2023Activity reportProject-TeamSODA
RNSR: 202224249S- Research center Inria Saclay Centre
- Team name: Computational and mathematical methods to understand health and society with data
- Domain:Digital Health, Biology and Earth
- Theme:Computational Neuroscience and Medicine
Keywords
Computer Science and Digital Science
- A3.3. Data and knowledge analysis
- A3.4. Machine learning and statistics
- A9.1. Knowledge
- A9.2. Machine learning
Other Research Topics and Application Domains
- B2.3. Epidemiology
- B9.1. Education
- B9.5.6. Data science
- B9.6.1. Psychology
- B9.6.3. Economy, Finance
1 Team members, visitors, external collaborators
Research Scientists
- Gael Varoquaux [Team leader, INRIA, Senior Researcher, HDR]
- Judith Abecassis [INRIA, ISFP]
- Marine Le Morvan [INRIA, Researcher]
- Jill Jenn Vie [INRIA, Researcher]
Post-Doctoral Fellows
- Riccardo Cappuzzo [INRIA, Post-Doctoral Fellow]
- Lihu Chen [INRIA, Post-Doctoral Fellow, from Jul 2023]
- Myung Kim [INRIA, Post-Doctoral Fellow]
- Clémence Reda [UNIV POTSDAM, Post-Doctoral Fellow, from Jul 2023]
PhD Students
- Julie Alberge [INRIA, from Oct 2023]
- Samuel Brasil De Albuquerque [INSERM]
- Bénédicte Colnet [INRIA, until Jun 2023]
- Alexis Cvetkov-Iliev [INRIA, until Jan 2023]
- Matthieu Doutreligne [HAS, from Nov 2023]
- Matthieu Doutreligne [HAS, until Sep 2023]
- Samuel Girard [INRIA, from Oct 2023]
- Leo Grinsztajn [INRIA]
- Felix Lefebvre [INRIA]
- Alexandre Perez [INRIA]
- Jovan Stojanovic [INRIA, from Oct 2023]
Technical Staff
- David Arturo Amor Quiroz [INRIA, Engineer]
- Franck Charras [INRIA, Engineer]
- Jerome Dockes [INRIA, Engineer, from Sep 2023]
- Jeremie Du Boisberranger [INRIA, Engineer]
- François Goupil [INRIA, Engineer]
- Olivier Grisel [INRIA, Engineer]
- Julien Jerphanion [INRIA, Engineer, until Mar 2023]
- Guillaume Lemaitre [INRIA, Engineer]
- Vincent Maladiere [INRIA, Engineer]
- Tomas Rigaux [INRIA, Engineer, until Sep 2023]
- Jovan Stojanovic [INRIA, Engineer, until Sep 2023]
Interns and Apprentices
- Julie Alberge [INRIA, Intern, from Apr 2023 until Aug 2023]
- Lilian Boulard [INRIA, Apprentice, until Sep 2023]
- Sami Boumaiza [INRIA, Intern, from May 2023 until Jul 2023]
- Samuel Girard [INRIA, Intern, from Apr 2023 until Sep 2023]
- Nour Oueghlani [INRIA, Intern, from Jun 2023 until Aug 2023]
Administrative Assistant
- Marie Enee [INRIA]
Visiting Scientist
- Ko Takeuchi [Kyoto university]
External Collaborators
- Theo Jolivet [AP/HP]
- Yann Lechelle [SENS DIGITAL, from Apr 2023]
- Anand-Arnaud Pajaniradjane [CENTRALESUPELEC, until Nov 2023]
2 Overall objectives
2.1 Context
2.1.1 Application context: richer data in health and social sciences
Opportunistic data accumulations, often observational, bare great promises for social and health sciences. But the data are too big and complex for standard statistical methodologies in these sciences.
Health databases
Increasingly rich health data is accumulated during routine clinical practice as well as for research. Its large coverage brings new promises for public health and personalized medicine, but it does not fit easily in standard biostatistical practice because it is not acquired and formatted for a specific medical question.
Social, educational, and behavioral sciences
Better data sheds new light on human behavior and psychology, for instance with on-line learning platforms. Machine learning can be used both as a model for human intelligence and as a tool to leverage these data, for instance improving education.
Likewise, activity traces can provide empirical evidence for economical or political science, but their complexity requires new statistical practices.
2.1.2 Related data-science challenges
Data management: preparing dirty data for analytics
Assembling, curating, and transforming data for data analysis is very labor intensive. These data-preparation steps are often considered the number one bottleneck to data-science. They mostly rely on data-management techniques. A typical problem is to establish correspondences between entries that denote the same entities but appear in different forms (entity linking, including deduplication and record linkage). Another time-consuming process is to join and aggregate data across multiple tables with repetitions at different levels (as with panel data in econometrics and epidemiology) to form a unique set of “features” to describe each individual. This process is related to database denormalization and might require schema alignment when performed across multiple data sources with imperfect correspondence in columns.
Progress in machine learning increasingly helps automating data preparation and processing data with less curation.
From machine learning to statistically-valid answers
Machine learning can be a tool to answer complex domain questions by providing non-parametric estimators. Yet, it still requires much work, eg to go beyond point estimators, to derive non-parametric procedures that account for a variety of bias (censoring, sampling biases, non-causal associations), or to provide theoretical and practical tools to assess validity of estimates and conclusion in weakly-parametric settings.
A question that is increasingly important in all applications of machine learning is that of auditing the model used in practice. This question arises in fundamental-research settings (medical research, political science...) for statistical validity, and in applications to assess societal biases, or safety of AI systems.
3 Research program
3.1 Table representation learning
Soda develops develop deep-learning methodology for relational databases, from tabular datasets to full relational databases. The stakes are i) to build machine-learning models that apply readily to the raw data so as to minimize manual cleaning, data formatting and integration, and ii) to extract reusable representations that reduce sample complexity on new databases by transforming the data in well-distributed vectors and bringing background information.
3.2 Mathematical aspects of statistical learning for data science
While complex models used in machine learning can be used as non-parametric estimators for a variety of statistical tasks or for decision making, the statistical procedures and validity criterion need to be reinvented. Soda contributes statistical tools and results for a variety of problems important to data science in health and social science (epidemiology, econometrics, education). Statistical topics of interest comprise:
- Missing values and survival analysis
- Causal inference
- Model validation and auditing
- Uncertainty quantification
3.3 Machine learning for health and social sciences
Soda targets applications in health and social sciences, as these can markedly benefit from advanced processing of richer datasets, can have a large societal impact, but fall out of mainstream machine-learning research, which focus on processing natural images, language, and voice. Rather, data surveying humans needs another focus: it is most of the time tabular, sparse, with a time dimension, and missing values. In term of application fields, we focus on the social sciences that rely on quantitative predictions or analysis across individuals, such as policy evaluation. Indeed, the same formal problems, addressed in the two research axes above, arise across various social sciences: epidemiology, education research, and economics. The challenge is to develop efficient and trustworthy machine learning methodology for these high-stakes applications.
3.4 Turn-key machine-learning tools for socio-economic impact
Societal and economical impact of machine learning requires easy-to-use practical tools that can be leveraged in non-specialized organizations such as hospitals or policy-making institutions.
Soda incorporates the core team working at Inria on scikit-learn, one of the most popular machine-learning tool world-wide. One of the missions of soda is to improve scikit-learn and its documentation, transferring outside of the lab the understanding of machine learning and data science accumulated by the various research efforts.
Soda works on other important software tools to foster growth and health of the Python data ecosystem in which scikit-learn is embedded.
4 Application domains
4.1 Precision medicine, public health, and epidemiology
Data management is the focus of the field of medical informatics as it is notably challenging in healthcare settings, due to the multiplicity of sources and the richness of the data that encompasses many modalities. We apply the our machine techniques for statistical analysis, including causal inference, in medicine to facilitate clinical research and public-health evidence. The central questions are that of personalized medicine –prediction at the individual level, for diagnosis, prognosis, or drug recommendation– and of public health –evaluation of treatments and policy, estimation of risk factors. The data on which we work are patient history and claims databases: mid-dimensional data with longitudinal coverage (as opposed to “omics” or imaging data, which is high dimensional and much less frequently available in clinical settings).
