Keywords
Computer Science and Digital Science
- A3.1.2. Data management, quering and storage
- A3.1.3. Distributed data
- A3.1.7. Open data
- A3.1.8. Big data (production, storage, transfer)
- A3.2.4. Semantic Web
- A3.3.3. Big data analysis
- A3.4.1. Supervised learning
- A3.4.2. Unsupervised learning
- A3.4.3. Reinforcement learning
- A3.4.4. Optimization and learning
- A3.4.6. Neural networks
- A3.4.8. Deep learning
- A5.1.4. Brain-computer interfaces, physiological computing
- A5.2. Data visualization
- A5.3.2. Sparse modeling and image representation
- A5.3.3. Pattern recognition
- A5.3.4. Registration
- A5.4.1. Object recognition
- A5.4.6. Object localization
- A5.9.2. Estimation, modeling
- A5.9.4. Signal processing over graphs
- A6.2.3. Probabilistic methods
- A6.2.4. Statistical methods
- A6.3.3. Data processing
- A6.3.4. Model reduction
- A9.2. Machine learning
- A9.3. Signal analysis
Other Research Topics and Application Domains
- B1.2. Neuroscience and cognitive science
- B1.2.1. Understanding and simulation of the brain and the nervous system
- B1.2.2. Cognitive science
- B2.1. Well being
- B2.2.2. Nervous system and endocrinology
- B2.2.6. Neurodegenerative diseases
- B2.5.1. Sensorimotor disabilities
- B2.5.2. Cognitive disabilities
- B2.6.1. Brain imaging
1 Team members, visitors, external collaborators
Research Scientists
- Emmanuel Caruyer [CNRS, Researcher]
- Julie Coloigner [CNRS, Researcher]
- Benoit Combès [Inria, Researcher, from Oct 2021]
- Olivier Commowick [Inria, Researcher, HDR]
- Claire Cury [Inria, Researcher]
- Fanny Degeilh [Inria, Starting Research Position, from Nov 2021]
- Camille Maumet [Inria, Researcher]
Faculty Members
- Pierre Maurel [Team leader, Univ de Rennes I, Associate Professor, HDR]
- Isabelle Bonan [Univ de Rennes I, Professor, HDR]
- Gilles Edan [Univ de Rennes I, Professor]
- Jean-Christophe Ferré [Univ de Rennes I, Professor, HDR]
- Francesca Galassi [Univ de Rennes I, Associate Professor, From Sept 2021]
- Jean-Yves Gauvrit [Univ de Rennes I, Professor, HDR]
- Raphael Truffet [École normale supérieure de Rennes, ATER, from Sep 2021]
Post-Doctoral Fellows
- Francesca Galassi [Univ de Rennes I, until Aug 2021]
- Lou Scotto Di Covella [Inria]
PhD Students
- Thomas Durantel [Univ de Rennes I]
- Mathis Fleury [Inria, until Feb 2021]
- Elodie Germani [Univ de Rennes I, from Oct 2021]
- Stephanie Leplaideur [Centre hospitalier régional et universitaire de Rennes, until Nov 2021]
- Giovanna Orru [Univ de Rennes I, until Jul 2021]
- Caroline Pinte [Univ de Rennes I, from Oct 2021]
- Xavier Rolland [CNRS]
- Jean Charles Roy [Univ de Rennes I, until Sep 2021]
- Raphael Truffet [Univ de Rennes I, until Aug 2021]
Technical Staff
- Hachim Bani [Inria, Ingineer]
- Élise Bannier [Centre hospitalier régional et universitaire de Rennes, Engineer]
- Benoit Combès [Centre hospitalier régional et universitaire de Rennes, Engineer, until Sep 2021]
- Isabelle Corouge [Univ de Rennes I, Engineer]
- Jean Come Douteau [Inria, Engineer, from Jun 2021]
- Quentin Duché [Univ de Rennes I, Engineer, from Nov 2021]
- Quentin Duché [Inria,, Engineer, until Oct 2021]
- Malo Gaubert [Rennes University Hospital, Engineer]
- Renaud Hédouin [Inria, Engineer]
- Nolwenn Jegou [University of Rennes 1, Engineer]
- Florent Leray [Inria, Engineer]
- Julien Louis [Inria, Engineer, until Feb 2021]
- Arthur Masson [Inria, Engineer]
Interns and Apprentices
- Ismail Abaakil [Centre hospitalier régional et universitaire de Rennes, until Jul 2021]
- Tristan Calas [Univ de Rennes I, from May 2021 until Jun 2021]
- Samia Djaiji [Univ de Rennes I, from May 2021 until Jun 2021]
- Nora El Graoui [Centre hospitalier régional et universitaire de Rennes, until Jul 2021]
- Nina Forde [Inria, from Aug 2021]
- Quentin Frecelle [Univ de Rennes I, until Jun 2021]
- Elodie Germani [Univ de Rennes I, until Jul 2021]
- Chloe Mercier [Inria, from Feb 2021 until Jul 2021]
- Etienne Objois [Univ de Rennes I, from May 2021 until Jul 2021]
- Caroline Pinte [Centre hospitalier régional et universitaire de Rennes, from Mar 2021 until Aug 2021]
Administrative Assistant
- Armelle Mozziconacci [CNRS]
Visiting Scientist
- Agustina Fragueiro [University of Studies G. d’Annunzio Chieti Pescara, from Nov 2021]
External Collaborators
- Pierre-Yves Jonin [Centre hospitalier régional et universitaire de Rennes, Neuropsychologist]
- Gabriel Robert [Univ de Rennes I, from Apr 2021, Psychiatrist]
2 Overall objectives
Empenn (means “Brain” in Breton language) ERL U1228 research team is jointly affiliated with Inria, Inserm (National Institute of Health and Scientific Research), CNRS (INS2I institute), and University of Rennes I. It is a team of IRISA/UMR CNRS 6074. Empenn is based in Rennes, at both the medical and science campuses. The team follows the “VisAGeS” one that was created for 12 years in 2006 by Inria, As for “VisAGeS”, Empenn hosts the accreditation number U1228 renewed by Inserm in 2017, after a competitive evaluation conducted by both HCERES and Inserm.
