Section: New Results
Adopting new computing paradigms
Participants : David James Sherman [correspondant] , Pascal Durrens, Natalia Golenetskaya, Florian Lajus, Xavier Calcas.
Analyses in comparative genomics are characteristically forms of datamining in high-dimension sets of relations between genes and gene products. For every linear increase in genomic data, these relations can grow at worst geometrically.
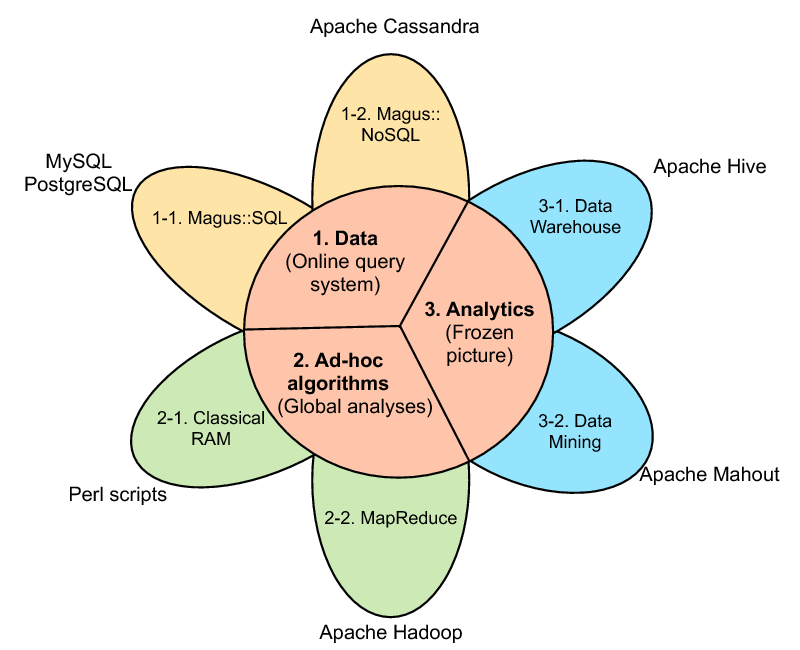
Natalia Golenetskaya's thesis[12] developed an integrated architecture that we call Tsvetok, which combines a novel NoSQL storage schema, domain-specific Map-Reduce algorithms, and existing resources to efficiently handle the fundamentally data-parallel analyses encountered in comparative genomics [48] , [42] , [51] . Tsvetok components are deployed in Magnome 's private cloud and have been extensively tested using data and use cases derived from log analyses of the Génolevures web resource. We designed Map-Reduce solutions for the principal whole-genome analyses used by Magnome for comparative genomics, in particular new distributed algorithms for systematic identification of gene fusion and fission events in eukaryote genomes, and large-scale consensus clustering for protein families. These examples illustrate two strategies that can be used to scale algorithms in a Map-Reduce setting[12] .
-
Converting classical graph-based algorithms with message propagation: instead of traversing a graph, which would incur high latency, information is sent forward in waves, and synchronized later. Some of the intermediate computations may be redundant, but overall running time is minimized.
-
Iterative sampling strategies, which run the standard algorithm on carefully chosen subsets, and later compute a consensus of the intermediate results. The iterations may take some time to converge, but the individual instances can be run within one machine.
|
Florian Lajus extended the Magus software platform to use the NoSQL storage components in Tsvetok, and has validated it on a large collection of fungal genomes. Xavier Calcas is currently integrating the Galaxy platform (http://usegalaxy.org ) with Magus.