We collaborate actively with AP-HP and Haute Autorité de Santé. APHP provides access to its very rich and complex data mart, with thousands of tables following millions of individuals, both a challenge and an opportunity, and we work with various medical specialists (neurology, diabetology, public health) on specific clinical questions related to prognostic, treatment evaluation, and risk factors. Haute Authorité de Santé collaborates with us by to answer public-health questions. These typically require causal inference. Finally, we are in close discussions with Institut Curie and HDH (health data hub), to start collaborations on respectively evaluation of oncology treatments using electronic health records and large language models for health databases.
4.2 Educational data mining
In educational data mining, we are interested in developing mathematical methods of learning to personalize education through adaptive assessment (developing algorithms that select questions for measuring efficiently the latent knowledge of examinees or for optimizing learning), recommending learning resources, generating exercises automatically. It is a challenging problem as it is hard to quantify learning, unlike in traditional reinforcement learning scenarios, and it is hard to measure the effect of courses on learning. This is why it is traditionally modeled as a partially-observable Markov decision process (POMDP). We are interested in modeling the evolution of uncertainty over the latent knowledge of examinees over time, for example using Bayesian approches, or model-based reinforcement learning.
Soda is actively collaborating with the national platform Pix.fr for certifying the digital competencies of all French citizens. Jill-Jênn Vie is one of the original core developers and they jointly received a Paris Region PhD grant in 2023 allowing them to co-supervise the PhD of Samuel Girard about optimizing human learning. Jill-Jênn Vie recently joined the scientific committee of the French Ministry of Education (CSEN, conseil scientifique de l'Éducation nationale), leading to collaborations with Franck Ramus and ongoing discussions with Camille Terrier, Marc Gurgand, Hugo Gimbert (MonProjetSup), and DEPP (direction de l'évaluation, de la prospective et de la performance). Through our exploratory action Orion, we are discussing potential collaborations with Louis Gleyo and Frédérique Alexandre-Bailly from ONISEP (Office national d'information sur les enseignements et les professions) to develop algorithms for orientation path personalization.
4.3 Data management
Data preparation for analytics is intrinsically related to data management. For instance, linked open data provides consistent views on data across silos, but integrating these data into a statistical model to answer a given question still requires a lot of user efforts. Database operation increasingly relies on machine learning. While Soda is in no way expert in database research, the analytic tools that we build for relational data are increasingly used for data management. We are collaborating with Paolo Papotti (Eurecom) on this topic.
4.4 Broader data science
The tools, practical and theoretical, that we develop are central to many applications of data science. For instance, we often discuss with banks and insurances, which use machine learning but face statistical problems that we tackle: censoring or other sampling biases, forecasting, uncertainty quantification. Marketing and business intelligence also face the same exact problems. Even more generally, data preparation from relational databases is a challenge is most data-science applications. We interact with data scientists in a broad set of applications via the user base of the software tools that we develop (eg scikit-learn) and the various courses and lectures that we give around these tools to industry audiences.
We have started a collaboration in economics (Margherita Comola, Paris School of Economics) on using machine learning to understanding communication strategies of politicians from social-network data.
4.5 Behavioral sciences
A methodological challenge in health and educational sciences common to behavioral science is that the quantities of interest are difficult to measure, e.g. intelligence or progress of a student. Supervised machine learning can infer proxies from indirect signs, such as psychological traits from brain imaging, diagnosis from clinical traces, or socio-economical status from demographics. This notion of proxies is central in policy evaluation, serving as indirect signals in causal inference, to provide secondary outcomes for treatment effect estimation or to control confounders not directly observed.
An ongoing project with Pass Culture (via Inria-Ministry of Culture convention) is to adapt the recommender system of the app to encourage diversity, i.e. not only optimize click-through rate, but making students discover new things. This is done by modeling this problem as contextual bandits, and a diversity term acts as regularizer in the objective function.
5 Social and environmental responsibility
5.1 Footprint of research activities
The main footprint of Soda's activity is the carbon footprint of our travels (surpassing our compute cost, as we seldom run very intensive computation). For this reason, we try to be careful with our long-distance travel and try to take the plane as little as possible. Not flying at all is not possible, as it would cut us off from the world-wide research community sometimes mediated by crucial conferences in North America. However, we favor online seminars, or on-premise talks accessible by train.
5.2 Impact of research results
While data science can improve health and education, working with personal data or providing decision tools that affect individuals comes with responsibilities.
First, overly optimistic claims, improper evaluations, or faulty statistical analysis can lead to premature usage of personal data. As collecting and handling personal data comes with privacy risks, a sober analysis of cost-benefit trade-offs is good practice. We strive to develop methodology for good evaluation 17, 20 and raise awareness of pitfalls 33, 32.
Second, Soda does not put any tools in production: none of the works of soda directly leads to automated decisions. Consequently none of our work has directly impacted individuals.
Finally, Soda works on pseudonymized data, and we leave the –pseudonymized– electronic health data on servers inside the protected environment of the hospital where they have been acquired and are used. Going further, Soda runs research on privacy-preserving synthetic data generation, to provide open datasets for research and development without privacy concerns.
6 Highlights of the year
6.1 Awards
Gaël Varoquaux highly-cited clarivate researcher
Gaël Varoquaux is on the list of the highly-cited clarivate researchers for year 2023.
7 New software, platforms, open data
7.1 New software
7.1.1 Scikit-learn
-
Keywords:
Clustering, Classification, Regression, Machine learning
-
Scientific Description:
Scikit-learn is a Python module integrating classic machine learning algorithms in the tightly-knit scientific Python world. It aims to provide simple and efficient solutions to learning problems, accessible to everybody and reusable in various contexts: machine-learning as a versatile tool for science and engineering.
-
Functional Description:
Scikit-learn can be used as a middleware for prediction tasks. For example, many web startups adapt Scikitlearn to predict buying behavior of users, provide product recommendations, detect trends or abusive behavior (fraud, spam). Scikit-learn is used to extract the structure of complex data (text, images) and classify such data with techniques relevant to the state of the art.
Easy to use, efficient and accessible to non datascience experts, Scikit-learn is an increasingly popular machine learning library in Python. In a data exploration step, the user can enter a few lines on an interactive (but non-graphical) interface and immediately sees the results of his request. Scikitlearn is a prediction engine . Scikit-learn is developed in open source, and available under the BSD license.
- URL:
- Publications:
-
Contact:
Olivier Grisel
-
Participants:
Alexandre Gramfort, Bertrand Thirion, Gael Varoquaux, Loic Esteve, Olivier Grisel, Guillaume Lemaitre, Jeremie Du Boisberranger, Julien Jerphanion
-
Partners:
Axa, BNP Parisbas Cardif, Dataiku, Nvidia, Chanel
7.1.2 joblib
-
Keywords:
Parallel computing, Cache
-
Functional Description:
Facilitate parallel computing and caching in Python.
- URL:
-
Contact:
Gael Varoquaux
7.1.3 skrub
-
Keyword:
Machine learning
-
Functional Description:
Joins, aggregates, and vectorizes tables to enable statistical learning, including with badly formated entries
- URL:
-
Contact:
Gael Varoquaux
7.2 Open data
KEN: Relational Data Embeddings for Feature Enrichment
For data science on common entities, such as cities, companies, famous people... augmenting the data at hand with information assembled from external sources may be key. For instance, estimating housing prices benefits from background information on the location: the population density, the average income... This information is present in external sources such as wikipedia. But assembling features that summarize the information is tedious manual work, which we seek to replace.