Through this unique partnership, the ambition of Empenn is to establish a multidisciplinary team bringing together researchers in information sciences and medicine. Our medium- and long-term objective is to introduce our basic research to clinical practice, while maintaining the excellence of our methodological research.
Our goal is to foster research in medical imaging, neuroinformatics and population cohorts. In particular, the Empenn team targets the detection and development of imaging biomarkers for brain diseases and focus its efforts on translating this research to clinics and clinical neurosciences at large.
In particular, the objective of Empenn is to propose new statistical and computing methods, and to measure and model brain morphological, structural and functional states in order to better diagnose, monitor and deliver treatment for mental, neurological and substance use disorders. We propose combining advanced instrumental devices and new computational models to provide advanced diagnosis, therapeutic and neuro-rehabilitation solutions for some of the major disorders of the developing and aging brain.
Generic and challenging research topics in this broad domain include finding new ways to compare models and data, assist decisions and interpretation, and develop feedback from experiments. These activities are performed in close collaboration with the Neurinfo in vivo imaging platform, which is a critical environment for the experimental implementation of our research on challenging clinical research projects and the development of new clinical applications.
3 Research program
3.1 Glossary
-
Magnetic Resonance Imaging
- MR - Magnetic Resonance
- MRI - Magnetic Resonance Imaging
- fMRI - Functional Magnetic Resonance Imaging
- DWI - Diffusion-Weighted Imaging
- ASL - Arterial Spin Labeling
-
Other modalities
- PET - Positron Emission Tomography
- EEG - Eletroencephalograpy
- NIRS - Near InfraRed Spectroscopy
-
Medical terminology
- MS - Multiple Sclerosis
- TBI - Traumatic Brain Injury
-
Methodological terminology
- GLM - General Linear Model
- MCM - Multi-compartment models
3.2 Scientific Foundations
The scientific foundations of our team concern the design and development of new computational solutions for biological images, signals and measurements. Our objective is to develop a better understanding of the normal and pathological brain, at different scales.
This includes imaging brain pathologies in order to better understand pathological behavior from the organ level to the cellular level, and even to the molecular level (PET-MR imaging), and the modelling of normal and pathological large groups of individuals (cohorts) from image descriptors. It also includes the challenge of the discovery of episodic findings (i.e. rare events in large volumes of images and data), data mining and knowledge discovery from image descriptors, the validation and certification of new drugs from imaging features, and, more generally, the integration of neuroimaging into neuroinformatics through the promotion and support of virtual organizations of biomedical actors by means of e-health technologies.
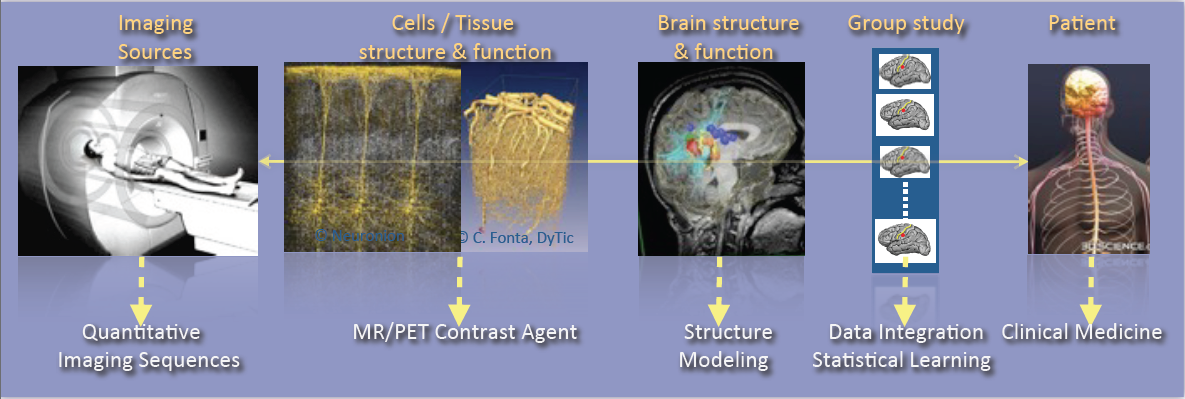
As shown in Figure 1, the research activities of the Empenn team closely link observations and models through the integration of clinical and multiscale data, and phenotypes (cellular, and later molecular, with structural or connectivity patterns in the first stage). Our ambition is to build personalized models of central nervous system organs and pathologies, and to compare these models with clinical research studies in order to establish a quantitative diagnosis, prevent the progression of diseases and provide new digital recovery strategies, while combining all these research areas with clinical validation. This approach is developed within a translational framework, where the data integration process to build the models is informed by specific clinical studies, and where the models are assessed regarding prospective clinical trials for diagnosis and therapy planning. All of these research activities are conducted in close collaboration with the Neurinfo platform, which benefited in 2018 from a new high-end 3T MRI system dedicated to research (3T Prisma™ system from Siemens), and through the development in the coming years of multimodal hybrid imaging (from the currently available EEG-MRI, to EEG-NIRS and PET-MRI in the future).
In this context, some of our major developments and newly arising issues and challenges will include:
- The generation of new descriptors to study brain structure and function (e.g. the combination of variations in brain perfusion with and without a contrast agent; changes in brain structure in relation to normal, pathological, functional or connectivity patterns; or the modeling of brain state during cognitive stimulation using neurofeedback).
- The integration of additional spatiotemporal and hybrid imaging sequences covering a larger range of observations, from the molecular level to the organ level, via the cellular level (arterial spin labeling, diffusion MRI, MR relaxometry, MR fingerprinting, MR cell labeling imaging, MR-PET molecular imaging, EEG-MRI functional imaging, EEG-NIRS-MRI, etc.).
- The creation of computational models through the data fusion of molecular, cellular (i.e. through dedicated ligands or nanocarriers), structural and functional image descriptors from group studies of normal and/or pathological subjects.
- The evaluation of these models in relation to acute pathologies, especially for the study of degenerative, psychiatric, traumatic or developmental brain diseases (primarily multiple sclerosis, stroke, traumatic brain injury (TBI) and depression, but applicable with a potential additional impact to epilepsy, Parkinson’s disease, dementia, Posttraumatic stress disorder, etc.) within a translational framework.
