Keywords
Computer Science and Digital Science
- A6.1.1. Continuous Modeling (PDE, ODE)
- A6.1.2. Stochastic Modeling
- A6.1.4. Multiscale modeling
- A6.2.1. Numerical analysis of PDE and ODE
- A6.2.6. Optimization
- A6.2.7. High performance computing
- A6.2.8. Computational geometry and meshes
- A6.3.1. Inverse problems
- A6.3.2. Data assimilation
- A6.3.3. Data processing
- A6.3.5. Uncertainty Quantification
Other Research Topics and Application Domains
- B1.1.2. Molecular and cellular biology
- B1.1.8. Mathematical biology
- B2.2.1. Cardiovascular and respiratory diseases
- B2.2.6. Neurodegenerative diseases
- B2.4.1. Pharmaco kinetics and dynamics
- B2.4.3. Surgery
- B2.6.1. Brain imaging
- B2.6.2. Cardiac imaging
1 Team members, visitors, external collaborators
Research Scientists
- Jacques Henry [INRIA, Emeritus, HDR]
- Peter Langfield [INRIA, Researcher]
- Michael Leguebe [INRIA, Researcher]
- Nejib Zemzemi [INRIA, Researcher]
Faculty Members
- Yves Coudière [Team leader, UNIV BORDEAUX, Professor, until Aug 2022, HDR]
- Mostafa Bendahmane [UNIV BORDEAUX, Associate Professor, HDR]
- Mark Potse [UNIV BORDEAUX]
- Lisl Weynans [UNIV BORDEAUX, Associate Professor, HDR]
Post-Doctoral Fellows
- Andony Arrieula [UNIV BORDEAUX, from Nov 2022]
- Mohamadou Malal Diallo [UNIV BORDEAUX, until Aug 2022]
- Laetitia Mottet [UNIV BORDEAUX, from Nov 2022]
PhD Students
- Andony Arrieula [Inria, until Oct 2022, Supervizor : Mark Potse]
- Simon Bihoreau [Inria, from Dec 2022, In MONC team, but cosupervized (Michael Leguèbe) in CARMEN]
- Oumayma Bouhamama [BORDEAUX INP, ATER, until Aug 2022, Supervizor : Lisl Weynans]
- Hassaan A. Bukhari [UNIV SARAGOSSE, Supervizor: Mark Potse]
- Zeina Chehade [UNIV BORDEAUX, Supervizor: Yves Coudière]
- Narimane Gassa [UNIV BORDEAUX, Supervizors : Nejib Zemzemi, Yves Coudière]
- Emma Lagracie [UNIV BORDEAUX, from Oct 2022, Supervizors: Lisl Weynans, Yves Coudière]
- Niami Nasr [UNIV BORDEAUX, Supervizor: Lisl Weynans]
- Valentin Pannetier [UNIV BORDEAUX, Supervizors: Yves Coudière, Michael Leguèbe]
- Bachar Tarraf [UNIV BORDEAUX, ATER, until Mar 2022, Supervizors: Yves Coudière, Michael Leguèbe]
Technical Staff
- Pauline Migerditichan [UNIV BORDEAUX, until Mar 2022]
- Corentin Prigent [UNIV BORDEAUX, Engineer, from Oct 2022]
- Chiheb Sakka [Inria, until Jul 2022, In STORM team]
Interns and Apprentices
- Victor Bousch [EINSERB-MATMECA, from Jun 2022 until Aug 2022]
- Léo Denis [UNIV LA ROCHELLE, from Mar 2022 until Aug 2022]
- Emma Lagracie [UMA-ENSTA, from Apr 2022 until Sep 2022]
Administrative Assistant
- Nathalie Robin [INRIA]
Visiting Scientists
- Hamza Ammar [LAMSIN, from May 2022 until Jun 2022, (Spicy Associated Team)]
- Abir Amri [LAMSIN, from Jun 2022 until Jun 2022, (PHC Utique, S. Ervedoza, IMB)]
- Carlos Fambuena Santos [UNIV POLYTECNICA VALENCIA, from May 2022 until Jul 2022, (PhD secondment from PersonalizeAF)]
- Khouloud Khordoghli [UNIV TUNIS, from May 2022 until Jun 2022, (Spicy Associated Team)]
- Beata Ondrusova [IMS - SLOVAK ACADEMY OF SCIENCES, from Oct 2022, (ECGI Consortium)]
- Jana Svehlikova [IMS - SLOVAK ACADEMY OF SCIENCES, from Oct 2022, (ECGI Consortium)]
External Collaborator
- Pierre-Elliott Bécue [ANSSI]
2 Overall objectives
The Carmen team develops and uses models and numerical methods to simulate the electrophysiology of the heart from the molecular to the whole-organ scale, and its relation to measurable signals inside the heart and on the body surface. It aims at:
- improving understanding of normal and pathological cardiac electrophysiology,
- improving the efficiency and accuracy of numerical models,
- exploiting all available electrical signals for diagnosis,
- improving understanding and guidance of ablative treatment of cardiac arrhythmia.
The numerical models developed, analyzed, and used by the team incorporate essentially the gating dynamics of the ion channels in the cardiac cell membranes and the heterogeneities of the cardiac tissue, coupling processes on the cellular scale into macroscopic reaction-diffusion models. The team also work on incorporating any new biological knowledge, at any scale, that helps to understand the mechanisms of arrythmias, their diagnosis or treatment. At the same time we use simpler or reduced models to solve the inverse problems related to non-invasive electrical imaging of the heart.
The fields involved in our research are: ordinary and partial differential equations (ODE & PDE), inverse problems, numerical analysis, high-performance computing, image segmentation, and mesh construction.
A main goal of the team is to contribute to the work packages defined in the project of IHU Liryc, an institute founded in 2011 that focuses on cardiac arrhythmia.
We cooperate with physiologists and cardiologists on several projects. The team is building new models and powerful simulation tools that will help to understand the mechanisms behind cardiac arrhythmias and to establish personalized and optimized treatments. A particular challenge consists in making the simulations reliable and accessible to the medical community.
3 Research program
3.1 Complex models for the propagation of cardiac action potentials
The contraction of the heart is coordinated by a complex electrical activation process which relies on about a million ion channels, pumps, and exchangers of various kinds in the membrane of each cardiac cell. Their interaction results in a periodic change in transmembrane potential called an action potential. Action potentials in the cardiac muscle propagate rapidly from cell to cell, synchronizing the contraction of the entire muscle to achieve an efficient pump function. The spatio-temporal pattern of this propagation is related both to the function of the cellular membrane and to the structural organization of the cells into tissues. Cardiac arrythmias originate from malfunctions in this process. The field of cardiac electrophysiology studies the multiscale organization of the cardiac activation process from the subcellular scale up to the scale of the body. It relates the molecular processes in the cell membranes to the propagation process through the multiscale structure of the tissue and organ, to measurable signals in the heart and to the electrocardiogram, an electrical signal on the torso surface.
Several improvements of current models of the propagation of action potentials are being developed in the Carmen team, based on previous work 42 and on the data available at IHU Liryc:
- Enrichment of the current monodomain and bidomain models 42, 53 by accounting for structural heterogeneities of the tissue at cellular and intermediate scales. Here we focus on multiscale analysis techniques applied to the various high-resolution structural data available at IHU Liryc.
- Coupling of the tissues from the different cardiac compartments and conduction systems. Here, we develop models that couple 1D, 2D and 3D phenomena described by reaction- diffusion PDEs.
These models are essential to improve our understanding of cardiac electrical dysfunction. To this aim, we use high-performance computing techniques in order to numerically explore the complexity of these models.
We use these model codes for applied studies in two important areas of cardiac electrophysiology: atrial fibrillation 46 and sudden-cardiac-death (SCD) syndromes 7, 649. This work is performed in collaboration with several physiologists and clinicians both at IHU Liryc and abroad.
3.2 Simplified models and inverse problems
The medical and clinical exploration of the cardiac electric signals is based on accurate reconstruction of the patterns of propagation of the action potential. The correct detection of these complex patterns by non-invasive electrical imaging techniques has to be developed. This involves solving inverse problems that cannot be addressed with the more complex models. We want both to develop simple and fast models of the propagation of cardiac action potentials and improve the solutions to the reconstruction questions of cardiac electrical imaging techniques.
These questions concern the reconstruction of diverse information, such as cardiac activation maps or, more generally, the whole cardiac electrical activity, from high-density body surface electrocardiograms. It is a possibly powerful diagnosis technique, which success would be considered as a breakthrough. Although widely studied during the last decade, the reconstructed activation maps, for instance, are highly inacurate and have a poor clinical interest. It remains a challenge for the scientific community to understand how body surface signals can better inform on the fine details of arrhythmic mechanisms.
The most usual method consists in solving a Laplace equation on the volume delimited by the body surface and the epicardial surface, for which we contribute by:
- studying in depth the dependance of the inverse problem on inhomogeneities in the torso, conductivity values, the geometry, electrode positions, etc., and
- improving the solution to the inverse problem by using new regularization strategies, factorization of boundary value problems, and the theory of optimal control.
In addition, we have started to explore many alternative approaches including:
- using complete propagation models in the inverse problem, like the bidomain or monodomain equations, for instance in order to localize electrical sources,
- constructing data-based models using e.g. statistical learning techniques, which would accurately represent some families of well-identified pathologies, or allow to combine physics and biology-informed models and clinical data, and
- constructing simpler models of the propagation of the activation front, based on eikonal or level-set equations.