We provide readily-computed vectorial representations of 5.7 millions entities (e.g. cities) that capture the information that can be aggregated across wikipedia.
https://soda-inria.github.io/ken_embeddings
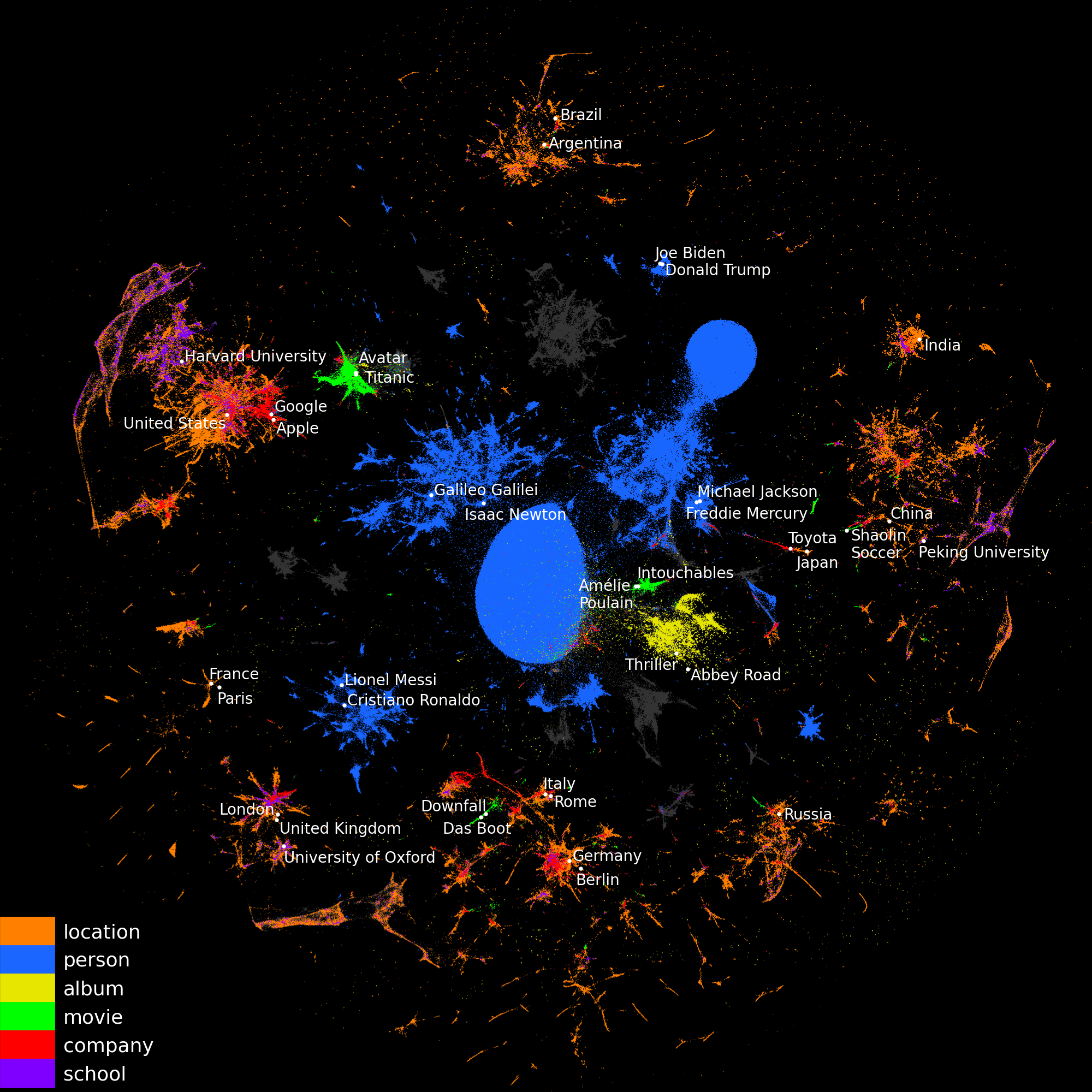
2D visualization of entity embeddings, colored by their types. UMAP was used to reduce their dimension from 200 to 2.
8 New results
8.1 Table representation learning
Participants: Gael Varoquaux.
Acronym Disambiguation: benchmark and model
14 Acronym Disambiguation (AD) is crucial for natural language understanding on various sources, including biomedical reports, scientific papers, and search engine queries. However, existing acronym disambiguation benchmarks and tools are limited to specific domains, and the size of prior benchmarks is rather small. To accelerate the research on acronym disambiguation, we construct a new benchmark named GLADIS with three components: (1) a much larger acronym dictionary with 1.5M acronyms and 6.4M long forms; (2) a pre-training corpus with 160 million sentences; (3) three datasets that cover the general, scientific, and biomedical domains. We then pre-train a language model, AcroBERT, on our constructed corpus for general acronym disambiguation, and show the challenges and values of our new benchmark.
The structure of positional encodings in language models