In terms of new major methodological challenges, we will address the development of models and algorithms to reconstruct, analyze and transform the images, and to manage the mass of data to store, distribute and “semanticize” (i.e. provide a logical division of the model’s components according to their meaning). As such, we expect to make methodological contributions in the fields of model inference; statistical analysis and modeling; the application of sparse representation (compressed sensing and dictionary learning) and machine learning (supervised/unsupervised classification and discrete model learning); data fusion (multimodal integration, registration, patch analysis, etc.); high-dimensional optimization; data integration; and brain-computer interfaces. As a team at the frontier between the digital sciences and clinical research in neuroscience, we do not claim to provide theoretical breakthroughs in these domains but rather to provide significant advances in using these algorithms through to the advanced applications we intend to address. In addition, we believe that by providing these significant advances using this set of algorithms, we will also contribute to exhibiting new theoretical problems that will fuel the domains of theoretical computer sciences and applied mathematics.
In summary, we expect to address the following major challenges:
- Developing new information processing methods able to detect imaging biomarkers in the context of mental, neurological, and substance use disorders.
- Providing new computational solutions for our target applications, allowing a more appropriate representation of data for image analysis and the detection of biomarkers specific to a form or grade of pathology, or specific to a population of subjects.
- Providing, for our target applications, new patient-adapted connectivity atlases for the study and characterization of diseases from quantitative MRI.
- Providing, for our target applications, new analytical models of dynamic regional perfusion, and deriving indices of dynamic brain local perfusion from normal and pathological populations.
- Investigating whether the theragnostics paradigm of rehabilitation from hybrid neurofeedback can be effective in some behavioral and disability pathologies.
These major advances will be primarily developed and validated in the context of several priority applications in which we expect to play a leading role: multiple sclerosis, stroke rehabilitation, and the study and treatment of depression.
4 Application domains
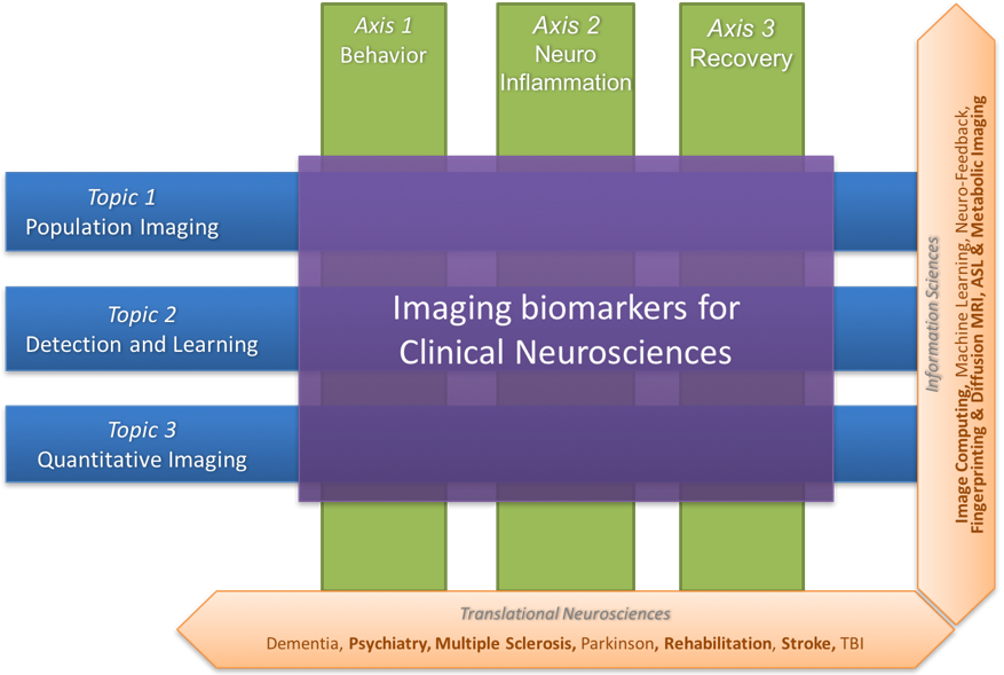
Figure 2 summarizes the scientific organization of the research team through three basic research topics in information sciences (Population Imaging, Detection and Learning, and Quantitative Imaging) and three translational axes on central nervous system diseases (Behavior, Neuro-inflammation and Recovery).
4.1 Basic research
4.1.1 Population imaging
One major objective of neuroimaging researchers and clinicians is to be able to stratify brain imaging data in order to derive new and more specific population models. In practice, this requires to set up large-scale experiments that, due to the lack of resources and capabilities to recruit locally subjects who meet specific inclusion criteria, motivates the need for sharing the load.
But, building and using multi-site large-scale resources pose specific challenges to deal with the huge quantity of data produced and their diversity. Empenn will focus on two challenges in particular:
- Providing computational environments for the computation and use of imaging biomarkers in the targeted brain diseases, a solution to be used by radiologists and neurologists/psychiatrists for the clinical follow-up of a large patient population.
- Modeling analytic variability of image processing pipelines to better understand and predict the behaviour of imaging biomarker detection solutions and improve reproducibility and productivity in clinical neuroimaging research.
4.1.2 Detection and learning
We intend to make significant contributions with major impacts in learning coupling models between functional recordings during neurofeedback procedures. These advances will provide a breakthrough in brain-computer interfaces for rehabilitation protocols. Our aim is to:
- Provide a computational environment that combines data-driven (machine learning) and Bayesian solutions to improve the detection of abnormal patterns in images through decision or evidence theory data fusion strategies. The major initial application will be for multiple sclerosis. Over the longer term, we also expect to adapt these methods to address a wider range of neurological diseases (epilepsy, stroke, tumors, etc.) in neonate and adult brains.
- Develop solutions for combining brain state measurements from multimodal sensors or sequences (e.g. fMRI, ASL, EEG, NIRS, etc.) with applications in the spatiotemporal reconstruction of brain activity from MRI-EEG or the combined detection of the endogenous hemodynamic and resting state network of the brain from ASL and NIRS. Over the longer term, the advent of new hybrid brain imaging sensors (e.g. PET-MRI) will require these methods to be extended to a larger spectrum of information combining structural, morphological, metabolic, electrophysiological and cellular/molecular information (e.g. through the use of specific ligands/nanocarriers).