3.3 Numerical techniques
We want our numerical simulations to be efficient, accurate, and reliable with respect to the needs of the medical community. Based on previous work on solving the monodomain and bidomain equations 44, 43, 52, 34, we will focus on:
- high-order numerical techniques with respect to the variables with physiological meaning, like velocity, AP duration and restitution properties and
- efficient, dedicated preconditioning techniques coupled with parallel computing.
Existing simulation tools used in our team rely, among others, on mixtures of explicit and implicit integration methods for ODEs, hybrid MPI-OpenMP parallellization, algebraic multigrid preconditioning, and Krylov solvers. New developments include high-order explicit integration methods and task-based dynamic parallellism.
3.4 Cardiac electrophysiology at the microscopic scale
Traditional numerical models of whole-heart physiology are based on the approximation of a perfect muscle using homogenisation methods. However, due to aging and cardiomyopathies, the cellular structure of the tissue changes. These modifications can give rise to life-threatening arrhythmias, the mechanisms of which we are investigating in collaboration with cardiologists at the IHU Liryc. For this research we are building models that describe the strong heterogeneity of the tissue at the cellular level.
The literature on this type of model is still very limited 59. Existing models are two-dimensional 50 or limited to idealized geometries, and use a linear (purely resistive) behaviour of the gap-juction channels that connect the cells. We propose a three-dimensional approach using realistic cellular geometry (Fig. 1), nonlinear gap-junction behaviour, and a numerical approach that can scale to hundreds of cells while maintaining a sub-micrometer spatial resolution (10 to 100 times smaller than the size of a cardiomyocyte). Following preliminary work in this area by us 38, 37, 36 and by others 59 we proposed a European project with 10 partner institutes and a 5.8M€ budget to develop software that can simulate such models, with micrometer resolution, on the scale of millions of cells, using future exascale supercomputers (microcard.eu). This project runs from April 2021 to October 2024, and involves also the Inria teams CAMUS, STORM and CARDAMOM as well as the Inria-led MMG Consortium.
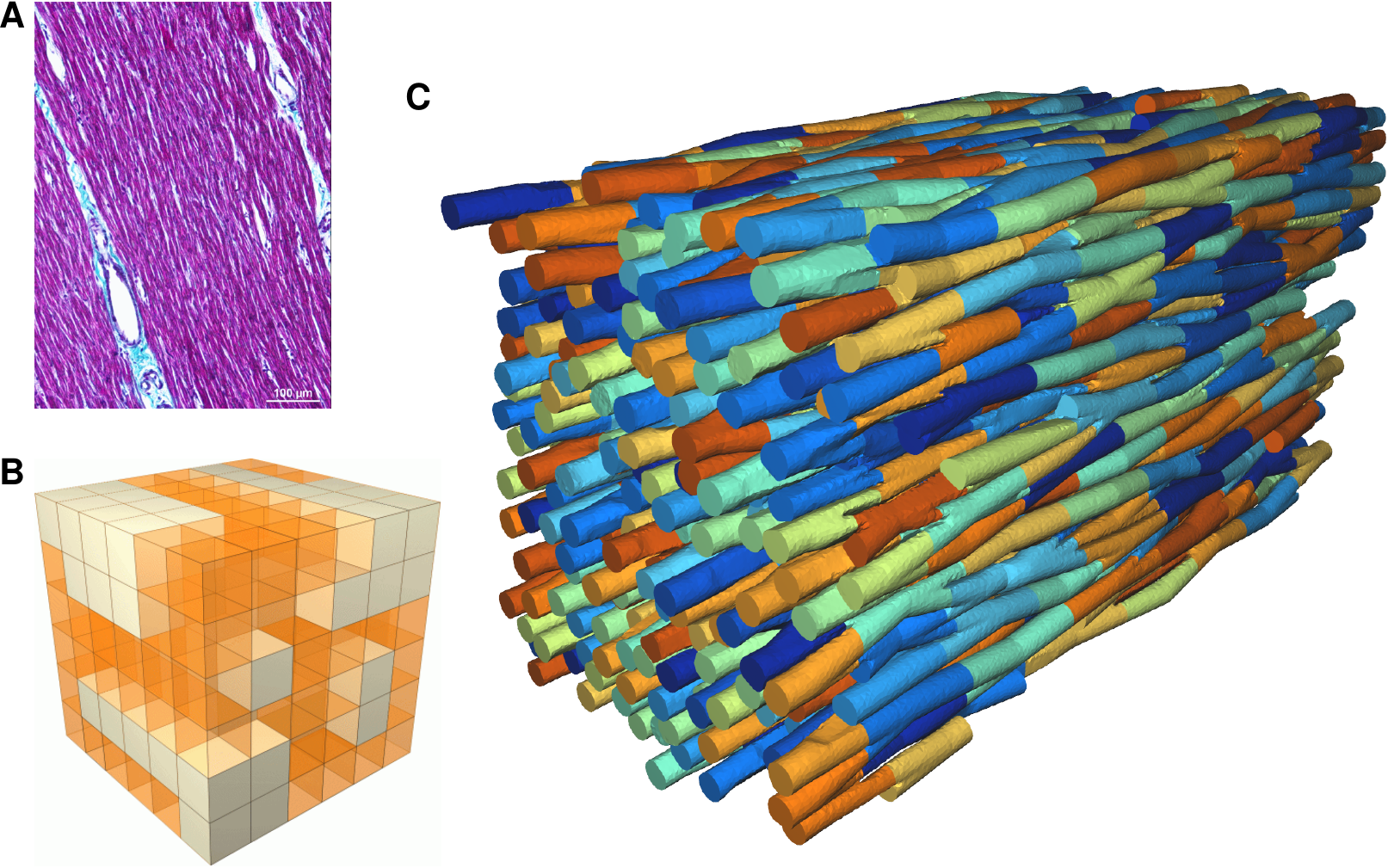
Image of the microstructure from histology, current, insufficient, representation of microstructural defects, and foreseen geometry to be used in microscopic models.
3.5 Models and tools for ablative therapies
Today, the most effective way to treat arrhythmias is to ablate selected regions of the cardiac tissue. As the lesions have no particular electric property, this creates conduction blocks that stop the disorganized propagation of action potentials. The ablation procedure consists in placing a catheter in contact with the targeted site and deliver energy into the tissue. The energy can be overheating by radio-frequency current, electroporating electric pulses or temperature drop (cryotherapy). In practice, the choice of the ablation site is done by the clinician based on previous signal measurements and imagery, and is also guided during the procedure with real-time measurement of the electric signal.
Our team works on several subjects related to ablation techniques that may improve the success rate of the treatments:
- accurate computation of electric fields generated by catheters: complex catheter shapes, contact models, tissue heterogeneities;
- models of creation of the lesions, either through temperature rise (radio-frequency) or electroporation; and
- localization tools to help clinicians target the optimal ablation sites, based on both data of previously ablated patients and synthetic data.
4 Application domains
4.1 Scientific context: IHU Liryc
The University Hospital of Bordeaux (CHU de Bordeaux) is equipped with a specialized cardiology hospital, the Hôpital Cardiologique du Haut-Lévêque, where the group of Professor Michel Haïssaguerre has established itself as a global leader in the field of cardiac electrophysiology 48, 47, 40. Their discoveries in the area of atrial fibrillation and sudden cardiac death syndromes are widely acclaimed, and the group is a national and international referral center for treatment of cardiac arrhythmia. Thus the group also sees large numbers of patients with rare cardiac diseases.
In 2011 the group has won the competition for a 40 million euro Investissements d'Avenir grant for the establishment of IHU Liryc, an institute that combines clinical, experimental, and numerical research in the area of cardiac arrhythmia. The institute works in all areas of modern cardiac electrophysiology: atrial arrhythmias, sudden death due to ventricular fibrillation, heart failure related to ventricular dyssynchrony, and metabolic disorders. It is recognized worldwide as one of the most important centers in this area.
The Carmen team was founded as a part of IHU Liryc. We bring applied mathematics and scientific computing closer to experimental and clinical cardiac electrophysiology. In collaboration with experimental and clinical researchers at Liryc we work to enhance fundamental knowledge of the normal and abnormal cardiac electrical activity and of the patterns of the electrocardiogram, and we develop new simulation tools for training, biological, and clinical applications.
4.2 Basic experimental electrophysiology
Our modeling is carried out in coordination with the experimental teams from IHU Liryc. It helps to write new concepts concerning the multiscale organisation of the cardiac action potentials that will serve our understanding in many electrical pathologies. For example, we model the structural heterogeneities at the cellular scale 39 (the MICROCARD project), and at an intermediate scale between the cellular and tissue scales.
At the atrial level, we apply our models to understand the mechanisms of complex arrythmias and the relation with the heterogeneities at the insertion of the pulmonary veins. We will model the heterogeneities specific to the atria, like fibrosis or fatty infiltration 55, 46. These heterogeneities are thought to play a major role in the development of atrial fibrillation.
At the ventricular level, we focus on (1) modeling the complex coupling between the Purkinje network and the ventricles, which is supposed to play a major role in sudden cardiac death, and (2) modeling the heteogeneities related to the complex organization and disorganization of the myocytes and fibroblasts, which is important in the study of infarct scars for instance.