15 Positional Encodings (PEs) are used to inject word-order information into transformer-based language models. They are needed from transformers to model sequences rather than sets of words, and they significantly enhance the quality of sentence representations. However, the embeddings capture specific inductive biases and the corresponding contribution to language models is not fully understood. Indeed recent findings highlight that various positional encodings are insensitive to word order. We conducted a systematic study of positional encodings in Bidirectional Masked Language Models (BERT-style). This study revealed two common properties, Locality and Symmetry, core to the function of PEs, that vary across the encodings used in practice (figure 2). We showed that these two properties are closely correlated with the performances of downstream tasks (figure 3). We quantified the weakness of current PEs by introducing two new probing tasks, on which current PEs perform poorly. We believe that these results are the basis for developing better PEs for transformer-based language models and that they explain the choice of local attention structure in the recent language model Mistral 7B.
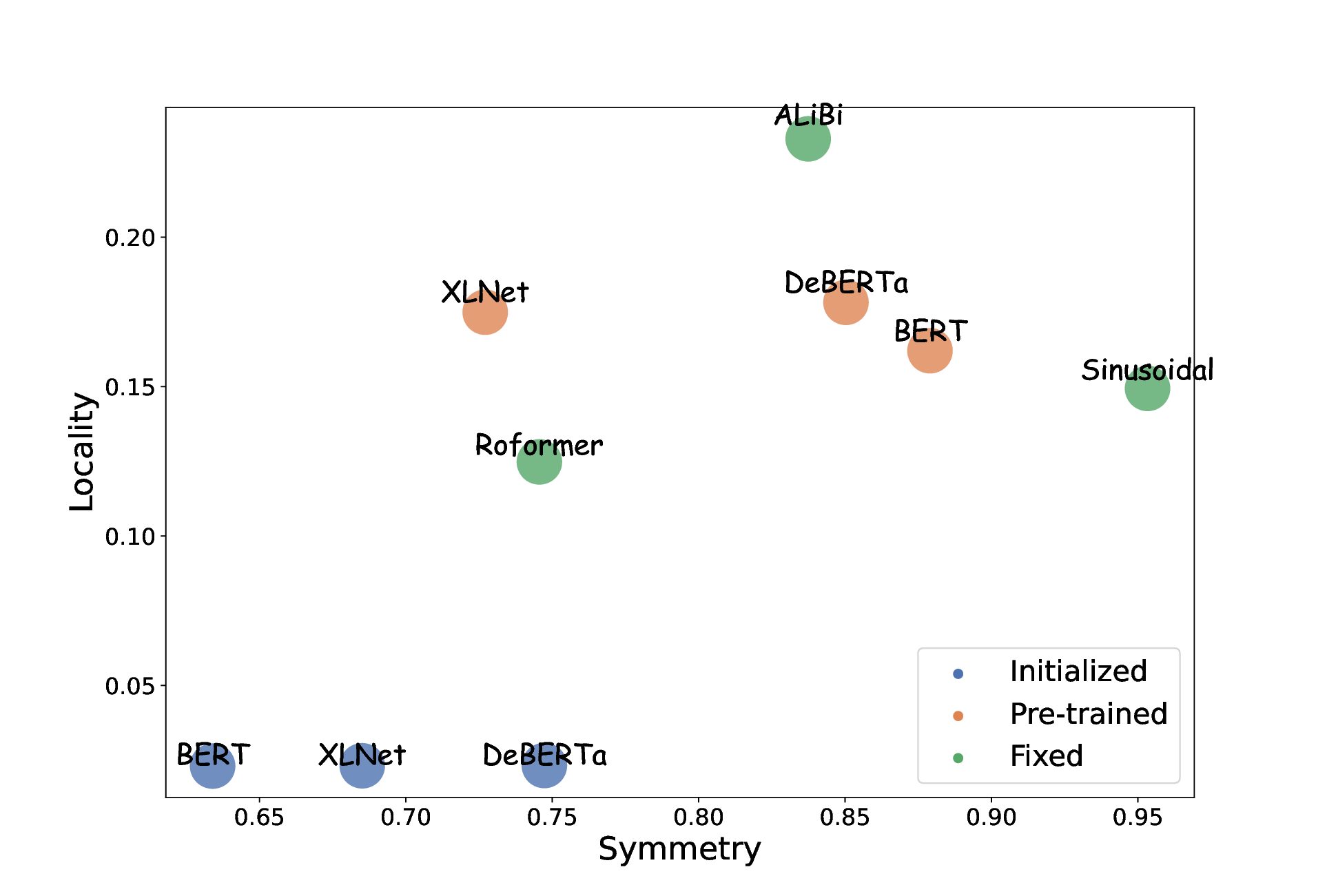
Locality and symmetry values of positional encodings
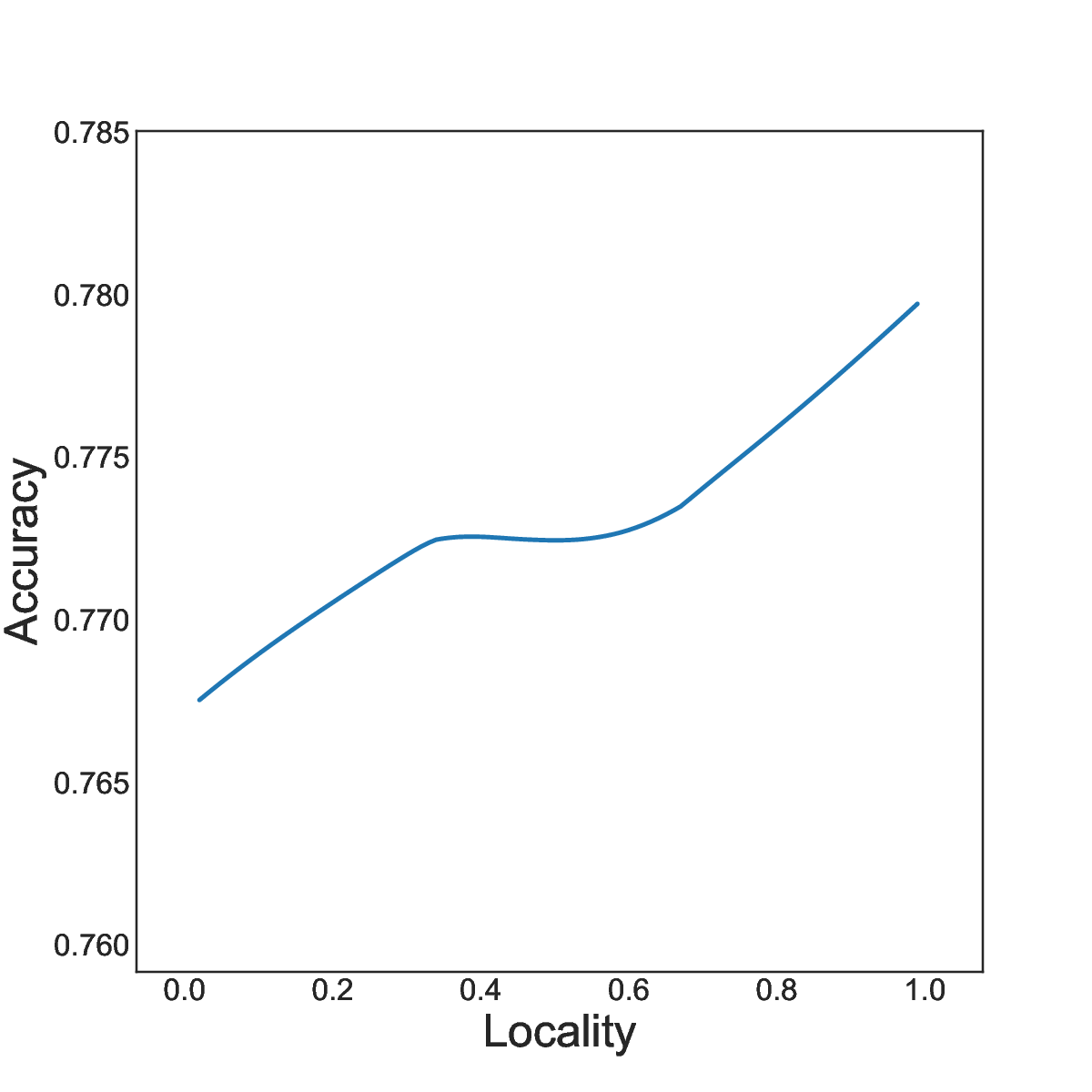
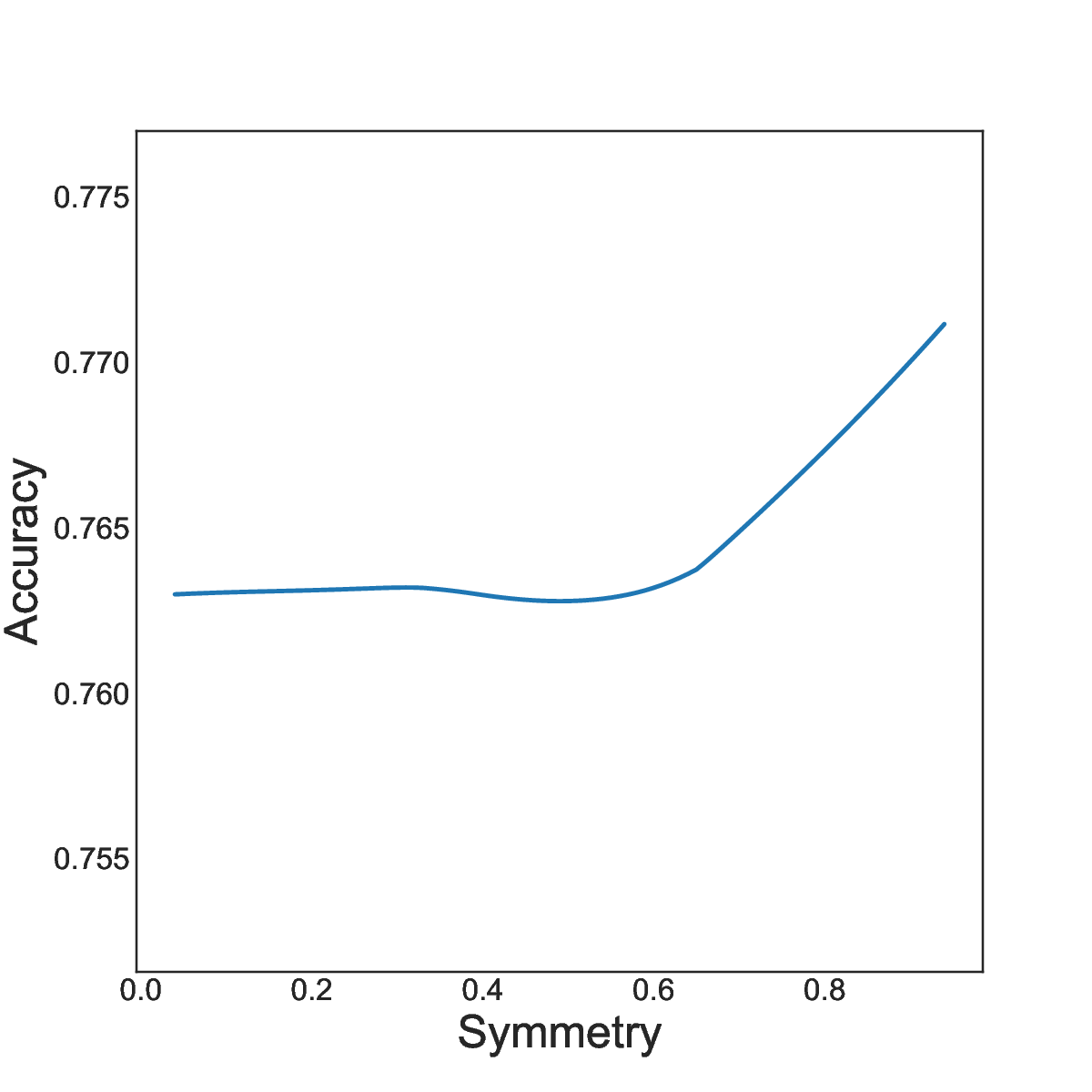
Study of locality and symmetry
Study of locality and symmetry
8.2 Mathematical aspects of statistical learning for data science
Participants: Marine Le Morvan, Gael Varoquaux.
Risk ratio, odds ratio, risk difference... Which causal measure is easier to generalize?
