Full Professor ("Professeur attaché") at Univ Grenoble Alpes for 3 years (2023 - 2026) in recognition of her teaching activity.
J. Geiselmann is full professor at Univ Grenoble Alpes. He therefore has a full teaching service (at least 192 hours per year) and administrative duties related to the organization and evaluation of the university course programs on all levels.
The following people have also contributed to courses last year.
Rand Asswad
- Practicals: Introduction to applied mathematics - image processing, L1, Computer Science (18 h)
- Course and practicals: Introduction to scientific programming, L1, Computer Science (24 h)
Ignacia Cancino Aguirre
- Practicals: Cellular biology, L2, Life Sciences, Univ Grenoble Alpes (10 h)
- Practicals: Biotechnology of membrane and cell systems, M1, TD, Univ Grenoble Alpes (10 h)
Eugenio Cinquemani
- Course: Statistics for systems biology, M1, Master AIRE (Interdisciplinary Approaches in Research and Education)/Learning planet institute, Université Paris Cité (20 h, also in charge of 20 h of practicals)
- Course: Modelling and identification of metabolic networks, M1, Phelma, INP Grenoble (4 h)
Hidde de Jong
- Course and practicals: Modeling and simulation of gene regulatory networks, M2, BIM, INSA de Lyon (25 h)
Aline Marguet
- Practicals: Biostatistics, M2, Univ Grenoble Alpes (27 h)
Charles Medous
- Practicals: Introduction to applied mathematics - image processing, L1, Computer Science (36 hours)
Delphine Ropers
- Course and practicals: Modelling in systems biology, M1, Phelma, INP Grenoble (16 h)
- Course and practicals: Cell systems biology and modelling cell functions, M1, Master ingéniérie de la santé, Univ Grenoble Alpes (24 h)
- Course: Modelling and simulation of genetic regulatory networks, M2, INSA de Toulouse (4 h)
- Course: Metabolic modelling with omics data, M2, IA4 Health International master course, Univ Grenoble Alpes (6 h)
10.2.2 Supervision
- PhD in progress: Rand Asswad, Development of control strategies for synthetic microbial consortia. Supervisors: Eugenio Cinquemani and Jean-Luc Gouzé (Inria - Univ Côte d'Azur)
- PhD in progress: Ignacia Cancino Aguirre, Computational analysis of metabolic strategies in pathogenic bacteria. Supervisors: Delphine Ropers and Hidde de Jong
- PhD in progress: Thibault Clavier Genetic growth control to maximize the bioproduction in E. coli. Supervisors: Johannes Geiselmann and Hidde de Jong
- PhD in progress: Charles Medous, Analysis of bacterial communities: stochastic modelling. Supervisors: Loren Coquille (Institut Fourier, Grenoble), Aline Marguet, Charline Smadi (Inrae Grenoble)
- PhD in progress: Emrys Reginato, Development, analysis, and inference of stochastic models of gene expression in growing cell populations. Supervisors: Eugenio Cinquemani and Aline Marguet
- PhD completed: Maaike Sangster, Development, characterization and control of E. coli communities on an automated experimental platform. Supervisors: Eugenio Cinquemani and Johannes Geiselmann
10.2.3 Juries
PhD thesis committees
| MICROCOSME member | Role | PhD student | University | Date |
| Eugenio Cinquemani | Maaike Sangster | Co-directeur | Univ Grenoble Alpes | May 2023 |
| Eugenio Cinquemani | Marielle Péré | Rapporteur | Univ Côte d'Azur | Jul 2023 |
| Hidde de Jong | Rapporteur | Leo Diaz | Univ Melbourne, Australia | Jul 2023 |
| Johannes Geiselmann | Examinateur | Clément Caffaratti | Univ Grenoble Alpes | Jan 2023 |
| Johannes Geiselmann | Rapporteur | Dimitrije Milunov | Univ Paris Cité | Feb 2023 |
| Johannes Geiselmann | Examinateur | Sara Badawi | Univ Grenoble Alpes | Mar 2023 |
| Johannes Geiselmann | Directeur | Maaike Sangster | Univ Grenoble Alpes | May 2023 |
| Johannes Geiselmann | Rapporteur | Lexuan Liu | Sorbonne Université | Sept 2023 |
| Johannes Geiselmann | Examinateur | Victor Simon | Univ Grenoble Alpes | Oct 2023 |
| Johannes Geiselmann | Examinateur | Agate Nidriche | Univ Grenoble Alpes | Dec 2023 |
| Johannes Geiselmann | Rapporteur | Yuliia Shymko | Univ Paris-Saclay | Dec 2023 |
| Johannes Geiselmann | Examinateur | Clémence Dupont Thibert | Univ Grenoble Alpes | Dec 2023 |
| Delphine Ropers | Rapporteur | Maxime Mahout | Univ Paris-Saclay | Nov 2023 |
| Delphine Ropers | Rapporteur | Matteo Bouvier | ENS Lyon | Dec 2023 |
Habilitation (HDR) committees
| MICROCOSME member | Role | HDR candidate | University | Date |
| Eugenio Cinquemani | Paolo Ballarini | Rapporteur | CentraleSupélec, Université Paris-Saclay | Jun 2023 |
PhD advisory committees
| MICROCOSME member | PhD student | University |
| Eugenio Cinquemani | Marielle Peré | Univ Côte d'Azur |
| Johannes Geiselmann | Clément Caffaratti | Univ Grenoble Alpes |
| Delphine Ropers | Paul Ahavi | Univ Paris Saclay |
10.2.4 Teaching administration
- Eugenio Cinquemani organizes a module on statistics in systems biology at Learning planet institute, Université Paris Cité.