4.3 Clinical electrophysiology
Treatment of cardiac arrhythmia is possible by pharmacological means, by implantation of pacemakers and defibrillators, and by curative ablation of diseased tissue by local heating, freezing or electroporation. In particular the ablative therapies create challenges that can be addressed by numerical means. Cardiologists would like to know, preferably by noninvasive means, where an arrhythmia originates and by what mechanism it is sustained.
We address this issue in the first place using inverse models, which attempt to estimate the cardiac activity from a (high-density) electrocardiogram. A new project aims to perform this estimation on-site in the catheterization laboratory and presenting the results, together with the cardiac anatomy, on the screen that the cardiologist uses to monitor the catheter positions 51, 35.
4.4 Application in Deep Brain Stimulation
Since 2017, we have been working with neurosurgeons from the Bordeaux University Hospital (Pr Cuny and Dr. Engelhardt) on improving the planning technique for deep brain surgery (DBS) for Parkinson's and Essential tremor diseases. DBS is the last resort to treat the symptoms of Parkinson's disease after the drug Levodopa. The surgery consists in placing electrodes in very specific regions of the patient's brain. These regions are unfortunately not visible on the 1.5 Tesla MRI, the most widely available MRI machines in hospitals. The most effective solution to date is to introduce 5 micro-electrodes (MER) to record the activity of neurons in the patient's brain and to prospect by moving the electrodes in order to find the best location. However, this approach renders the surgery very cumbersome because the patient must be awake during the exploration phase. In addition, this phase takes at least 3 hours and mobilizes a neurologist with his staff. The total duration of the operation is between 7 and 8 hours. Many elderly patients do not tolerate this surgery. We have proposed an approach that avoids the prospecting phase and performs surgery under general anesthesia. The idea is to learn on pairs of clinical landmarks and the position of active electrodes in order to predict the optimal position of the DBS from a pre-operative image. This approach simplifies and standardizes surgery planning. We tested several approaches doi:10.3389/fneur.2021.620360. We continue to seek approaches to fully automate the targeting process. We carried out a proof of concept by learning on the clinical database of the Bordeaux University Hospital. The clinical validation of our approach is in progress through a clinical trial which includes patients from the University Hospitals of Bordeaux and Lyon. Pr Cuny has submitted a phase 3 national clinical research hospital project (PHRCN) including 11 CHUs in France which has been accepted by the General Directorate for Care Offers (DGOS). The aim is to compare our new approach to the ones used in the other centers. Inria Bordeaux is a partner in this project and we maintain the OptimDBS software and solve any technical problem related to the compatibility of the MRIs exported by our software and the surgical robots in the different centers.
5 Social and environmental responsibility
5.1 Footprint of research activities
We avoid flying whenever we can. For example, we used trains to travel to Nice (10 hours) and Budapest (2 days). One of us traveled to the 2022 CinC meeting in Tampere (Finland) by bicycle.
5.2 Impact of research results
The MICROCARD project, which we coordinate, has energy efficiency as one of its goals. To this end, our partners in the STORM and CAMUS teams are developing methods to increase the time- and energy-efficiency of cardiac simulation codes.
6 Highlights of the year
- Organization of the PersonalizeAF (H2020-MSCA-ITN) Bordeaux summer school
- Organization of the 2nd MICROCARD Workshop (EuroHPC JU)
- Kick-off of the associated team SPICY
- Evaluation of the team
7 New software and platforms
7.1 CEPS
No new feature of high relevance was added to CEPS this year. However, the software is currently being deeply rewritten for the H2020 SimCardioTest project. In this project, it will be used as the core computing engine of a cloud based platform for in silico trials, targeting the design of pacemakers.
- The pacemaker model that is derived (see section 8.3) requires to changes deeply the architecture of CEPS.
- In parallel, in order to comply with software quality criteria described by the FDA in the V&V40 standard, CEPS is now being monitored through the SonarQube platform of Inria, and a much more thorough testing suite is being written.
7.2 Mmg and ParMmg
Mmg and ParMmg are intensively used in the MICROCARD project to automatically generate 3D meshes of cardiac tissues from imaging data. The CARMEN teams contributes to Mmg, with the main objective of making the software robust when dealing with complex meshes provided by Orobix (italian partner company in MICROCARD).
ParMmg is a parallel unstructured mesh adaptation software using iterative remeshing and repartitionning. It uses the Mmg software to perform the sequential remeshing steps. A new version of Mmg (5.7.0) has been released this year, and improvement of the level-set discretization is being investigated. In addition, a new version of ParMmg is under development to be able to run on large heterogeneous supercomputers.
7.3 New software
7.3.1 CEPS
-
Name:
Cardiac ElectroPhysiology Simulation
-
Keywords:
Simulation, Health, Mesh, Cardiac, 3D, Cardiac Electrophysiology
-
Scientific Description:
As compared to other existing softwares, CEPS aims at providing a more general framework of integration for new methods or models and a better efficiency in parallel. CEPS is designed to run on massively parallel architectures, and to make use of state-of-the-art and well known computing libraries to achieve realistic and complex heart simulations. CEPS also includes software engineering and validation tools.
-
Functional Description:
CEPS is a modular high-performance computing software for performing numerical simulations in cardiac electrophysiology. It is based on modules : - management of geometries represented by meshes in 3D, 2D or 1D (volumes, surfaces, trees), - model simulation of cellular electrophysiology, - calculating the tissue propagation of the action potentials in the cardiac geometries, - calculation of extracardiac potentials, - time approximation methods in order 2, 3 and 4 specific to electrocardiography.
- URL:
-
Contact:
Michael Leguebe
-
Participants:
Mehdi Juhoor, Nejib Zemzemi, Antoine Gerard, Charlie Douanla Lontsi, Pierre-Elliott Bécue, Marc Fuentes, Yves Coudière, Michael Leguebe, Andjela Davidovic, Pauline Migerditichan, Florian Caro
-
Partners:
Université de Bordeaux, Fondation Bordeaux Université, CHU de Bordeaux, Inria
7.3.2 OptimDBS
-
Name:
Optimizing the Deep Brain Stimulation
-
Keywords:
Image analysis, Deep brain stimulation, Statistical learning
-
Functional Description:
Targeting software for deep brain stimulation
- URL:
-
Contact:
Nejib Zemzemi
-
Participants:
Nejib Zemzemi, Louise-Amelie Schmitt, Emmanuel Cuny, Julien Engelhardt
-
Partner:
CHU de Bordeaux
7.3.3 MUSIC - Carmen plugins
-
Name:
Carmen plugins for multi-modality imaging in Cardiology
-
Keywords:
Image segmentation, Mesh generation, Image filter, Numerical simulations, Cardiac Electrophysiology, Inverse problem, Finite element modelling, Visualization, 3D interaction, Registration
-
Scientific Description:
Carmen plugins is a collection of toolboxes and pipelines allowing the following functionalities: - Segmenting and filtering the heart, the surrounding organs and the body surface. - Generate/optimize surface and volume of computational meshes for the segmented organs. - Generate fibers orientations for Atria and Ventricles. - Interactively annotate meshes. - Read map and visualize electrical information collected from medical devices. - Numerical simulation of the forward problem using finite elements method. - Method of fundamental solutions for solving the ECGI inverse problem combined with the regularization methods including CRESO,Zero Crossing GCV, RGCV, ADPC and Ucurve. - Landmark based mesh registration. - ECGI pipeline: including segmentation, mesh generation, identification of the vest electrodes.
-
Functional Description:
Carmen plugins is a collection of toolboxes and pipelines allowing the following functionalities: - Segmenting and filtering the heart, the surrounding organs and the body surface. - Generate/optimize surface and volume of computational meshes for the segmented organs. - Generate fibers orientations for Atria and Ventricles. - Interactively annotate meshes. - Read map and visualize electrical information collected from medical devices. - Numerical simulation of the forward problem using finite elements method. - Method of fundamental solutions for solving the ECGI inverse problem combined with the regularization methods including CRESO,Zero Crossing GCV, RGCV, ADPC and Ucurve. - Landmark based mesh registration. - ECGI pipeline: including segmentation, mesh generation, identification of the vest electrodes.
- Publications:
-
Contact:
Nejib Zemzemi
-
Participants:
Hubert Cochet, Florent Collot, Mathilde Merle, Maxime Sermesant, Julien Castelneau, Mehdi Juhoor, Pauline Migerditichan, Nejib Zemzemi, Yves Coudière
-
Partners:
Université de Bordeaux, IHU - LIRYC
Participants: Mark Potse, Andony Arrieula.
7.4 New platforms
CEMPACK
CEMPACK is a collection of software that was previously archived in different places. It includes the high-performance simulation code Propag and a suite of software to create geometric models, prepare inputs for Propag, and analyse its outputs. In 2017 the code was collected in an archive on Inria's GitLab platform. The main components of CEMPACK are the following.