1 There are many measures to report so-called treatment or causal effect: absolute difference, ratio, odds ratio, number needed to treat, and so on. The choice of a measure, eg absolute versus relative, is often debated because it leads to different appreciations of the same phenomenon; but it also implies different heterogeneity of treatment effect. In addition some measures – but not all – have appealing properties such as collapsibility, matching the intuition of a population summary. We review common measures and their pros and cons typically brought forward. Doing so, we clarify notions of collapsibility and treatment effect heterogeneity, unifying different existing definitions. Our main contribution is to propose to reverse the thinking: rather than starting from the measure, we start from a non-parametric generative model of the outcome. Depending on the nature of the outcome, some causal measures disentangle treatment modulations from baseline risk. Therefore, our analysis outlines an understanding what heterogeneity and homogeneity of treatment effect mean, not through the lens of the measure, but through the lens of the covariates. Our goal is the generalization of causal measures. We show that different sets of covariates are needed to generalize an effect to a different target population depending on (i) the causal measure of interest, (ii) the nature of the outcome, and (iii) the generalization's method itself (generalizing either conditional outcome or local effects).
Validating probabilistic classifiers: beyond calibration
3 Ensuring that a classifier gives reliable confidence scores is essential for informed decision-making. For instance, before using a clinical prognostic model, we want to establish that for a given individual is attributes probabilities of different clinical outcomes that can be indeed trusted. To this end, recent work has focused on miscalibration, i.e., the over or under confidence of model scores. Yet calibration is not enough: even a perfectly calibrated classifier with the best possible accuracy can have confidence scores that are far from the true posterior probabilities, if it is over-confident for some samples and under-confident for others. This is captured by the grouping loss, created by samples with the same confidence scores but different true posterior probabilities. Proper scoring rule theory shows that given the calibration loss, the missing piece to characterize individual errors is the grouping loss. While there are many estimators of the calibration loss, none exists for the grouping loss in standard settings. We propose an estimator to approximate the grouping loss 17. We show that modern neural network architectures in vision and NLP exhibit grouping loss, notably in distribution shifts settings, which highlights the importance of pre-production validation.
8.3 Machine learning for health and social sciences
Participants: Gael Varoquaux, Judith Abécassis, Jill-Jênn Vie.
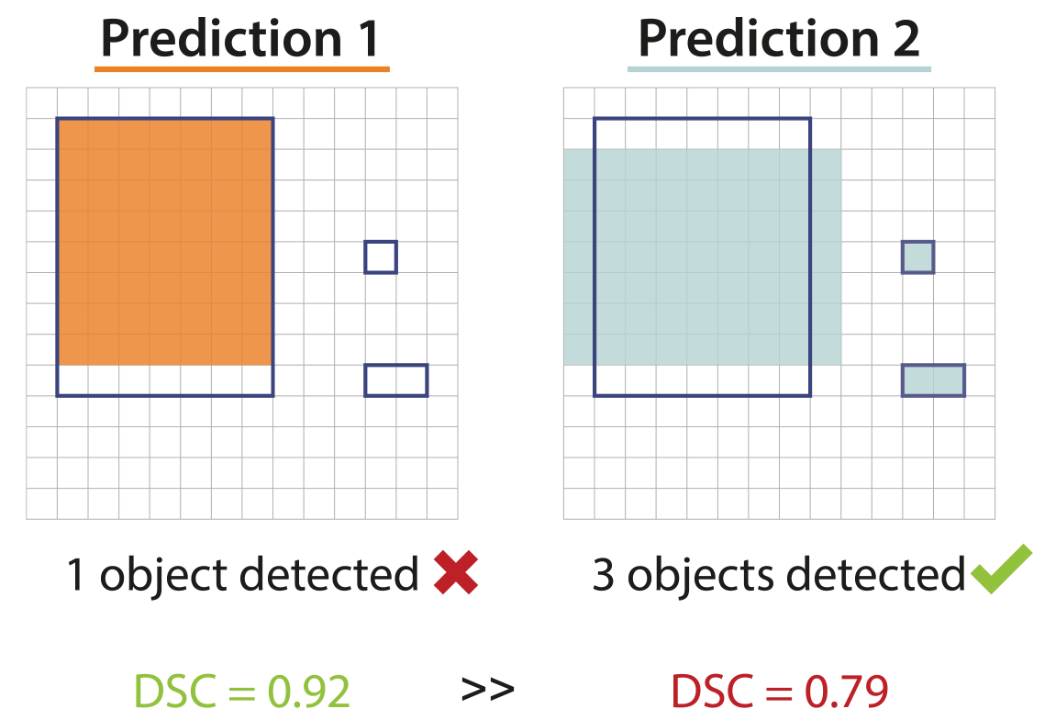
1 object detected, versus 3 objects detected
Understanding metric-related pitfalls in image analysis validation
4 Validation metrics are key for the reliable tracking of scientific progress and for bridging the current chasm between artificial intelligence (AI) research and its translation into practice. However, increasing evidence shows that particularly in image analysis, metrics are often chosen inadequately in relation to the underlying research problem. This could be attributed to a lack of accessibility of metric-related knowledge: While taking into account the individual strengths, weaknesses, and limitations of validation metrics is a critical prerequisite to making educated choices, the relevant knowledge is currently scattered and poorly accessible to individual researchers. Based on a multi-stage Delphi process conducted by a multidisciplinary expert consortium as well as extensive community feedback, we provided the first reliable and comprehensive common point of access to information on pitfalls related to validation metrics in image analysis. Focusing on biomedical image analysis but with the potential of transfer to other fields, the addressed pitfalls generalize across application domains and are categorized according to a newly created, domain-agnostic taxonomy. To facilitate comprehension, illustrations and specific examples accompany each pitfall. As a structured body of information accessible to researchers of all levels of expertise, this work enhances global comprehension of a key topic in image analysis validation.
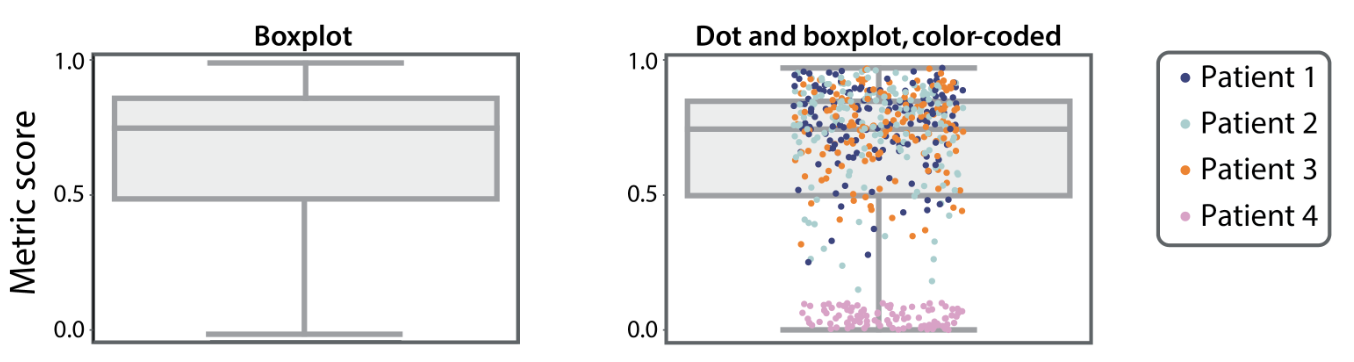
Heterogeneity in the scores may be invisible in a box plot. Adding the relevant stratus information on a scatter plot may reveal it.
Synchronous female bilateral breast cancers
2 Synchronous bilateral breast cancer is a very rare situation where tumors in both breasts are detected in a patient. This configuration provides insights to understand the relationships among tumor, host (shared for the two tumors), immunity and response to treatment. However, little evidence exists regarding immune infiltration and response to treatment in sBBCs. With Institut Curie we analysed the health records of 404 patients with sBBCs (fig:cancerflowchart), out of 17,575 female patients with non-metastatic breast cancer between 2005 and 2015. We showed that the impact of the subtype of breast cancer on levels of tumor infiltrating lymphocytes (TIL) and on pathologic complete response rates differs according to the concordant or discordant subtype of breast cancer of the contralateral tumor: luminal breast tumors with a discordant contralateral tumor had higher TIL levels and higher pCR rates than those with a concordant contralateral tumor. Our study indicates that tumor-intrinsic characteristics may have a role in the association of tumor immunity and pCR and demonstrates that the characteristics of the contralateral tumor are also associated with immune infiltration and response to treatment.