4.1.3 Quantitative imaging
The Empenn research group focuses on the development of several quantitative techniques in magnetic resonance imaging of the brain. These methods allow for characterization of both the function and the structure of the brain with high precision. Arterial spin labelling (ASL) is a contrast agent-free imaging technique which labels arterial blood water as an endogenous tracer for perfusion and can measure resting-state cerebral blood flow. We are interested in estimating multiparametric hemodynamics using ASL, such as combined cerebral blood flow and arterial transit times, and derive statistical descriptors to represent significant differences between groups. In addition to quantitative perfusion parameters, our contributions on tissue compartment imaging aim at delineating neural circuits and characterize their microstructure properties, using both diffusion MRI and relaxometry. In diffusion MRI, arbitrary gradient waveforms were shown to exhibit higher sensitivity to microstructure parameters than standard pulsed gradients. We work on the optimization of sampling protocols in this domain, with the objective to propose sequences compatible with in vivo acquisition. Complementary to diffusion MRI, we develop methods for the reconstruction of myelin-bound, extra-axonal and cerebrospinal fluid water using multi-compartment modelling of the T2-relaxometry signal. We combine these techniques with tractography to identify trajectories of pathologies associated to the evolution of these microstructural parameters along specific fiber bundles in the brain white matter.
4.2 Translational resarch
4.2.1 Behavior
Advances in the field of in vivo imaging offer new opportunities for addressing the management of resistant affective disorders and their consequences (suicide risk and socio-professional impact), and the management of spatial cognition disorders after stroke and their consequences (postural perturbations and the loss of autonomy). Our objective, and the main challenge in this context, will be to introduce medical image computing methods to the multidisciplinary field of behavioral disorders (cognitive disorders, particularly spatial and postural control disorders or anterograde memory impairment, mood disorders, notably resistant depression, schizophrenic disorders, pervasive developmental disorders, attention disorders, etc.) in order to gain a better understanding of the pathology and devise innovative therapeutic approaches.
We also expect to become a major player in the future and make important contributions with significant impacts, primarily in drug-resistant depression in young and old populations. In particular, we expect to provide new image-related metrics combining perfusion, metabolism and microstructural information regarding the brain in order to better characterize pathologies, provide prospective evolution values and potentially provide new brain stimulation targets that could be used in neurofeedback rehabilitation protocols or other types of brain stimulation procedures.
We aim to provide new imaging markers of mental diseases, especially in the context of mood disorders. The new biomarkers will be derived from the metabolic (ASL and later ASL+PET) point of view as well as from the microstructural point of view (multicompartment diffusion MRI and relaxometry). Similarly, we expect to exhibit imaging biomarker regularities combining metabolic and structural information. Over the longer term, we expect these biomarkers to be the target of neurofeedback rehabilitation procedures. Also, over the longer term, we expect to supplement the MRI markers with molecular markers coming from new PET tracers, especially those associated with serotonin intake, at one time point or during a rehabilitation protocol under hybrid PET-EEG-MRI neurofeedback procedures.
4.2.2 Neuroinflammation
Some of the major ongoing research issues regarding neuroimaging of neuro-inflammatory diseases concern the definition of new biomarkers to track the development of the pathology using high- dimensional data (e.g. nD+t MRI). This includes the use of white matter-specific imaging, such as magnetization transfer MRI, relaxometry and diffusion-weighted imaging (DW-MRI). Our objective is (1) to develop information-processing tools to tag the spatiotemporal evolutions of Multiple Sclerosis patterns at the brain parenchyma and spinal cord levels from their different signatures (inflammatory cells visible with USPIO or Gd contrast agents on MRI, persistent black holes, eloquent regional atrophy and microstructure signatures); and (2) to test these new tools on new imaging cohorts. In this respect, we for instance conduct studies on brain and spinal cord imaging, continuing on from the PHRC multicentric EMISEP project (PI: G. Edan), as it is very likely that lesions in the spine will directly affect the ambulatory ability of the patient (and thereby the clinical scores). In order to extend this experiment to a larger MS population, based on our expertise from the OFSEP cohort, we also plan to improve the MS therapeutic decision process through the MUSIC project (Multiple Sclerosis Imaging Check out, a public/private project). Our goal is to develop and assess a standardized monitoring tool that provides a robust, long-term computerized MRI follow-up that will become the gold standard in clinical practice for therapeutic decisions in MS treatment. As part of this project, Empenn will share its expertise in data management systems (Shanoir and FLI-IAM) and automatic processing tools (through the medInria and Anima software repositories) to extract quantitative indices from the images.
4.2.3 Recovery
Mental and neurological disorders are the leading cause of years lived with a disability. Treatment-resistant depression affects approximately 2% of the European population. Meanwhile, in the case of brain disorders, almost 1.5 million Europeans (15 million people worldwide) suffer a stroke event each year. Current recovery methods for brain disorders and traumatic brain injuries remain limited, preventing many from achieving full recuperation. We propose to address the issue of brain recovery by introducing new advances from recent breakthroughs in computational medical imaging, data processing and human-machine interfaces, and demonstrate how these new concepts can be used, in particular for the treatment of stroke and major depressive disorders.
We ambition to combine advanced instrumental devices (hybrid EEG, NIRS and MRI platforms), with new hybrid brain computer interface paradigms and new computational models to provide neurofeedback-based therapeutic and neuro-rehabilitation paradigms in some of the major mental and neurological disorders of the developmental and aging brain.
Neurofeedback involves using a brain-computer interface that provides an individual with real-time biofeedback about his or her brain activity in the form of sensory feedback. It enables individuals to learn to better control their brain activity, which can be measured in real time using various non-invasive sensors as described above. Although EEG is currently the only modality used by clinical practitioners in that context, it lacks specificity due to its low spatial resolution. Dynamic research into fMRI-neurofeedback has held promise for treating depression, chronic pain and stroke, since it offers the prospect of real-time imagery of the activity in deep brain structures with high spatial resolution. However, the low temporal resolution and high cost of fMRI-neurofeedback has hampered the development of many applications. We believe that the future belongs to hybrid responses that combine multimodal sensors and intend to demonstrate this in the Empenn project.