-
Propag-5.1
Applied modeling studies performed by the Carmen team in collaboration with IHU Liryc and foreign partners 756, 46, 45, 41 rely on high-performance computations on the national supercomputers Irene, Zay, and Adastra. The Propag-5 code is optimized for these systems. It is the result of a decades-long development first at the Université de Montréal in Canada, then at Maastricht University in the Netherlands, and finally at the Institute of Computational Science of the Università della Svizzera italiana in Lugano, Switzerland. Since 2016 most of the development on Propag has been done by M. Potse at the Carmen team 58. The code scales excellently to large core counts 57 and, as it is controlled completely with command-line flags and configuration files, it can be used by non-programmers. It also features:
- a plugin system for membrane models,
- a completely parallel workflow, including the initial anatomy input and mesh partitioning, which allows it to work with meshes of more than nodes,
- a flexible output scheme allowing hundreds of different state variables and transient variables to be output to file, when desired, using any spatial and temporal subsampling,
- a configurable, LUSTRE-aware parallel output system in which groups of processes write HDF5/netCDF files, and
- CWEB documentation of the entire code base.
The code has been stable and reliable for many years. It can be considered the workhorse for our HPC work until CEPS takes over.
-
Gepetto
The Gepetto suite, named after a famous model maker, transforms a surface mesh of the heart into a set of (semi-)structured meshes for use by the Propag software or others. It creates the different fiber orientations in the model, including the transmurally rotating ventricular fibers and the various bundle structures in the atria (figure 2), and creates layers with possibly different electrophysiological properties across the wall. A practically important function is that it automatically builds the matching heart and torso meshes that Propag uses to simulate potentials in the torso (at a resolution of 1 mm) after projecting simulation results from the heart model (at 0.1 to 0.2 mm) on the coarser torso mesh 54. Like Propag, the Gepetto software results from a long-term development that started in Montreal, Canada, around 2002. The code for atrial fiber structure was developed by our team.
-
Blender plugins
Blender is a free software package for the production of 3-D models, renderings, and animations, comparable to commercial software such as Cinema4D. CEMPACK includes a set of plugins for Blender that facilitate the production of anatomical models and the visualization of measured and simulated data. It uses the MMG remeshing library, which is developed by the CARDAMOM team at Inria Bordeaux. As of 2022, the segmentation work is mostly done with the MUSICardio software, but we still use Blender for finishing touches and high-quality visualization.
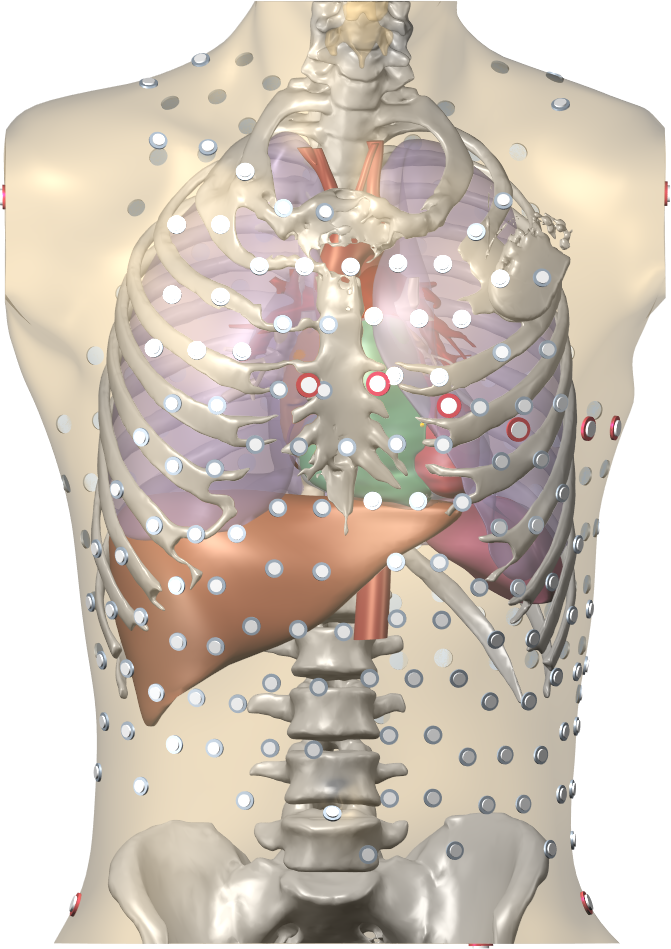
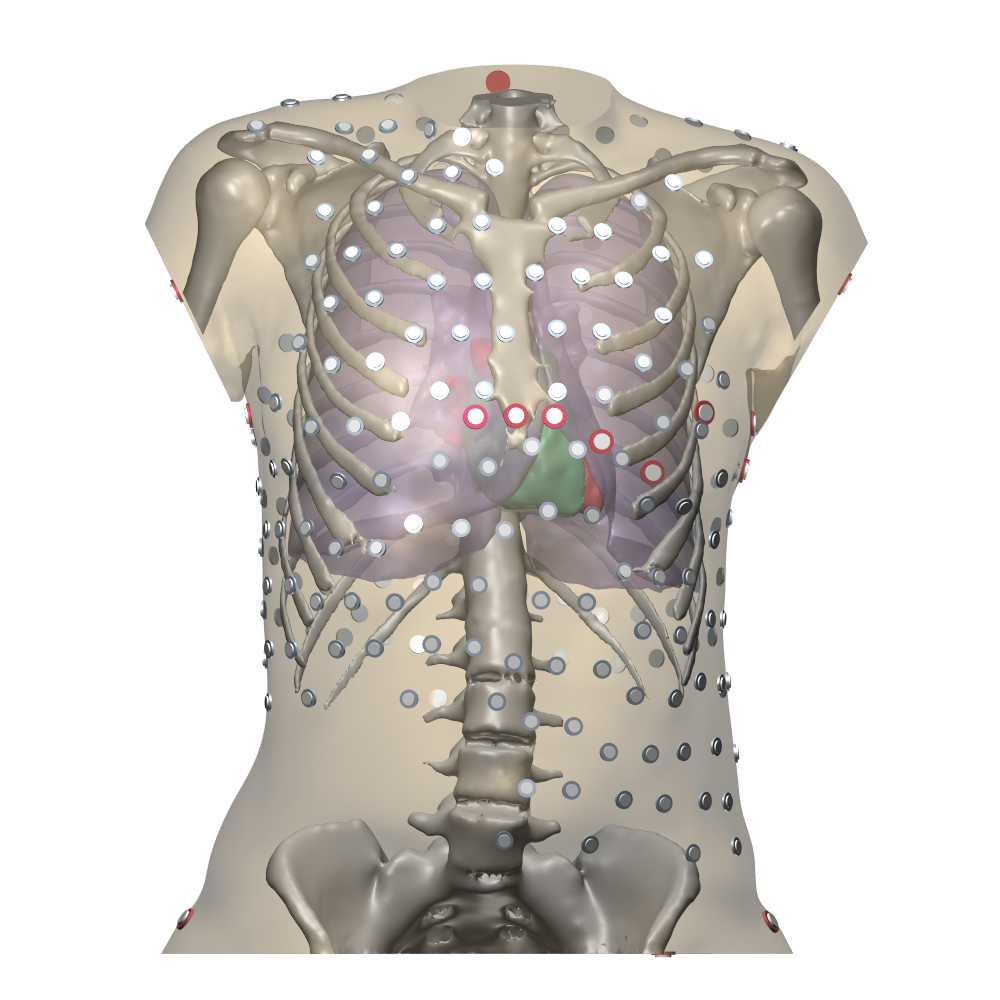
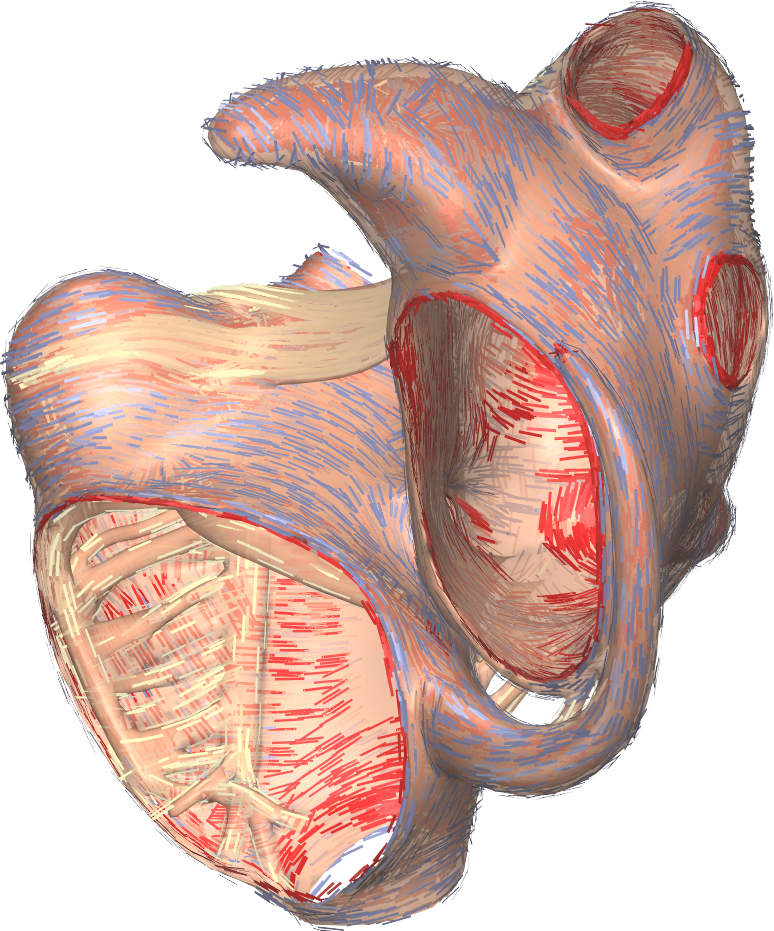
Participants: Mostafa Bendahmane, Yves Coudière, Jacques Henry, Peter Langfield, Michael Leguèbe, Mark Potse, Lisl Weynans, Nejib Zemzemi.