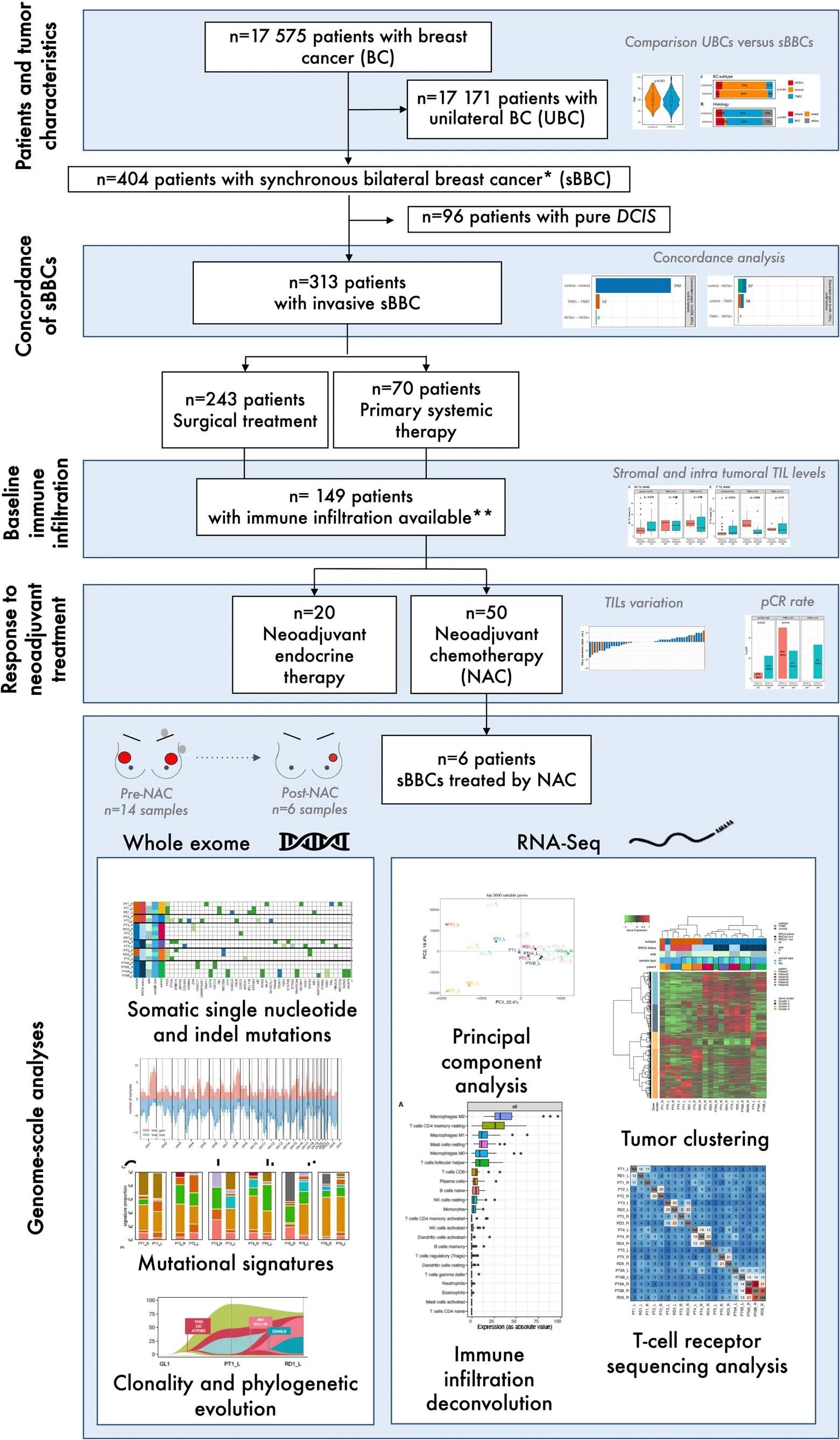
Flowchart representing the multi-source analysis of a cohort of Synchronous bilateral breast cancer patients
Additionally, on a subset of patients with available frozen tissue, tumor sequencing revealed that left and right tumors were independent regarding somatic mutations, copy number alterations and clonal phylogeny, whereas primary tumor and residual disease were closely related both from the somatic mutation and from the transcriptomic point of view.
Learning path personalization as recommending nodes on a bipartite graph
5 Adaptive learning is an area of educational technology that consists in delivering personalized learning experiences to address the unique needs of each learner. An important subfield of adaptive learning is learning path personalization: it aims at designing systems that recommend sequences of educational activities to maximize students' learning outcomes. In this work we framed learning path personalization as recommendation of nodes on a bipartite graph of keywords to documents and learned a policy for recommending documents based on prior user feedback, using reinforcement learning. Our model is based on a graph neural network, as those can be trained on some graphs and be reused on new graphs, no matter the number of nodes, making it a scalable approach as new documents go. We evaluated on simulated data, and showed good results compared to a baseline, even in the low data regime.
8.4 Turn-key machine-learning tools for socio-economic impact
Participants: Jeremie Du Boisberranger, Olivier Grisel, Guillaume Lemaitre, Gael Varoquaux.
New release of scikit-learn
Scikit-learn is always improving, adding features for better and easier machine learning in Python. We list below a few highlights that are certainly not exhaustive but illustrate the continuous progress made.
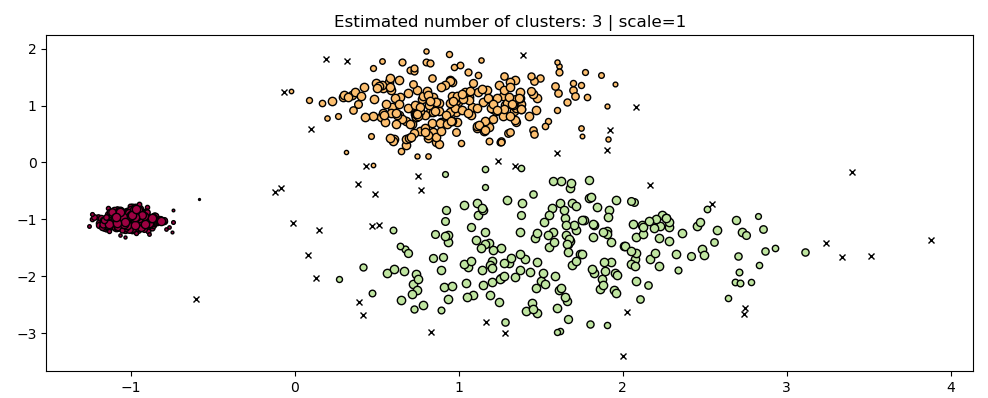
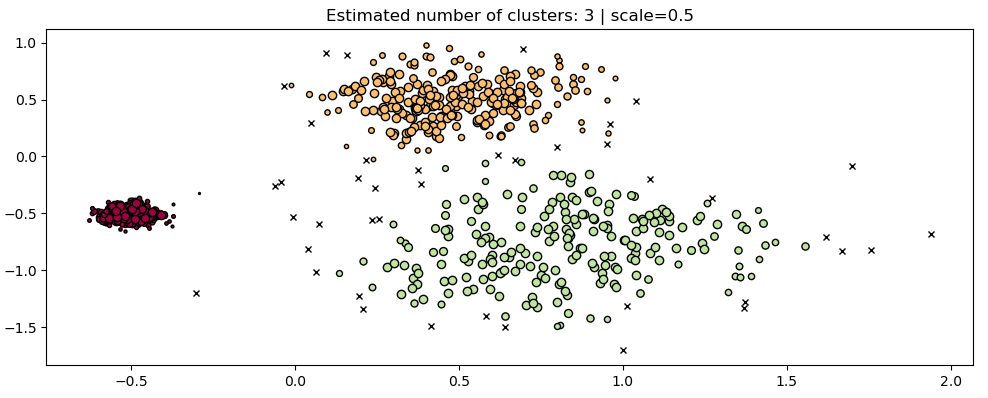
Two visualization of clusters estimated with the HDBScan algorithm, the figure on the right is estimated on data with a scale twice as small as on the left
Two visualization of clusters estimated with the HDBScan algorithm, the figure on the right is estimated on data with a scale twice as small as on the left
Release 1.3 (June 2023), with a large number of changes; the most notable ones are:
- Addition of HDBSCAN, a modern hierarchical density-based clustering algorithm. Similarly to OPTICS, it can be seen as a generalization of DBSCAN by allowing for hierarchical instead of flat clustering, however it varies in its approach from OPTICS. This algorithm is very robust with respect to its hyperparameters' values and can be used on a wide variety of data without much, if any, tuning.