5 Social and environmental responsibility
Francesca Galassi is part of the Women in MICCAI (WiM) Committee since Oct 2021 - until 2023. The mission of the WiM Committee is to strengthen and widen the representation of female scientists in the MICCAI community by pursuing policies that encourage more female participation in the field and ensure fair and equitable career promotion for female faculty and students - policies that assist in overcoming implicit gender bias within the community. WiM’s activities include: the coordination with the organizers of the annual MICCAI conferences, the organization of networking events within the community particularly at the MICCAI conference on an annual basis, the development and maintenance of online discussion and promotion platforms (e.g. Twitter, Facebook, LinkedIn, Google Groups), and interfacing and advising the MICCAI board on related matters.
6 Highlights of the year
In 2021 :
- Mathis Fleury successfully defended his PhD thesis.
- Raphaël Truffet successfully defended his PhD thesis.
- Pierre Maurel successfully defended his HDR.
- Benoit Combès has been recruited as permanent Inria researcher.
- Francesca Galassi has been recruited as MCF of UR1.
- Quentin Duché has been recruited as research engineer of UR1.
- Fanny Degeilh won an "Inria Starting Research Position - Call 2021" in June 2021.
- An international challenge on Multiple Sclerosis new lesions segmentation was organized by the team.
- An international challenge on Diffusion-weighted MRI reconstruction was organized by the team.
7 New software and platforms
7.1 New software
7.1.1 Anima
-
Keywords:
Filtering, Medical imaging, Diffusion imaging, Registration, Relaxometry
-
Scientific Description:
Anima is a set of libraries and tools developed by the team as a common repository of research algorithms. As of now, it contains tools for image registration, statistical analysis (group comparison, patient to group comparison), diffusion imaging (model estimation, tractography, etc.), quantitative MRI processing (quantitative relaxation times estimation, MR simulation), image denoising and filtering, and segmentation tools. All of these tools are based on stable libraries (ITK, VTK), making it simple to maintain.
-
Functional Description:
Anima is a set of libraries and tools in command line mode for processing and analysing medical images.
- URL:
-
Contact:
Olivier Commowick
-
Participants:
Aymeric Stamm, Fang Cao, Florent Leray, Guillaume Pasquier, Laurence Catanese, Olivier Commowick, Renaud Hedouin, René-Paul Debroize
7.1.2 MedINRIA
-
Keywords:
Visualization, DWI, Health, Segmentation, Medical imaging
-
Scientific Description:
MedInria aims at creating an easily extensible platform for the distribution of research algorithms developed at Inria for medical image processing. This project has been funded by the D2T (ADT MedInria-NT) in 2010, renewed in 2012. A fast-track ADT was awarded in 2017 to transition the software core to more recent dependencies and study the possibility of a consortium creation.The Empenn team leads this Inria national project and participates in the development of the common core architecture and features of the software as well as in the development of specific plugins for the team's algorithm.
-
Functional Description:
MedInria is a free software platform dedicated to medical data visualization and processing.
- URL:
-
Contact:
Olivier Commowick
-
Participants:
Maxime Sermesant, Olivier Commowick, Théodore Papadopoulo
-
Partners:
HARVARD Medical School, IHU - LIRYC, NIH
7.1.3 autoMRI
-
Keywords:
FMRI, MRI, ASL, FASL, SPM, Automation
-
Scientific Description:
This software is highly configurable in order to fit a wide range of needs. Pre-processing includes segmentation of anatomical data, as well as co-registration, spatial normalization and atlas building of all data types. The analysis pipelines perform either within-group analysis or between-group or one subject-versus-group comparison, and produce statistical maps of regions with significant differences. These pipelines can be applied to structural data to exhibit patterns of atrophy or lesions, to ASL (both pulsed or pseudo-continuous sequences) data to detect perfusion abnormalities, to functional data - either BOLD or ASL - to outline brain activations related to block or event-related paradigms. New functionalities have been implemented to facilitate the management and processing of data coming from complex projects.
-
Functional Description:
AutoMRI is based on MATLAB and the SPM12 toolbox and provides complete pipelines to pre-process and analyze various types of images (anatomical, functional, perfusion).
- URL:
-
Contact:
Isabelle Corouge
-
Participants:
Camille Maumet, Elise Bannier, Isabelle Corouge, Pierre Maurel, Quentin Duché, Julie Coloigner
7.1.4 ShanoirUploader
-
Name:
ShanoirUploader (SHAring NeurOImaging Resources Uploader)
-
Keywords:
Webservices, PACS, Medical imaging, Neuroimaging, DICOM, Health, Biology, Java, Shanoir
-
Scientific Description:
ShanoirUploader is a desktop application on base of JavaWebStart (JWS). The application can be downloaded and installed using an internet browser. It interacts with a PACS to query and retrieve the data stored on it. After this ShanoirUploader sends the data to a Shanoir server instance in order to import these data. This application bypasses the situation, that in most of the clinical network infrastructures a server to server connection is complicated to set up between the PACS and a Shanoir server instance.
-
Functional Description:
ShanoirUploader is a Java desktop application that transfers data securely between a PACS and a Shanoir server instance (e.g., within a hospital). It uses either a DICOM query/retrieve connection or a local CD/DVD access to search and access images from a local PACS or the local CD/DVD. After having retrieved the data, the DICOM files are locally anonymized and then uploaded to the Shanoir server. A possible integration of a hash creation application for patient identifiers is provided as well. The primary goals of that application are to enable mass data transfers between different remote server instances and therefore reduce the waiting time of the users, when importing data into Shanoir. Most of the time during import is spent with data transfers.
- URL:
-
Contact:
Michael Kain
-
Participants:
Christian Barillot, Inès Fakhfakh, Justine Guillaumont, Michael Kain, Yao Chi
7.1.5 Shanoir-NG
-
Name:
Shanoir-NG (SHAring NeurOImaging Resources - Next Generation)
-
Keywords:
Neuroimaging, DICOM, Nifti
-
Functional Description:
Shanoir-NG is a complete technological remake of the first version of the Shanoir application, but maintaining the key concepts of Shanoir.