8 New results
8.1 Analysis of partial differential equations
- Global Lipschitz stability of multiple space dependent ionic parameters for the MEA/hiPSC-CM assays 9. The work is now published in Inverse Problems journal (DOI 10.1088/1361-6420/aca70d). This work is devoted to analyze the inverse problem of determining two space dependent ionic parameters of a nonlinear reaction diffusion system arainsing from MEA/hiPSC-CM assays model described in 33. We consider the classical phenomenological model in cardiac electrophysiology of FitzHugh-Nagumo to describe the ionic exchanges at the microscopic level. Our main result is the uniqueness and a Lipschitz stability estimate for two ionic parameters (k, γ) of the model using sub-boundary observations over an interval of time. The key ingredients are global Carleman-type estimates with a suitable observations acting on a part of the boundary.
8.2 Numerical analysis and development of numerical methods
-
The foundations of several projects that use numerical continuation techniques have been established. Most notably, a method for computing isochrons in higher-dimensional models has been developed. To demonstrate the method, it has been applied to the four dimensional Hodgkin-Huxley model. The computed isochrons are consistent with previous depictions, but the method is also able to uncover complicated isochron geometry arising from numerical sensitivities in the model that were not previously resolved.
Additionally, the development of a toolbox with the aim of studying propagating waves has started. In practice, this is a numerical continuation solver based on the Auto package, but with spatial domains that have appropriate discretizations and solution representations. The toolbox is implemented partially in Python as a prototype, where further detailed analysis and testing can be performed.
- In the context of Electrical Impedance Tomography, we proposed a new system of equations to obtain a unique solution for the Complete Electrode Model and proved the convergence of an immersed boundary method to solve numerically this model.
- In the context of the MICROCARD project, we have extended the standard two point flux approximation finite volume method to the microscopic bidomain equations, proved its convergence, and obtained some error estimate. A prototype code is currently being implemented. The work was presented at the DDFV workshop in Marseille, October 17th-21st, 2022.
8.3 Modeling and inverse problems
- The Ph.D. thesis of Bachar Tarraf was defended in February 2022 29. We extended our simple model of cardiac mitochondria to include ROS production and mPTP opening. We tried several machine learning techniques to generate a classification of mitochondria in different respiratory media and try to reproduce data from our collaborators within the Mitocard ANR project. However, the data were hardly exploitable to establish a reference classification from the start, and we were not able to determine specifically any parameter from our extended model.
-
Modeling pacemakers: in the context of the H2020 SimCardioTest European project, we derived a simple model of electric stimulation by a pacemaker, in collaboration with Microport CRM, manufacturer. To our knowledge, the study of cardiac stimulation is done along two research axes that are dissociated. On one hand, extensive literature can be found on the polarization phenomena that occur at the surface of an electrode when it delivers current into an electrolyte. On the other hand, stimulation of cardiac tissue is generally made through a simple rectangle function that acts as source term of the monodomain or bidomain equations, representing a direct injection of current into the tissue.
We are now coupling a model of pacemaker and a model of electrode contact to a stimulation model. In order to determine the parameters of the contact, that depend on the shape and material of each pacemaker electrode, we used data from bench tests performed at Microport. The model represents the contact by a simple capacitance in parallel with a resistance. Even if the model is quite simple, we already faced difficulties to calibrate the parameters. Even if all the physics could not be explained by this model, we obtained fits that differ by 2% (in L2 relative error) from the data, which is suitable for using them as input to a cardiac model.
We have coupled this contact model to a 0D model a cardiac cell stimulation in order to determine the parameters of the complete model that influence the voltage and duration of stimulation that are required to excite the cells, determining the so-called Lapicque curve. The results of this study will be submitted to FIMH 2023. We are also currentlty implementing a coupled 3D pacemaker/bidomain model.
-
In the framework of the international ECGI consortium, we continue the work on the assessement of the Effect of the segmentation uncertanties on the ECGI forward and inverse problem solutions.
This year we had many publications at the international conference of Computing in Cardiology 2022 that hold in Tampere. In the first study 26 we quantify the dependence of a cardiac propagation model on segmentation variability. We used statistical shape modeling and polynomial Chaos (PC) to capture segmentation variability dependence and applied its affects to a propagation model. We evaluated the predicted local activation times (LATs) an body surface potentials (BSPs) from two modeling pipelines: an Eikonal propagation model and a surface-based fastest route model. The predicted uncertainty due to segmentation shape variability was distributed near the base of the heart and near high amplitude torso potential regions. Our results suggest that modeling pipelines may have to accommodate segmentation errors if regions of interest correspond to high segmentation error. Further, even small errors could proliferate if modeling results are used to to feed further computations, such as ECGI.
In the second study 18, we generated a statistical shape model from segmentations of the same patient and generated 262 cardiac geometries to run in an ECG forward computation of body surface potentials (BSPs) using an equivalent dipole layer cardiac source model and 5 ventricular stimulation protocols. Variability between simulated BSPs for all models and protocols was assessed using Pearson's correlation coefficient (CC). Compared to the BSPs of the mean cardiac shape model, the lowest variability (average CC = 0.98 ± 0.03) was found for apical pacing whereas the highest variability (average CC = 0.90 ± 0.23) was found for right ventricular free wall pacing. Furthermore, low amplitude BSPs show a larger variation in QRS morphology compared to high amplitude signals. The results indicate that the uncertainty in cardiac shape has a significant impact on ECGI.
In the third study 17, we analysed the effect of segmentation variability on the ECGI estimation of the cardiac activity with 262 shape models generated from fifteen different segmentations. Therefore, we designed two test cases: with and without shape model uncertainty. Moreover, we used four cases for ectopic ventricular excitation and compared the ECGI results in terms of reconstructed activation times and excitation origins. The preliminary results indicate that a small variation of the activation maps can be observed with a model uncertainty but no significant effect on the source localization is observed.
- In the context of the project about electrical impedance tomography, we developed a method to reconstruct not only the conductivities inside the domain of interest, but also the shape of the domain itself and the location of the electrodes. The code for the inverse problem of Electrical Impedance Tomography was validated in sequential 2D, and parallelized.
- In collaboration with Jérôme Fehrenbach (Univ. Toulouse), we developed a fast algorithm to estimate the location of activation based on an eikonal equation.
8.4 Clinical electrophysiology
- Andony Arrieula defended his PhD thesis in October 2022 27. In this thesis he developed two methods to help find suitable ablation sites for premature ventricular beats during a cardiac catheterization procedure. The first method uses ECG data from beats stimulated with the catheter at known positions, accurately measured with the clinical catheter localization system. Using linear interpolation it can subsequently infer the location of a premature beat from a measured ECG. The main challenge here was to deal with both underdetermined and overdetermined linear systems. The second method is entirely noninvasive: after training with thousands of simulated ECGs, a machine-learning technique predicts the site of origin from an ECG and with help from an anatomical model of the heart extracted from clinical imaging data. Both methods were tested extensively on simulated data and on clinical data.
8.5 High performance computing
- We used high-performance computing resources principally for the work described in section 8.4.
- We provided simulated data to test machine-learning and signal analysis methods 23, 21.
- We used simulations to investigate whether inter-individual differences in cell composition across the ventricular wall may explain variability in the ECG response to serum potassium and calcium variations 19.
- We tested a method aimed to accelerate the initialization of whole-heart models with strongly abnormal parameter settings 20.
8.6 Microscopic models
- Within the MICROCARD project the CARMEN team is also involved in the construction of synthetic models of the cardiac muscle at the cellular scale. These models are to complement the imaging-derived models, which will be very limited in size. We reported a first method for this purpose at the ECCOMAS meeting in June 24. At the time of the meeting we were still limited to models of a few dozen cells. Thanks to the improvements in the Mmg software, released in version 5.7.0 in December 2022, we have leaped to thousands of cells in the last few days of the year.
- As detailed above, a finite volume numerical method was defined and analyzed to wolve the microscopic bidomain esuations on these geometries.
9 Bilateral contracts and grants with industry
Participants: Nejib Zemzemi.
9.1 Bilateral Grants with Industry
RebrAIn and Inria contracted an agreement allowing Nejib Zemzemi to pass half of his time working at RebrAIn. The startup RebrAIn has been co-founded by N. Zemzemi and E. Cuny.
Lisl Weynans obtained a grant from EDF Foundation to found a research project about Electrical Impedance Tomography.
10 Partnerships and cooperations
Participants: Mostafa Bendahmane, Yves Coudière, Jacques Henry, Peter Langfield, Michael Leguèbe, Mark Potse, Lisl Weynans, Nejib Zemzemi.