- Delphine Ropers organizes a module on the mathematical modelling of biological systems at PHELMA, INP Grenoble.
- Hidde de Jong organizes a module on the modelling of genetic and metabolic networks at INSA de Lyon.
10.3 Popularization
10.3.1 Interventions
Charles Medous
| Title | Event and location | Date |
| Mathematical games for children from primary (CE2) to high school | Club maths "Les maths Autrement", Institut Fourier, Univ Grenoble Alpes | Organisation and animation of 2 sessions of 2 hours/month |
Aline Marguet
| Title | Event and location | Date |
| Modélisation mathématique des populations de cellules | Invited talk at the "cérémonie de remise de prix des Olympiades de Mathématiques", Académie de Grenoble | May 2023 |
| Modélisation mathématique des populations de cellules | Seminar to high school students (2de), Fête de la Science, Montbonnot |
11 Scientific production
11.1 Major publications
- 1 articleResource allocation accounts for the large variability of rate-yield phenotypes across bacterial strains.eLife12May 2023, 1-29HALDOI
- 2 articleEstimation of time-varying growth, uptake and excretion rates from dynamic metabolomics data.Bioinformatics33142017, i301-i310HALDOI
- 3 articleStochastic reaction networks with input processes: Analysis and application to gene expression inference.Automatica1012019, 150-156HALDOI
- 4 articleCompetitive effects in bacterial mRNA decay.Journal of Theoretical Biology504November 2020HALDOIback to text
- 5 articleDynamical allocation of cellular resources as an optimal control problem: Novel insights into microbial growth strategies.PLoS Computational Biology123March 2016, e1004802HALDOI
- 6 articleStatistical estimation in a randomly structured branching population.Stochastic Processes and their Applications129122019, 5236-5277HALDOI
- 7 articleA synthetic growth switch based on controlled expression of RNA polymerase.Molecular Systems Biology1111November 2015, 840HALback to textback to text
- 8 articleWhat population reveals about individual cell identity: Single-cell parameter estimation of models of gene expression in yeast.PLoS Computational Biology122February 2016, e1004706HALDOI
- 9 articleInheritance and variability of kinetic gene expression parameters in microbial cells: modeling and inference from lineage tree data.Bioinformatics35142019, i586-i595HALDOIback to text
- 10 articleEnhanced production of heterologous proteins by a synthetic microbial community: Conditions and trade-offs.PLoS Computational Biology1642020, e1007795HALDOIback to textback to text
- 11 articleThe Csr System Regulates Escherichia coli Fitness by Controlling Glycogen Accumulation and Energy Levels.mBio85October 2017, 1-14HALDOI
- 12 articleThe post-transcriptional regulatory system CSR controls the balance of metabolic pools in upper glycolysis of Escherichia coli.Molecular Microbiology1004May 2016, 686-700HALDOIback to textback to text
- 13 articleAcetate metabolism and the inhibition of bacterial growth by acetate.Journal of Bacteriology20113July 2019, 147 - 166HALDOI
11.2 Publications of the year
International journals
Invited conferences
Scientific book chapters
Edition (books, proceedings, special issue of a journal)
Doctoral dissertations and habilitation theses
Reports & preprints
11.3 Cited publications
- 27 articleMultiomics Study of Bacterial Growth Arrest in a Synthetic Biology Application.ACS Synthetic Biology1011November 2021, 2910-2926HALDOIback to text
- 28 articleThe essential role of mRNA degradation in understanding and engineering E. coli metabolism.Biotechnology Advances2022, 1-13HALDOIback to text