Why did we take this big effort to implement Shanoir-NG from scratch? • Over the years of the existence and usage of Shanoir the technological basis of Shanoir has become outdated and most of its original technical frameworks (JBoss 4, Java Server Faces (JSF), Richfaces, JBoss Seam) are not supported and maintained anymore. • Furthermore the architectures and technologies for developing web applications have dynamically progressed in the last 5 years. The arrival of Single-Page-Applications (SPAs), like Gmail and Twitter, the Docker containerization technology and microservices architectures have dramatically changed the way we develop web applications today. • This lead to the consequence, that only migrating Shanoir to newer versions of the existing libraries and code, was far from being sufficient to extend the lifetime and long-time usage of Shanoir. That is why we started to develop Shanoir-NG from scratch with a new architecture (microservices and REST) and new technologies, while keeping most of its functionalities.
Shanoir-NG (SHAring NeurOImaging Resources) is an open-source neuroinformatics platform designed to share, archive, search and visualize neuroimaging data.
It provides a user-friendly secure web access and offers an intuitive workflow to facilitate the collecting and retrieving of neuroimaging data from multiple sources and a wizzard to make the completion of metadata easy. Shanoir-NG comes along many features such as anonymization of data (based on standard profiles), support for multi-centric clinical studies on subjects or group of subjects.
Shanoir-NG offers an ontology-based data organization (OntoNeuroLOG). Among other things, this facilitates the reuse of data and metadata, the integration of processed data and provides traceability trough an evolutionary approach. Shanoir-NG allows researchers, clinicians, PhD students and engineers to undertake quality research projects with an emphasis on remote collaboration. As a secured Jakarta EE web application, it therefore allows you safely storing and archiving, with no more requirements than a computer with an internet connection!
Shanoir-NG has been extended for preclinical data too, it manages your study meta-data and preclinical images: • Pathology models, therapies, anesthetics and physiological data • Imports Bruker file format
Furthermore, Shanoir-NG is not only a web application: it is also a complete neuroinformatics platform in which you can easily integrate your existing processing tools or develop your own ones: see ShanoirTk or ShanoirUploader to import your data directly from the PACS in the hospital.
Using cross-data navigation and advanced search criteria (new Solr search module), the user scan quickly point to a subset of data to be downloaded. Client side applications have as well been developed to illustrate how to locally access and exploit data though the available web services. With regards to security, the system requires authentication and user rights are adjustable for each hosted study. A study responsible can thereby define the users allowed to see, download or import data into his study or simply make it public.
Shanoir-NG serves neuroimaging researchers in organizing efficiently their studies while cooperating with other laboratories. By managing patient privacy, Shanoir allows the exploitation of clinical data in a research context. It is finally a handy solution to publish and share data with a broader community.
It supports the following formats: DICOM (MR, CT, PT, NM), processed datasets (NIfTI), Bruker, EEG(BrainVision/EDF), big zip files
-
News of the Year:
Shanoir-NG is a complete technological remake of the first version of the Shanoir application, but maintaining the key concepts of Shanoir.
- URL:
-
Contact:
Michael Kain
-
Participants:
Christian Barillot, Mathieu Simon, Michael Kain, Yao Chi, Aneta Morawin, Arnaud Touboulic, Inès Fakhfakh, Anthony Baire
-
Partners:
CHU Grenoble, INSERM, CNRS, Université Grenoble Alpes, Université de Strasbourg
7.1.6 LongiSeg4MS
-
Name:
Longitudinal Segmentation For Multiple Sclerosis
-
Keywords:
3D, Brain MRI, Deep learning, Detection
-
Functional Description:
LongiSeg4MS is an automatic new multiple sclerosis (MS) lesion detection tool based on longitudinal data and using deep learning. The system uses FLAIR, T1 or T2 modalities, or a combination of those. The input is 2, 4 or 6 images (2 FLAIR, 2 FLAIR and 2 T1, etc.), a set of modalities for each time point, and outputs a segmentation map describing the location of new MS lesions.
- URL:
-
Contact:
Arthur Masson
-
Partner:
OFSEP
7.1.7 Anima medInria plugins
-
Keywords:
IRM, Medical imaging, Diffusion imaging
-
Functional Description:
Plugins for the medInria software based on the open source software Anima developed in the Visages / Empenn team. These plugins are interfaces between anima and medinria allowing to use Anima functionalities within the clinical user interface provided by medInria. The current functionalities included in the plugins are right now: image registration, denoising, quantitative imaging (relaxometry), and model estimation and visualization from diffusion imaging.
- URL:
-
Contact:
Olivier Commowick
-
Participants:
Olivier Commowick, René-Paul Debroize, Guillaume Pasquier
7.2 New platforms
7.2.1 The Neurinfo Platform
Empenn is the founding actor of an experimental research platform which was installed in August 2009 at the University Hospital of Rennes. The University of Rennes 1, Inria, CNRS for the academic side, and the University Hospital of Rennes and the Cancer Institute “Eugene Marquis” for the clinical side, are partners of this neuroinformatics platform called Neurinfo (https://www.neurinfo.org). Concerning the Neurinfo Platform, the activity domain is a continuum between methodological and technological research built around specific clinical research projects. On the medical field, the translational research domain mainly concerns medical imaging and more specifically the clinical neurosciences. Among them are multiple sclerosis, epilepsy, neurodegenerative, neurodevelopmental and psychiatric diseases, surgical procedures of brain lesions, neuro-oncology and radiotherapy planning. Beyond these central nervous system applications, the platform is also open to alternative applications. Neurinfo ambitions to support the emergence of research projects based on their level of innovation, their pluri-disciplinarity and their ability to foster collaborations between different actors (public and private research entities, different medical specialties, different scientific profiles). In this context, a research 3T MRI system (Siemens Verio) was acquired in summer 2009 in order to develop the clinical research in the domain of morphological, functional, structural and cellular in-vivo imaging. A new 3T Siemens Prisma MRI scanner was installed at the Neuroinfo platform in February 2018. In 2014, an equipment for simultaneous recording of EEG and MRI images was acquired from Brain Product. In 2015, a mock scanner for experimental set-up was acquired as well as a High Performance Computing environment made of one large computing cluster and a data center that is shared and operated by the Inria center and IRISA (UMR CNRS 6074). The computation cluster (480 cores) and the data center (up to 150 TB) are dedicated to host and process imaging data produced by the Neurinfo platform, but also by other research partners that share their protocols on the Neurinfo neuroinformatics system (currently more than 60 sites).