- Addition of TargetEncoder which is a categorical encoding based on target mean conditioned on the value of the category.
- Decision trees now natively handle missing values.
- Added the class ValidationCurveDisplay that allows easy plotting of validation curves
- The gradient-boosting models now support the Gamma-deviance loss function, useful for modeling strictly positive targets with a right-skewed distribution
- Similarly to OneHotEncoder, the OrdinalEncoder now supports aggregating infrequent categories into a single output for each feature.
skrub
skrub is a much younger package that strives to facilitate statistical learning on relational data, even when it is messy. Skrub is an evolution of the dirty-cat package, giving it a broader scope.
Release 0.1 (Dec 2023)
- The TableVectorizer (to vectorize a table)
suitable for automatic usage:
- automatic casting of types in transform,
- avoid dimensionality explosion when a feature has two unique values, by using a OneHotEncoder that drops one of the two vectors.
- transform can now return features, without modification.
- DatetimeEncoder can transform a datetime column into several numerical columns (year, month, day, hour, minute, second, ...). It is now the default transformer used in the TableVectorizer for datetime columns.
- AggJoiner: performs a join on an external table and aggregates the results in case of a one-to-many match. Useful for assembling features across multiple tables for learning.
joblib
joblib is a very simple computation engine in Python that is massively used worldwide, including as a dependency of packages such as scikit-learn for parallel computing.
Release 1.3 (August 2023). Many changes to follow evolutions of the ecosystem and improve behaviors (eg better error handling). Major changes are:
- Add limits on age and number of items on the cache
- Parallel computing can now return results asynchronously and dynamically map-reduce like behavior to decrease memory usage
9 Bilateral contracts and grants with industry
Participants: Gael Varoquaux.
9.1 Bilateral contracts with industry
scikit-learn consortium
scikit-learn is development is funded via a consortium with industry partners hosted by the Inria foundation. The consortium currently comprises hugginface, dataiku, BNP-Parisbas-cardiff, AXA, and Channel which joined in 2023. It has an operating budget of around 250 k€ yearly, and employs 5 engineers. Priorities are set by a bi-annual technical committee where industry partners sit together with community members in a joint discussion. Gaël Varoquaux is in charge at Soda.
Intel donation
Intel has partnered with Soda in the context of its “oneAPI centers of excellence”. The donation, an amount of 100 k€ yearly for two years (2022 and 2023), aims to establish a high-performance computational back-end to speed up scikit-learn on GPUs using SYCL. Different backends are explored: a plugin external to the scikit-learn codebase to extend it or the use of the array API to run computation on a dedicated array-computing engine. Gaël Varoquaux is in charge at Soda.
Pass Culture
Within the Ministry of Culture-Inria convention, Samuel Girard and Jill-Jênn Vie have been involved in a short-term (4 months, 18k) and long-term (12 months, 77k, will start in 2024, already funded in 2023) partnership to improve the diversity of recommendations in Pass Culture (used by 3M students in France). We are in discussions to hire an engineer from April 2024.
Collaboration with Haute Autorité de Santé
We had a 3-year long collaboration with Haute Autorité de Santé (HAS) on using health data mart for policy evaluation, which finished at the end of 2023. The collaboration was mediated by Matthieu Doutreligne –part time HAS, part time Soda– and came with a budget of 50 k€. Gaël Varoquaux is in charge at Soda.
9.2 Bilateral Grants with Industry
Collaboration with public interest group Pix
Jill-Jênn Vie applied to Paris Region PhD 2023 with Pix (certification of digital competencies, 6M active users), about optimizing human learning. Samuel Girard PhD is on this funding (105k from région Île-de-France, 20k from Pix).
Plan de relance with Dataiku
– Soda has a 24 months post-doc funded by “plan de relance” jointly with Dataiku on using embeddings for database analytics. The post-doc started beginning of November 2022. Gaël Varoquaux is in charge at Soda.
10 Partnerships and cooperations
10.1 International initiatives
10.1.1 Inria associate team not involved in an IIL or an international program
Participants: Gaël Varoquaux, Jill-Jenn Vie.
OPALE
-
Title:
Optimal policy active learning for education
-
Duration:
2021 -> 2023
-
Coordinator:
Koh Takeuchi (takeuchi@i.kyoto-u.ac.jp)
-
Partners:
- Kyoto University (Japan)
-
Inria contact:
Jill-Jênn Vie
-
Summary:
Our research project is to explore reinforcement learning and causal inference to learn policies for collecting student data, so that we can understand how the students learn, and which lessons/exercises in a course have a strong impact on learning for which students. This has benefits both for the students that can reflect on their knowledge, and the teachers that can receive feedback and recommendations to improve their teaching: for example, if the teacher hesitates between several learning resources to master a topic, our algorithm would be able to quantify which lesson is most suitable for a given student. We plan to implement our algorithms in actual platforms to try the learned policies on real students, with public and private partnerships (for example Pix, a project initially based at the French Ministry of Education, which is now the public standardized test for certifying digital competencies in France).
10.2 International research visitors
10.2.1 Visits of international scientists
Participants: Clémence Réda.
Other international visits to the team
Clémence Réda
-
Status
Post-Doc
-
Institution of origin:
Rostock Universität
-
Country:
Germany
-
Dates:
July–October 2023 (4 months)
- Context of the visit:
-
Mobility program/type of mobility:
research stay
10.2.2 Visits to international teams
Participants: Alexandre Perez.
Research stays abroad
Alexandre Perez-Level
-
Visited institution:
Stanford
-
Country:
USA
-
Dates:
sept 1st – dec 20th 2023
-
Context of the visit:
Research stay hosted by Professor Kojevo of the Computer-Science department
-
Mobility program/type of mobility:
research stay
10.3 European initiatives
10.3.1 Horizon Europe
Intercept T2D
Intercept is a collaborative research project on transition to Type-2 diabetes and risk factors of health degradation. The partners comprise several hospitals across Europe, as well as INSERM in France for biochemistry and genomics. Soda is involved in close collaboration with APHP on work package 5 to establish epidemiological evidence from routine-care data. In particular, the goals are to establish risk factors of diabetes complications as well as to study the causal effect of inflammation and inflammation-reducing drug on long-term patient health. The funding for Soda is 700 k€ and the project started in 2023. Gaël Varoquaux is in charge at Soda, lead of work package 5.
10.3.2 Other european programs/initiatives
Jill-Jênn Vie submitted a ERC named OTHeLLO: Optimal Transport of Human Learners to their Learning Objectives.
Jill-Jênn Vie has been in the advisory board of the 2021–2024 Erasmus+ AI4T project (ID: 626154-EPP-1-2020-2-FR-EPPKA3-PI-POLICY).
10.4 National initiatives
PEPR Santé Numérique
Soda is part of the “PEPR Santé Numérique” in the SMATCH subgroup that focuses on evidence of clinical efficacy. Soda will address two questions. The first question, addressed in collaboration with the PreMedical team, is that of external validity of randomized trials: how much is the treatment effect measured in a randomized clinical trial affected by the sampling bias of the trial, the difference between the study population and the intended target population. The second question, addressed in collaboration with the Heka team, is that of defining guidelines to evaluate software as a medical device. One particular challenge that we will tackle is to give procedures and recommendations to evaluate an update to a software used in clinical decision making using historical data rather than a trial. The project started end of 2023. Gaël Varoquaux is in charge at Soda.
10.5 Regional initiatives
DataIA funding: data wrangling software
The DataIA institute is funding an engineer for 2 years to work on data-wrangling software. A first version of the corresponding software project, named skrub has been released: (website). Gaël Varoquaux is in charge at Soda.