10.1 International initiatives
10.1.1 Associate Teams in the framework of an Inria International Lab or in the framework of an Inria International Program
SPICY
-
Title:
Stochastic forward and inverse Problems In Cardiac electrophysiologY
-
Date:
From 2021
-
Coordinator:
Mourad Bellasoued (mourad.bellassoued@enit.utm.tn)
-
Partners:
- Université de Tunis El Manar Tunis (Tunisie)
-
Inria contact:
Mostafa Bendahmane
-
Summary:
The electrocardiography imaging inverse problem is frequently solved using the deterministic quasi-static models. These models don't take into account the heart dynamic in time, channel noise and external random perturbations acting in the torso. Recent numerical studies in the direct problem have shown that such randomness cannot be suppressed. Occasionally deterministic equations give qualitatively incorrect results. Therefore, it is important to quantify the nature of the noise and choose an appropriate model incorporating randomness. In our project, we study the inverse problem constrained by the stochastic monodomain or bidomain equations in electrocardiology. The state equations consist in a coupled stochastic reaction-diffusion system modelling the propagation of the intracelullar and extracellular electrical potentials, and stochastic ionic currents in the heart. These equations are coupled to the stochastic quasi-static elliptic equation in the torso. Thus, we will demonstrate that the novel concept of applying the stochastic model will be useful to improve noninvasive reconstruction of electrical heart activity. We will perform numerical experiments representing the effect of the stochastic heart dynamic on the inverse solutions. Moreover, we will study the stability result for the conductivities and numerically solve the parameters estimations problem in the stochastic model.
10.1.2 Other International Programs
ECOS Sud
-
Title:
Virtual Element Methods for Bidomain Model For Cardiac Electrophysiology
-
Partner Institution:
University of the Bío-Bío, Concepción, Chile
-
Coordinators:
Mostafa Bendahmane (Inria contact), Veronica Anaya (vanaya@ubiobio.cl)
-
Date
From 2021
-
Description:
Our project is framed within the area of mathematical modeling and numerical simulations in physiology, particularly we explore the cardiac electrical potentials and we develop several applications using VEM discretizations to bidomain model. For instance, primal and mixed formulations will be considered. The main motivation to study this model is that it constitutes a stepping stone towards the more complex models with important applications in cardiac dynamics where we could have complex domains. Thus, we can explote the capability of VEM to use general meshes. We also plan to derive a posteriori error estimators for the VEM and use it to drive space/time adaptive schemes.
10.1.3 International research visitors
Cf. section Team Members, Visiting scientists.
10.1.4 Visits to international teams
Narimane Gassa
-
Visited institution:
EP-Solutions (industrial company)
-
Country:
Switzerland
-
Dates:
2 months
-
Context of the visit:
PersonalizeAF
-
Type of mobility:
Reasearch stay abroad
10.2 European initiatives
10.2.1 H2020 projects
MICROCARD:
MICROCARD project on cordis.europa.eu
-
Title:
Numerical modeling of cardiac electrophysiology at the cellular scale
-
Duration:
From April 1, 2021 to September 30, 2024
-
Partners:
- Université de Bordeaux (Liryc, IMB), coordinator
- Inria (centers Bordeaux and Nancy; teams CARMEN, STORM, CAMUS, CARDAMOM)
- MEGWARE Computer Vertrieb und Service GMBH, Germany
- SIMULA Research Laboratory AS, Norway
- Université de Strasbourg, France
- Konrad-Zuse-Zentrum für Informationstechnik Berlin (ZIB), Germany
- Università della Svizzera italiana (USI), Switzerland
- Karlsruhe Institute for Technology (KIT), Germany
- Università degli studi di Pavia, Italy
- Bordeaux INP, France
- Numericor GMBH, Austria
- Orobix SR, Italy
-
Inria contact:
Mark POTSE
-
Coordinator:
Mark POTSE
-
Summary:
Numerical models of cardiac electrophysiology are highly sophisticated and widely used, but to match observations in aging and diseased hearts they need to move from a continuum approach to a representation of individual cells and their interconnections. This implies a different, harder numerical problem and a 10,000-fold increase in problem size. Exascale computers will be needed to run such models.
In the MICROCARD project we develop an exascale application platform for cardiac electrophysiology simulations that is usable for cell-by-cell simulations. The platform is co-designed by HPC experts, numerical scientists, biomedical engineers, and biomedical scientists, from academia and industry. We develop, in concert, numerical schemes suitable for exascale parallelism, problem-tailored linear-system solvers and preconditioners, and a compiler to translate high-level model descriptions into optimized, energy-efficient system code for heterogeneous computing systems. The code will be parallelized with a recently developed runtime system that is resilient to hardware failures and will use an energy-aware task placement strategy.
The platform will be applied in real-life use cases with high impact in the biomedical domain and will showcase HPC in this area where it is painfully underused. It will be made accessible for a wide range of users both as code and through a web interface.
We further employ our HPC and biomedical expertise to accelerate the development of parallel segmentation and (re)meshing software, necessary to create the extremely large and complex meshes needed from available large volumes of microscopy data.
The platform will be adaptable to similar biological systems such as nerves, and components of the platform will be reusable in a wide range of applications.
SimCardioTest:
Simcardiotest project on cordis.europa.eu
-
Title:
In silico testing and certification of healthcare products
-
Duration:
From January 1, 2021 to December 30, 2024
-
Partners:
- UNIVERSITE DE BORDEAUX, France
- UNIVERSIDAD POMPEU FABRA, Spain
- UNIVERSITAT POLITECNICA DE VALENCIA, Spain
- SIMULA RESEARCH LABORATORY AS, Norway
- INSILICOTRIALS TECHNOLOGIES S.P.A., Italy
- SORIN CRM SAS, France
- EXACTCURE, France
- BOSTON SCIENTIFIC SCIMED INC, USA
- VIRTUAL PHYSIOLOGICAL HUMAN INSTITUTE FOR INTEGRATIVE BIOMEDICAL RESEARCH VZW, Belgium
-
Inria contact:
Yves Coudière
-
Coordinator:
Maxime Sermesant
-
Summary:
Computer modelling and simulation have the power to increase speed and reduce costs in most product development pipelines. The EU-funded SimCardioTest project aims to implement computer modelling, simulation and artificial intelligence to design and test cardiac drugs and medical devices. Scientists will establish a platform for running in silico trials and obtaining scientific evidence based on controlled investigations. The simulation of disease conditions and cohort characteristics has the potential to overcome clinical trial limitations, such as under-representation of groups. It also reduces the size and duration of human clinical trials as well as animal testing, and offers robust, personalised information. Leveraging in silico technology in healthcare will expedite product and drug certification and offer patients the best possible care.
PersonalizeAF:
PersonalizeAF project on cordis.europa.eu
-
Title:
Personalized Therapies for Atrial Fibrillation. A Translational Approach
-
Duration:
From 1st February 2020 to 31st January 2024
-
Partners:
- KARLSRUHER INSTITUT FUER TECHNOLOGIE Germany
- UNIVERSITEIT MAASTRICHT Netherlands
- UNIVERSITAETSKLINIKUM FREIBURG, Germany
- UNIVERSITE DE BORDEAUX, France
- THE UNIVERSITY OF OXFORD United Kingdom
- CONSORCI INSTITUT D'INVESTIGACIONS BIOMEDIQUES AUGUST PI I SUNYER, Spain
- ALMA MATER STUDIORUM - UNIVERSITA DI BOLOGNA, Italy
- SIMULA RESEARCH LABORATORY AS, Norway
- FUNDACION PARA LA INVESTIGACION BIOMEDICA DEL HOSPITAL GREGORIO MARANON, Spain
- NCARDIA SERVICES BV, Netherlands
- FUNDACION PARA LA INVESTIGACION DEL HOSPITAL UNIVERSITARIO LA FE DE LA COMUNIDAD VALENCIANA, Spain
-
Inria contact:
Nejib Zemzemi
- Coordinator:
-
Summary:
Atrial Fibrillation (AF) is the most common cardiac arrhythmia affecting more than 6 million Europeans with a cost exceeding 1% of the EU health care system budget (13.5 billion annually). New treatment strategies and the progress achieved in research on AF mechanisms and substrate evaluation methods to date have not been commensurate with an equivalent development of the knowledge and technologies required to individually characterize each patient in search of the most efficient therapy.
PersonalizeAF addresses this challenge by delivering an innovative multinational, multi-sectorial, and multidisciplinary research and training programme in new technologies and novel strategies for individualized characterization of AF substrate to and increase treatments’ efficiency.
From the research point of view, PersonalizeAF will integrate data and knowledge from in-vitro, in silico, ex vivo and in vivo animal and human models to: 1) generate an individual description of the state of the atrial muscle identifying the disease mechanisms and characteristics; 2) understanding the potential effect that different therapies have on different atrial substrates; and 3) combining this information to generate a specific profile of the patient and the best therapy for each patient.
With this purpose, PersonalizeAF partnership aggregates relevant scientific staff from the academic and clinical world with highly specialised biomedical companies which will be involved in a high-level personalised training programme that will train a new generation of highly skilled professionals and guarantee ESRs and future PhD students outstanding Career Opportunities in the biomedical engineering, cardiology services and medical devices sectors. PersonalizeAF will disseminate results to a wide spectrum of stakeholders, create awareness in the general public about atrial fibrillation and encourage vocational careers among young students.
10.3 National initiatives
ANR Exacard.
Ended Nov. 2022.
Granted by ANR in July 2018, EXACARD is a collaborative project with computer scientists from Labri that targets hexascale computing in cardiac electrophysiology, it serves as a sandbox to prepare the project EuroHPC MICROCARD. PI for Bordeaux University/Liryc: Yves Coudière.
ANR MITOCARD.
Ended in Apr. 2022.