In 2019, an MRI and EEG-compatible fNIRS system was acquired through a co-funding from the INS2I institute of CNRS and FEDER. At the end of 2019, GIS IBISA awarded the Neurinfo platform with a complementary funding that will be dedicated to supplement the current system with additional sensors (from 8x8 optodes to 16x16 optodes). In 2021, the Regional Council of Britanny granted the platform a funding to provide engineer support for one year to develop and integrate this new imaging system.
8 New results
8.1 Basic research
8.1.1 Population imaging
Population imaging is fundamental when it comes to evaluate clinical biomarkers. The team contributions on this aspect are summarised in this section. We studied how analytical variability can impact fMRI results and proposed recommendations and neuroinformatics models to describe the data. We also maintained our clinical interest regarding several pathologies by exploring brain function and connectivity. Also, technical recommendations regarding multicentric imaging protocols were proposed.
Consensus-based guidance for conducting and reporting multi-analyst studies
Participants: Emmanuel Caruyer.
Any large dataset can be analyzed in a number of ways, and it is possible that the use of different analysis strategies will lead to different results and conclusions. One way to assess whether the results obtained depend on the analysis strategy chosen is to employ multiple analysts and leave each of them free to follow their own approach. In this work, we presented consensus-based guidance for conducting and reporting such multi-analyst studies, and we discussed how broader adoption of the multi-analyst approach has the potential to strengthen the robustness of results and conclusions obtained from analyses of datasets in basic and applied research. Associated publication: 10.
Structural and functional interplay in anxiety related classification: a graph signal processing approach
Participants: Julie Coloigner, Pierre Maurel.
Anxiety disorders are one of the most common mental health conditions with a high rate of everyday life disability. Connectivity is steadily gaining relevance to increase our knowledge of psychiatric diseases. Graph signal processing (GSP) is a new framework to integrate structural connectivity and brain function. We propose here a graph-based analysis using GSP metrics and classification procedure, to identify anxiety biomarkers. Results suggest that the joint consideration of structure-function features improves their discriminatory accuracy, and our understanding of the pathophysiology of anxiety. Associated publication: 41.
Isolating the sources of pipeline-variability in group-level task-fMRI results
Participants: Camille Maumet.
While the development of tools and techniques has broadened our horizons for comprehending the complexities of the human brain, a growing body of research has highlighted the pitfalls of such methodological plurality. In a recent study, we found that the choice of software package used to run the analysis pipeline can have a considerable impact on the final group-level results of a task-fMRI investigation (Bowring et al., 2019, BMN ). In this study we revisited our work, seeking to identify the stages of the pipeline where the greatest variation between analysis software is induced. We carried out further analyses on the three datasets evaluated in BMN, employing a common processing strategy across parts of the analysis workflow and then utilizing procedures from three software packages (AFNI, FSL and SPM) across the remaining steps of the pipeline. We used quantitative methods to compare the statistical maps and isolated the main stages of the workflow where the three packages diverge. Across all datasets, we found that variation between the packages’ results was largely attributable to a handful of individual analysis stages, and that these sources of variability were heterogeneous across the datasets (e.g. choice of first-level signal model had the most impact for the ds000001 dataset, while first-level noise model was more influential for ds000109 dataset). We also observed areas of the analysis workflow where changing the software package causes minimal differences in the final results, finding that the group-level results were largely unaffected by which software package is used to model the low-frequency fMRI drifts. Associated publications: 13, 43.
This work was done in collaboration with Alex Bowring and Tom Nichols from the University of Oxford.
Towards efficient fMRI data re-use: can we run between-group analyses with datasets processed differently?
Participants: Xavier Rolland, Pierre Maurel, Camille Maumet.
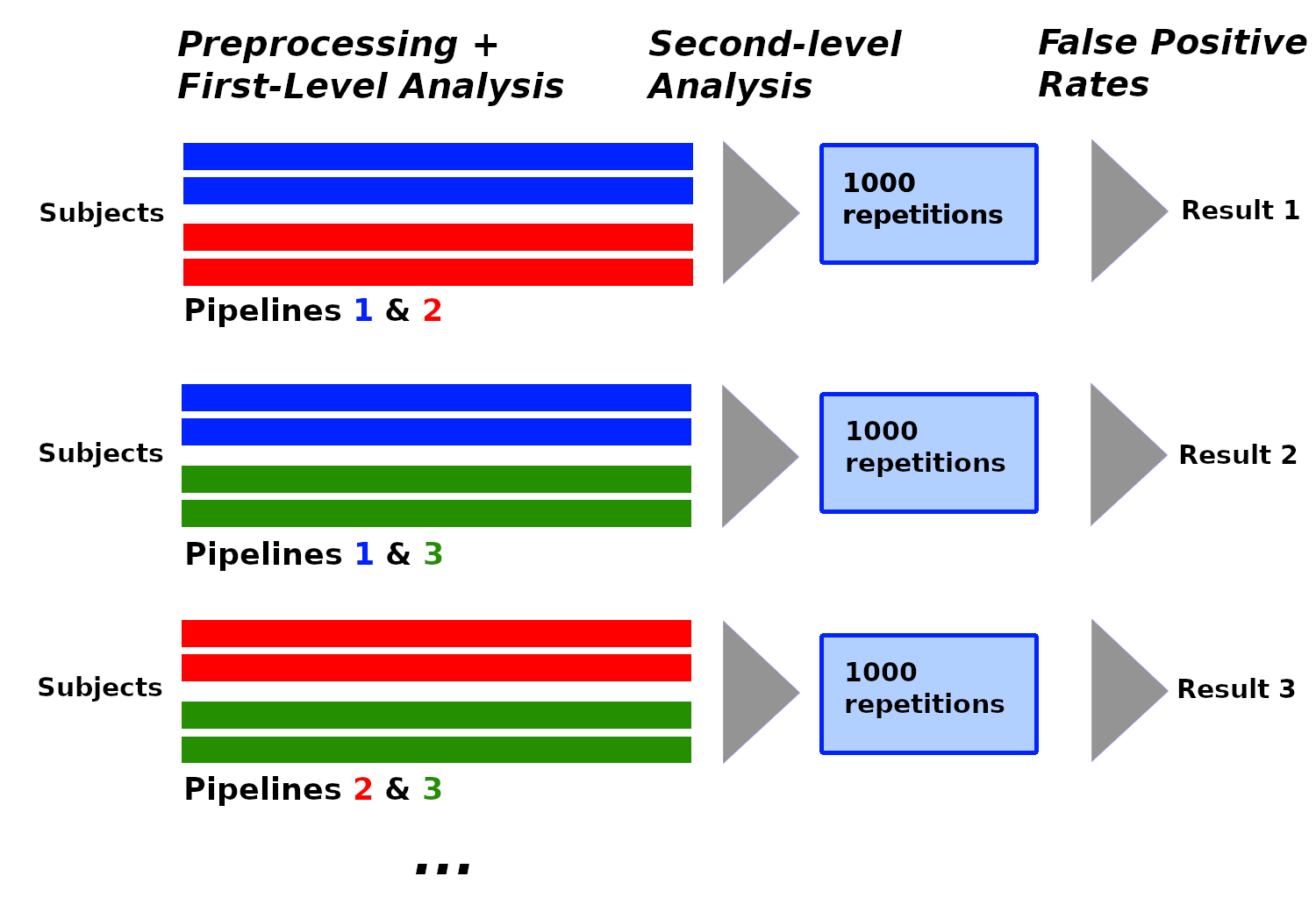
In recent years, the lack of reproducibility of research findings has become an important source of concerns in many scientific fields, including functional Magnetic Resonance Imaging (fMRI). The low statistical power often observed in fMRI studies was identified as one of the leading causes of irreproducibility. The development of data sharing in the field of neuroimaging opens up new opportunities to achieve larger sample sizes by reusing existing data. FMRI studies use subject data processed with pipelines, and although most shared datasets currently include raw data, we may expect to see an increasing proportion of processed data among shared subject data in the future, for privacy and sustainability reasons. Pipelines consist of multiple steps, each with multiple possible methodological choices: this existing variability in terms of processing and analysis (analytical variability) has an impact on the results.