11 Dissemination
11.1 Promoting scientific activities
11.1.1 Scientific events: organisation
Member of the organizing committees
-
Gaël Varoquaux
Organizer of the 2nd “Table Relational Learning” workshop, at NeurIPS (website)
11.1.2 Scientific events: selection
Member of the conference program committees
-
Gaël Varoquaux
Area chair: NeurIPS, NeurIPS datasets and benchmarks, ICLR, ICML
-
Jill-Jênn Vie
Workshop chair: PAKDD 2023
Publicity chair: EDM 2023, EDM 2024
Reviewer
-
Gaël Varoquaux
AISTATS, IJCAI, NeurIPS workshop selection
-
Marine Le Morvan
ICML
-
Judith Abécassis
NeurIPS, ICLR
-
Jill-Jênn Vie
NeurIPS, LAK, PAKDD, ICCE, AI4ED workshop at AAAI
11.1.3 Journal
Reviewer - reviewing activities
-
Gaël Varoquaux
JMLR
11.1.4 Invited talks
-
Gaël Varoquaux
Keynote: IDA (Intelligent Data Analysis, Louvain la Neuve), JDSE (Orsay), Journée Française d'IA en imagerie biologique (Paris), RSE days (Swansea)
-
Marine Le Morvan
-
Young Statisticians and Probabilists (YSP) Days, Paris, France, January 2023
Introduction to missing values.
-
PyLadies Meetup, Paris, France, April 2023
Learning with missing values: theoretical insights and application to health databases.
-
-
Jill-Jênn Vie
-
Journée Groupe thématique numérique Intelligence Artificielle et Éducation Ouverte, Nantes, janvier 2023
Fairness et confidentialité en IA pour l'éducation : risques et opportunités
-
Atelier La diversité de la recherche en EIAH par l'exemple, conférence Environnements informatiques pour l'apprentissage humain, Brest, juin 2023
Traçage des connaissances et optimisation de l'apprentissage humain
-
Journées scientifiques Inria, Bordeaux, septembre 2023
Apprendre de données d'humains : applications du crowdsourcing et de l'inférence causale pour l'éducation
-
Cercle Ekitia – Les données synthétiques : encadrement et enjeux des techniques et des usages, en ligne, novembre 2023
Techniques de génération de données synthétiques, pertinence et risques de leur utilisation
-
-
Bénédicte Colnet
-
Paris Women in Machine Learning & Data Science, Paris, France, February 2023
Reweighting the RCT for generalization: finite sample error and variable selection
-
International Online Causal Inference Seminar, May 2023
Risk ratio, odds ratio, risk difference... Which measure is easier to generalize?
-
PyLadies Meetup, Paris, France, April 2023
Machine learning and causal inference: Toward new clinical evidence?
-
Bits in Bio Meetup, November 2023, Paris, France
Combining randomized and observational data: Toward new clinical evidence?
-
11.1.5 Leadership within the scientific community
-
Jill-Jênn Vie
Secrétaire de la Société informatique de France (SIF) jusqu'au 1er février 2023
11.1.6 Scientific expertise
-
Gaël Varoquaux
Member of the National Committee of experts on artificial intelligence (Comité IA) www.gouvernement.fr/communique/comite-de-lintelligence-artificielle
-
Jill-Jênn Vie
Member of the Scientific Committee of the French Ministry of Education (CSEN)
Scientific committe of Inria project Regalia
Advisory board of the Erasmus+ AI4T project
11.1.7 Research administration
-
Gaël Varoquaux
Stanford Open Source Center (OSPO) Advisory Board Member
11.2 Teaching - Supervision - Juries
Courses
-
Gaël Varoquaux
- Machine learning for social sciences, EHESS, 8h
- Real-world data science = messy data, Nepal Machine Learning Summer School, 1.5h, Katmandu (remote talk)
- Learning on messy tabular data, Hi-Paris Summer School, 1.5h
- Machine learning for data science with scikit-learn, African Institute for Mathematical Science, 3h, Kigali
- Machine learning, data science, inria academie, 1.5h
-
Marine Le Morvan
- Advanced Machine Learning, Ecole Polytechnique, 21h
- Refresher Course in Artificial Intelligence, Ecole Polytechnique, 15h
- Statistics and machine learning with missing values, Université Paris Dauphine, 6h
- Learning with trees, Ecole Polytechnique Executive education, 7h
-
Olivier Grisel
- Deep Learning (M2), Université Bretagne Sud, 21h
- Machine Learning (M1), Université Bretagne Sud, 21h
-
Jill-Jênn Vie
- Deep learning: do it yourself, ENS, 7.5h eq. TD
- INF471S ICPC–SWERC training (advanced algorithms), École polytechnique, 60h
- PDV302 ICPC–SWERC training (advanced algorithms), Bachelor École polytechnique, 24h
- Préparation au SWERC, ENS Paris-Saclay, 39h eq. TD
- CSE301 Introduction to Haskell, Bachelor École polytechnique, 1h30
E-learning
-
Machine learning with Scikit-learn MOOC
40 hours of learning starting as an introduction to machine learning and covering more advanced topics such as data preparation and model selection. Accessible on inria.github.io/scikit-learn-mooc, and designed by Loïc Esteve , Arturo Amor , Guillaume Lemaître , Olivier Grisel , Gaël Varoquaux .
11.2.1 Supervision
-
Gaël Varoquaux
Supervising the following PhD students: Julie Alberge, Lihu Chen, Bénédicte Colnet, Alexis Cvetkov-Iliev, Matthieu Doutreligne, Alexandre Perez, Jovan Stojanovic.
Also supervising undergraduate intern Anand-Arnaud Pajaniradjane and apprentice master Lilian Boulard.
-
Marine Le Morvan
Supervising Alexandre Perez (PhD student).
-
Jill-Jênn Vie
Supervising Samuel Girard and Jean Vassoyan (PhD students), Tomas Rigaux (engineer).
-
Judith Abecassis
Supervising Julie Alberge (PhD student) and Sami Boumaiza (intern).
-
Bénédicte Colnet
Supervising Nour Oueghlani (intern).
11.2.2 Juries
-
Gaël Varoquaux
was “examinateur” for the defense of Romain Peressoni (Bordeaux).
Comité de suivi de thèse: Lawrence Stewart (Paris), Alan Balendran (Paris)
-
Jill-Jênn Vie
Jury of the agrégation d'informatique, June, Paris.
Jury of the PhD defense of Olivier Allègre (Paris)
Comité de thèse of Mélina Verger (Paris), Erva Nihan Kandemir (Paris).
11.3 Popularization
11.3.1 Articles and contents
-
Gaël Varoquaux
- Podcast (1.5 hour): I, scientist Youtube link
- Podcast (45mn): Café Data Youtube link
- Podcast (1.5 hour): Super Data science Youtube link
11.3.2 Education
-
Gaël Varoquaux
- Course (1.5 hour, on machine learning for tabular data) at AI for Ukraine (website)
- Talk (1 hour): introduction to machine learning for the FHU Neurovasculaire
- Talk (30 hour): introduction to AI for health and society to executives of the “ministère de l'intérieur”.
-
Judith Abécassis
- Conference for Junior High School students - Week of Mathematics (March 2023)
- Introduction to causal inference (1 day course) - L'Oréal (June 2023)
- Masterclass IA et santé - journée Santé Inria (December 2023)
11.3.3 Interventions
-
Judith Abécassis
- Atelier-Rencontre Inria x France Biotech - Onco (May 2023)
- Intervention Chiche, Lycée Hélène Boucher, May 2023
-
Jill-Jênn Vie
Intervention Chiche Algorithmes de recommandation comment ça marche ?, Lycée international de Palaiseau, novembre 2023
-
Samuel Girard
Intervention Chiche, Inria Rocquencourt, December 2023
-
Bénédicte Colnet
Intervention Chiche
12 Scientific production
12.1 Major publications
- 1 misc Risk ratio, odds ratio, risk difference... Which causal measure is easier to generalize? 2023 HAL back to text
- 2 articleEvolution of synchronous female bilateral breast cancers and response to treatment.Nature MedicineMarch 2023HALDOIback to text
- 3 inproceedingsBeyond calibration: estimating the grouping loss of modern neural networks.ICLR ProceedingsICLR 2023 – The Eleventh International Conference on Learning RepresentationsKigali, Rwanda2023HALback to text
- 4 miscUnderstanding metric-related pitfalls in image analysis validation.December 2023HALback to text
- 5 inproceedingsTowards Scalable Adaptive Learning with Graph Neural Networks and Reinforcement Learning.EDM 2023 - 16th International Conference on Educational Data MiningBangalore, IndiaMay 2023HALback to text
12.2 Publications of the year
International journals
International peer-reviewed conferences
Scientific book chapters
Doctoral dissertations and habilitation theses
Reports & preprints