The MITOCARD project (Electrophysiology of Cardiac Mitochondria), coordinated by S. Arbault (Université de Bordeaux, ISM), was granted by the ANR in July 2017. The objective of MITOCARD is to improve understanding of cardiac physiology by integrating the mitochondrial properties of cell signaling in the comprehensive view of cardiac energetics and rhythm pathologies. It was recently demonstrated that in the heart, in striking contrast with skeletal muscle, a parallel activation by calcium of mitochondria and myofibrils occurs during contraction, which indicates that mitochondria actively participate in Ca2+ signaling in the cardiomyocyte. We hypothesize that the mitochondrial permeability transition pore (mPTP), by rhythmically depolarizing inner mitochondrial membrane, plays a crucial role in mitochondrial Ca2+ regulation and, as a result, of cardiomyocyte Ca2+ homeostasis. Moreover, mitochondrial reactive oxygen species (ROS) may play a key role in the regulation of the mPTP by sensing mitochondrial energetics balance. Consequently, a deeper understanding of mitochondrial electrophysiology is mandatory to decipher their exact role in the heart's excitation-contraction coupling processes. However, this is currently prevented by the absence of adequate methodological tools (lack of sensitivity or selectivity, time resolution, averaged responses of numerous biological entities). The MITOCARD project will solve that issue by developing analytical tools and biophysical approaches to monitor kinetically and quantitatively the Ca2+ handling by isolated mitochondria in the cardiomyocyte.
MITOCARD is a multi-disciplinary project involving 4 partners of different scientific fields: the CARMEN team as well as:
-
ISM,
the largest chemistry laboratory of the Université de Bordeaux, where the necessary measurement methods will be developed,
-
Liryc,
where mitochondria are studied at all levels of integration from the isolated mitochondrion to the intact heart,
-
LAAS,
the MiCrosystèmes d'Analyse (MICA) group at the Laboratory of Analysis and Architecture of Systems, which develops the biological microsensors for this project.
The project will:
- develop chips integrating 4 different electrochemical microsensors to monitor in real-time key mitochondrial signaling parameters: Ca2+, membrane potential, quinone reduction status, O2 consumption, and ROS production,
- develop microwell arrays integrating ring nanoelectrodes to trap single mitochondria within micrometric chambers and measure locally by combined fluorescence microscopy and electrochemical techniques intra- (by fluorescence) and extra-mitochondrial (electrochemistry) metabolites,
- develop a mathematical model of mitochondrial Ca2+ and ROS handling built on existing knowledge, new hypotheses, and the measured data.
The model may serve both to assess biological assumptions on the role of mitochondria in Ca2+ signaling and to integrate pathological data and provide clues for their global understanding.
ANR MAESTRO.
The ANR project MAESTRO (Magnetic Signal detection of ventricular arrhythmOgenic substrates), coordinated by Prof. Michel Haïssaguerre (IHU Liryc), has a computational component for which we recruited a postdoc in December 2022, directed by Mark Potse.
GENCI.
GENCI project A0130307379, “Interaction between tissue structure and ion-channel function in cardiac arrhythmia,” coordinated by Mark Potse, comprises 2.35 million core-hours on the national supercomputers Zay and Joliot-Curie. Compared to previous years it is a modest allocation. This is because most of our computational needs in 2023 are either smaller or larger than the national scale.
11 Dissemination
11.1 Promoting scientific activities
11.1.1 Organization of scientific events
Nejib Zemzemi Organized the Bordeaux Summer school: Experimental electrophysiology and heart computer models summer school. From June 20th to July 1st. The school was followed by the PersonalizeAF workshop (organized by Nejib Zemzemi) from the 4th to the 5th of July 2022.
Conference program committees
- Nejib Zemzemi and Mark Potse reviewed abstracts for Computing in Cardiology and Heart Rhythm Society.
- Peter Langfield was a moderator at the World Congress on Biomechanics in July.
- Mostafa Bendahmane was a member of Scientific committee in Workshop on Mathematics for the Health Sciences at American university of Beirut, Lebanon, 25-26 October 2022.
11.1.2 Journals
We reviewed for numerous journals in the fields of applied mathematics, biomedical engineering, and cardiology. Additionally:
- Mark Potse is an associate editor of Frontiers in Cardiac Electrophysiology and Journal of Electrocardiology.
- Journal Member of the editorial boards: Mostafa Bendahmane is an editor of "Moroccan journal of pure and applied analysis"
11.1.3 Invited talks
- Mark Potse. The MICROCARD project and gender. #RESET your project with gender, 15 November 2022. (webinar)
- Yves Coudière and Zeina Chehade. Towards finite volume methods for the cardiac micromodel. Workshop on Discrete Duality Finite Volume Method and Applications, Marseille, France, October 2022.
- Yves Coudière, Very high order finite volume methods for a reaction-diffusion problem, and a parallel implementation. Lecture at the Discrete Duality Finite Volume Method and Applications, Marseille, France, October 2022.
- Yves Coudière, A tour on equations that model cardiac electrical activity, and some related numerical difficulties. Lecture Series on Mathematical Biology, October 6th, 2022 organized by the Center for Advanced Mathematical Sciences (CAMS) of the American University of Beyruth
- Yves Coudière, Modeling the propagation of cardiac action potential in hearts with structural heterogeneities. Workshop on Mathematics for the Health Sciences, Departement of Mathematics of the American University of Beyruth, October, 25th-26th, 2022.
- Mark Potse. Modeling the heart, cell by cell. HiPEAC 2022, Budapest, June 2022. (HeLP-DC Workshop on Heterogeneous and Low-Power Data Center technologies)
- Mark Potse. MICROCARD: modeling the heart cell by cell. TERATEC Forum, Paris, June 2022. (workshop HPC Technologies and Health)
- Mark Potse. MICROCARD: Numerical modeling of cardiac electrophysiology at the cellular scale. OpenCARP Contributor Meeting, online, 14 March 2022.
- Lisl Weynans, Journées Maths Bio Santé, Besançon, Octobre 2022
- Lisl Weynans: Workshop Méthodes Frontières Immergées en Nouvelle-Aquitaine,04-05 Octobre 2022
- Lisl Weynans: séminaire IDEFIX-MEDISIM-POEMS, Mai 2022
- Mostafa Bendahmane. Recent progress on homogenization of the bidomain and tridomain models in electrocardiology. American University of Beirut, 28 October 2022.
- Mostafa Bendahmane. Recent progress on homogenization of the bidomain and tridomain models in electrocardiology. Mocasim 20 December 2022, University Cadi Ayyad, Marrakech, Morocco.
11.2 Teaching - Supervision - Juries
11.2.1 Teaching
The 2 assistant professors and 1 professor of the team teach at several levels of the Bordeaux University programs in Mathematics, Neurosciences, and Medicine (respectively, 192, 192 and 96 h/year on average). The researchers also have a regular teaching activity, contributing to several courses in the Applied Mathematics at the Bachelor and Master levels (between 16 and 72 h/year).
The PhD students who ask for it are used to teach between 32 and 64 h/year, usually courses of general mathematics in L1 or mathematics for biologists in L1 or L2.
Teaching responsibilities at the University of Bordeaux:
- Yves Coudière: Master MAS (Mathématiques Appliquées, Statistiques), parcours MNCHP (Modélisation Numérique Calcul Haute Performance), until August 2022.
- Lisl Weynans: Licence Mathématique parcours ingénierie mathématique,
- Lisl Weynans: Mineure Mathématiques du parcours International de la Licence,
- Mostafa Bendahmane: Responsable de la mobilité internationale des étudiants de Licence MIASHS.
Courses (L for Bachelor level, M for Master level):
- Numerical analysis (L2)
- Programming for scientific computing with C++ (L3)
- Solving sparse linear systems (L3)
- Programming in Fortran (M1)
- Numerical approximation of PDEs: Finite Differences, Finite Elements, Finite Volumes (M1, M2)
- Supervision of programming projects (L3, M1)
- Mathematical modelling in L2 parcours medicine and physics
- Linear Algebra, Optimization under constraints (L2 and Essca school)
- Analysis, L2
- Computational Neurosciences, M2
- Neuropsychology and Psychophysiology, L3
- Physics for students in medicine, one lecture on cardiac modelling
- Graduate program EUR Digital Public Health, Bordeaux University, March 2019. Philosophy of science and the role of numerical models: introductory course in the module Modeling in life science
11.2.2 Supervision
Cf section Team Members, PhD students.
11.2.3 Juries
- Lisl Weynans: Sixtine Michel (Univ. Bordeaux), jury member
- Lisl Weynans: Mirco Cialella (Univ. Bordeaux), jury member
- Lisl Weynans: Matthias Averseng (Onera, ISAE Toulouse), jury member
- Lisl Weynans: Meriem Zefzouf (Univ. Montpellier), reviewer
- Nejib Zemzemi: Abir Amri (LAMSIN. Tunisia), jury member
- Mark Potse: Éric Irakoze (Université de Montréal), jury member
- Yves Coudière: Giulia Bellezza (Univ. Bordeaux), president of the jury
- Yves Coudière: Oumayma Bouhamama (Univ. Bordeaux), president of the jury
- Yves Coudière: Edmée Eyraud (Univ. Bordeaux), jury member, PhD thesis on chronic obstructive pulmonary disease with mathematical modelling
11.3 Popularization
11.3.1 Mediation responsibilities
- Lisl Weynans co-organizes the event "Moi Mathématicienne, Moi Informaticienne", which is a one-week internship for young girls to discover mathematics and informatics and the jobs associated: webpage of the event. This event has received the "Coup de coeur du jury Talents U" prize of Bordeaux University.
