Section: New Results
Inter-individual spatio-temporal registration strategies applied to 3D microscopy image sequences of Arabidopsis floral meristems
Participants : Gaël Michelin, Grégoire Malandain.
This work is made in collaboration with Yassin Refahi (Sainsbury Lab., University of Cambridge), Jan Traas (ENS Lyon) and Christophe Godin (Inria Virtual Plants team, Montpellier).
In developmental biology, the study of model organisms such as the plant Arabidopsis thaliana aims at understanding genetic mechanisms responsible of morphogenesis. Today, fluorescent confocal microscopy is a means for in vivo imaging of organs of interest such as Arabidopsis floral meristems at cell level with a high spatio-temporal resolution. To handle such 3D+ image sequences, adapted computer-assisted methods are highly desirable. Moreover, the inter-individual development variability quantification requires the ability to register spatio-temporal image sequences from a population of individuals.
In the related work, we propose a dedicated tool for the inter-individual spatio-temporal sequence-to-sequence registration applied to developing Arabidopsis flower meristems. We also discuss the different strategies that may be adopted by the user for the method application in order to assist the choice of parameters for the registration method such as:
-
the initialization of the image-to-sequence registration optimization process;
-
the initialization "propagation" strategy for sequence-to-sequence registration;
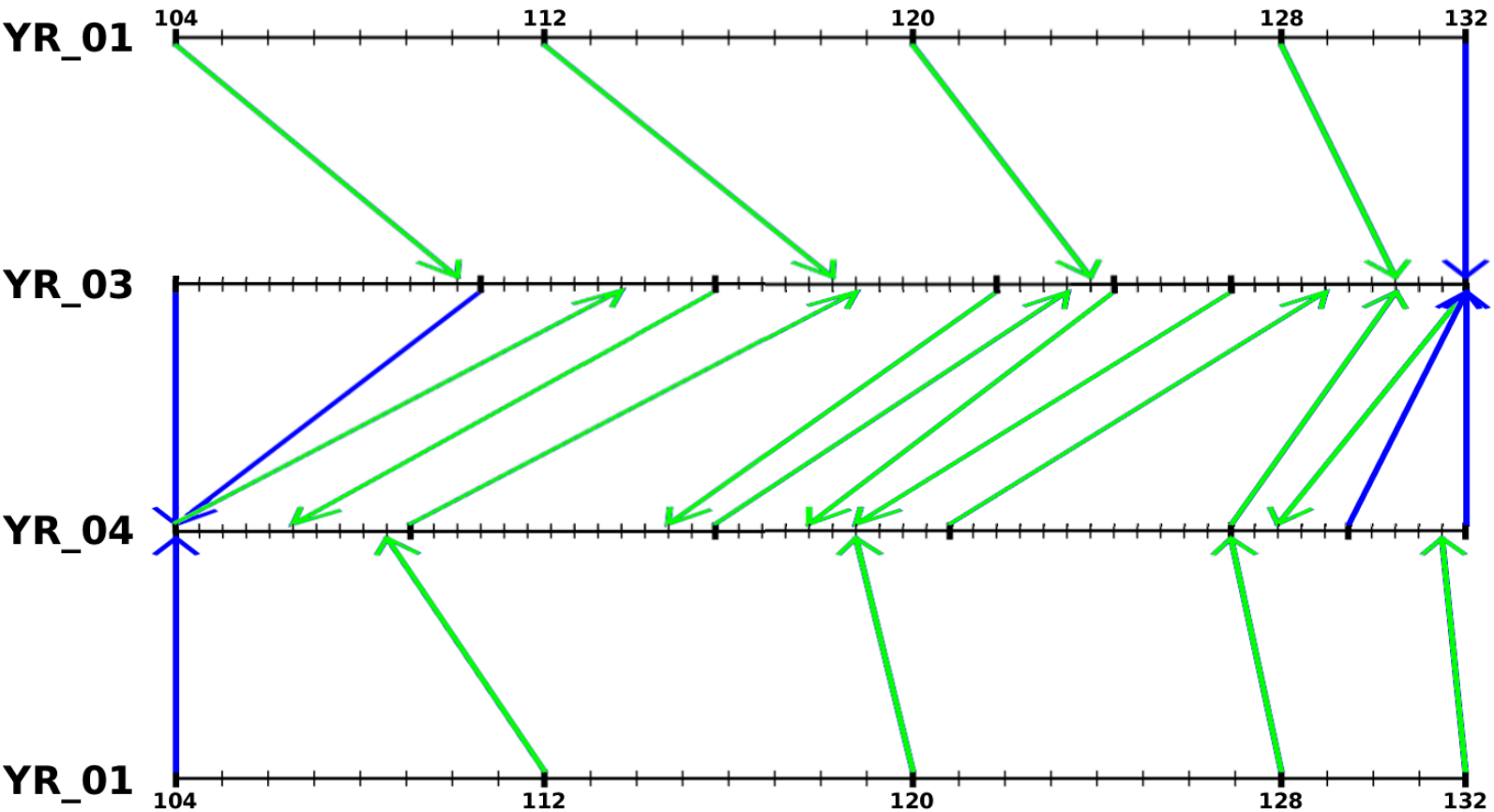
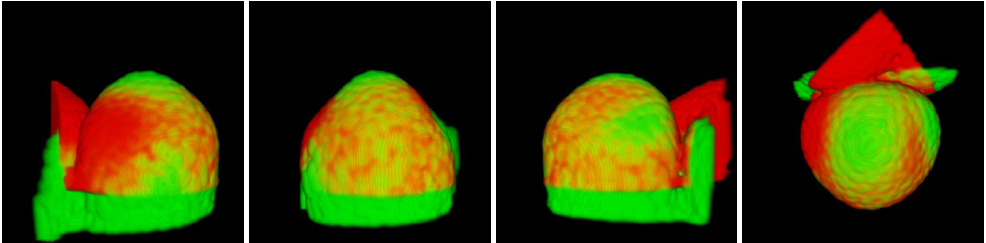
Figure 8 shows the result of the temporal registration between three interpolated image sequences of developing Arabidopsis floral meristems. Figure 9 shows an example of spatial registration between two images from different floral meristems at the same developing stage.