11.3.2 Articles
- Mark Potse co-authored with Emmanuelle Saillard (STORM team, Inria) an article in The Conversation in which we explain how exascale supercomputers will benefit cardiology research 32.
11.3.3 Education
- Yves Coudière: CHICHE at Lycée (High school) F. Magendie, Bordeaux.
- DECLIC event at Gustave Eiffel High-School, Bordeaux.
11.3.4 Interventions
- Nejib Zemzemi participate to a round-table Forum néo-aquitain on artificial intelligence (AI) and robotics, at the NAIA.R 2022 event. The round-table focused on using AI for health care problems, and was entitled (French) "utilisation de l'IA en santé: Nouvelles technologies, nouveaux remèdes?"
11.3.5 Interviews
- The SimCardioTest project has produced a series of interviews for its Youtube channel:
- During the European Researchers’ Night, NEDC at Bordeaux, Yves Coudière, Narimane Gassa, Niami Nasr and Valentin Pannetier held a booth to promote scientific activities of the team, and the three ongoing european projects thanks to an illustrative video: PersonalizeAF, MICROCARD and SimCardioTest.
- At this occasion, Yves Coudière was interviewed by Radio Campus Bordeaux
- Still at the occasion of the NEDC, a short video was produced by the online media Curieux, which is based on a interview of Yves Coudière. It has been shared on various social media, like tiktok.
12 Scientific production
12.1 Major publications
- 1 articleConvergence of discrete duality finite volume schemes for the cardiac bidomain model.Networks and Heterogeneous Media622011, 195-240
- 2 articleA mathematical model of Purkinje-Muscle Junctions.Mathematical Biosciences and Engineering842011, 915-930
- 3 articleExistence And Uniqueness Of The Solution For The Bidomain Model Used In Cardiac Electrophysiology.Nonlinear Anal. Real World Appl.1012009, 458-482URL: http://hal.archives-ouvertes.fr/hal-00101458/fr
- 4 articleA 2D/3D Discrete Duality Finite Volume Scheme. Application to ECG simulation.International Journal on Finite Volumes612009, URL: http://hal.archives-ouvertes.fr/hal-00328251/fr
- 5 articleStability And Convergence Of A Finite Volume Method For Two Systems Of Reaction-Diffusion Equations In Electro-Cardiology.Nonlinear Anal. Real World Appl.742006, 916--935URL: http://hal.archives-ouvertes.fr/hal-00016816/fr
- 6 articleThe Early Repolarization Pattern; A Consensus Paper.Journal of the American College of Cardiology66resulting from the Symposium on J Wave Patterns and a J Wave Syndrome, Glasgow, August 2013. Defines terminology for the J point: Jo, Jp, Jt. Provisionally accepted 19 May 2015.2015, 470-477URL: http://dx.doi.org/10.1016/j.jacc.2015.05.033
- 7 articleReduced Sodium Current in the Lateral Ventricular Wall Induces Inferolateral J-Waves.Front Physiol7365August 2016
- 8 articlePreconditioning the bidomain model with almost linear complexity.Journal of Computational Physics2311January 2012, 82--97URL: http://www.sciencedirect.com/science/article/pii/S0021999111005122
12.2 Publications of the year
International journals
International peer-reviewed conferences
Conferences without proceedings
Edition (books, proceedings, special issue of a journal)
Doctoral dissertations and habilitation theses
Reports & preprints
12.3 Other
Scientific popularization
12.4 Cited publications
- 33 articleIn silico assessment of the effects of various compounds in MEA/hiPSC-CM assays: Modelling and numerical simulations.Journal of Pharmacological and Toxicological Methods892018, 59-72
- 34 articleConvergence of discrete duality finite volume schemes for the cardiac bidomain model.Networks and Heterogeneous Media622011, 195-240
- 35 inproceedingsAn Improved Iterative Pace-Mapping Algorithm to Detect the Origin of Premature Ventricular Contractions.Computing in Cardiology47abstract nr 62, really early. Oral presentation.RiminiComputing in Cardiology2020, 62
- 36 inproceedingsModélisation et simulation de l'électrophysiologie cardiaque à l'échelle microscopique.43e Congrès National d'Analyse Numérique (CANUM)oral presentationSMAIObernai, Alsace, FranceMay 2016, URL: http://smai.emath.fr/canum2016/resumesPDF/peb/Abstract.pdf
- 37 miscTheoretical and Numerical Study of Cardiac Electrophysiology Problems at the Microscopic Scale..PosterJuly 2016
- 38 inproceedingsA Three-Dimensional Computational Model of Action Potential Propagation Through a Network of Individual Cells.Computing in Cardiology 2017Rennes, FranceSeptember 2017, 1-4
- 39 inproceedingsMicroscopic Simulation of the Cardiac Electrophysiology: A Study of the Influence of Different Gap Junctions Models.Computing in CardiologyMaastricht, NetherlandsSeptember 2018
- 40 articleImpact of Septal Radiofrequency Ventricular Tachycardia Ablation; Insights From Magnetic Resonance Imaging.Circulation1302014, 716-718URL: https://www.ahajournals.org/doi/10.1161/CIRCULATIONAHA.114.010175
- 41 inproceedingsDo we need to enforce the homogeneous Neuman condition on the Torso for solving the inverse electrocardiographic problem by using the method of fundamental solution ?Computing in Cardiology 201643Computing in Cardiology 2016Vancouver, CanadaSeptember 2016, 425-428
- 42 articleOptimal monodomain approximations of the bidomain equations used in cardiac electrophysiology.Mathematical Models and Methods in Applied Sciences246February 2014, 1115-1140
- 43 articleA 2D/3D Discrete Duality Finite Volume Scheme. Application to ECG simulation.International Journal on Finite Volumes612009, URL: http://hal.archives-ouvertes.fr/hal-00328251/fr
- 44 articleStability And Convergence Of A Finite Volume Method For Two Systems Of Reaction-Diffusion Equations In Electro-Cardiology.Nonlinear Anal. Real World Appl.742006, 916--935URL: http://hal.archives-ouvertes.fr/hal-00016816/fr
- 45 articleSpatially Coherent Activation Maps for Electrocardiographic Imaging.IEEE Transactions on Biomedical Engineering64May 2017, 1149-1156
- 46 inproceedingsEpicardial Fibrosis Explains Increased Transmural Conduction in a Computer Model of Atrial Fibrillation .Computing in CardiologyVancouver, CanadaSeptember 2016
- 47 articleSudden cardiac arrest associated with early repolarization.N. Engl. J. Med.3582008, 2016--2023
- 48 articleSpontaneous initiation of atrial fibrillation by ectopic beats originating in the pulmonary veins.N. Engl. J. Med.3391998, 659-666URL: https://www.nejm.org/doi/full/10.1056/NEJM199809033391003
- 49 articleMechanism of Right Precordial ST-Segment Elevation in Structural Heart Disease: Excitation Failure by Current-to-Load Mismatch.Heart Rhythm72010, 238-248URL: http://dx.doi.org/10.1016/j.hrthm.2009.10.007
- 50 articleA microstructural model of reentry arising from focal breakthrough at sites of source-load mismatch in a central region of slow conduction.Am. J. Physiol. Heart Circ. Physiol.3062014, H1341-1352
- 51 inproceedingsA new ECG-based method to guide catheter ablation of ventricular tachycardia.iMAging and eLectrical TechnologiesUppsala, SwedenApril 2018
- 52 articlePreconditioning the bidomain model with almost linear complexity.Journal of Computational Physics2311January 2012, 82--97URL: http://www.sciencedirect.com/science/article/pii/S0021999111005122
- 53 articleA Comparison of monodomain and bidomain reaction-diffusion.IEEE Transactions on Biomedical Engineering53122006, 2425-2435URL: http://dx.doi.org/10.1109/TBME.2006.880875
- 54 articleCardiac Anisotropy in Boundary-Element Models for the Electrocardiogram.Medical and Biological Engineering and Computing472009, 719--729URL: http://dx.doi.org/10.1007/s11517-009-0472-x
- 55 inproceedingsAnatomically-induced Fibrillation in a 3D model of the Human Atria.Computing in CardiologyMaastricht, NetherlandsSeptember 2018
- 56 miscRegional conduction slowing can explain inferolateral J waves and their attenuation by sodium channel blockers.PosterSeptember 2016
- 57 inproceedingsFeasibility of Whole-Heart Electrophysiological Models With Near-Cellular Resolution.CinC 2020 - Computing in CardiologyRimini / Virtual, ItalySeptember 2020
- 58 articleScalable and Accurate ECG Simulation for Reaction-Diffusion Models of the Human Heart.Frontiers in Physiology9April 2018, 370
- 59 articleA Cell-Based Framework for Numerical Modeling of Electrical Conduction in Cardiac Tissue.Front. Phys.5This is very very similar to what we were doing with PEB... Looks like we're scooped at least for the approach, but we do have a few abstracts ([becue:cinc17], [becue16a], MMCE meeting in Ottawa November 2017). Note it's in Frontiers in (Biomedical) Physics, not Physiology.2017, 48