We investigated the impact of analytical variability when combining subject data processed differently in between-group analyses. We created a set of pipelines for subject-level processing that we applied on data from the Human Connectome Project (n=1080). We then performed between-group analyses with subject data processed with different pipelines, under the null hypothesis (making any detection a false positive) using two different software packages: SPM and FSL. We compared the estimated false positive rates obtained to the nominal false positive rate.
We found that the analytical variability induced by the studied parameters was found to be acceptable for some of these analyses and redhibitory for others. We concluded that different processed subject data cannot be combined without taking into account the processing applied on these data.
Recommendations for the implementation of multicenter studies with MRI
Participants: Elise Bannier.
MRI is becoming increasingly important in clinical research studies, both in terms of inclusion criteria and endpoints. Markers based on MRI acquisitions are now regularly considered as primary endpoints. Especially in multicenter studies, the management of MRI acquisitions has to consider the heterogeneity of MRI systems in terms of manufacturers, magnetic fields, receiving coils and software versions. This is due to the number of important parameters that can be specified to obtain an MRI acquisition, the variability between the solutions proposed by the manufacturers and the regular technical innovations. The objective is to detail the specificities to be taken into account and the people to be involved in order to carry out an MRI study whatever the organ concerned by the imaging. Based on the experience and expertise of the members of the Réseau d'entraide multicentrique en IRM (REMI), we proposed detailed recommendations to the French-speaking community in order to better take into account the specificities of MRI at all stages of the study and to participate in the improvement of the quality of MRI research. These recommendations also take into account the specificities of MRI research platforms and clinical imaging services, as well as regulatory constraints.
BIDS-prov: a provenance framework for BIDS
Participants: Remi Adon, Camille Maumet.
Differences in details of analysis pipelines have a non negligible impact on neuroimaging results: end-to-end pipelines, neuroimaging software package, software versions and operating system have all been shown to introduce some level of variations. In order to interpret and compare scientific results as well as enable data reuse, researchers need a precise description of a hierarchy of data manipulation and transformations steps from original data to a finding. This description or ‘provenance’ includes information about data, software (versions, parameters, etc.) and people. The Brain Imaging Data Structure (BIDS) has been well adopted in the neuroimaging community and provides structured file hierarchies with JSON metadata files to represent many different aspects of a brain datasets including: raw data across modalities (MRI, EEG, iEEG) but also some derived data. However, BIDS does not capture details of transformations within a BIDS dataset (e.g., DICOMs to BIDS files and BIDS derivatives). We proposed a formal provenance framework for BIDS. Associated publication: 42.
BIDS Statistical Models - An implementation-independent representation of General Linear Models
Participants: Camille Maumet.
The general linear model (GLM) is a mainstay of neuroimaging analysis, and particularly task-based functional MRI (fMRI) analysis. As such, there is an independent implementation in all major tool suites. This proliferation provides access to the technique to researchers from varied backgrounds, but each implementation has its own input specification. Consequently, it is non-trivial to compare methods across studies that use different tool suites or to construct the same model across suites in order to compare the tools themselves. The Brain Imaging Data Structure (BIDS) is a standard for organizing data from a broad range of neuroscientific experiments, including neuroimaging data, experimental events and physiological recordings. Because experiments are often carefully designed to elicit neural responses corresponding to specific proposed cognitive mechanisms, it is useful to include the intended model in the dataset as a means of documenting the experiment and providing a software-independent guide to reproduce the analysis. We introduced BIDS Stats-Models, a specification for describing how a GLM or similar model should be fit to a BIDS dataset. Associated publication: 44. This work was performed as part of an international collaboration led by Christopher Markiewicz from Stanford University.
8.1.2 Detection and learning
In this section, we summarised different contributions focusing on information extraction from medical image data. First, in the field of medical imaging, machine learning methods can be used to detect subtle brain abnormalities, in order to improve the quality of a diagnosis, a prognostic or a disease understanding. This year, we developed and assessed a machine learning-based approach to improve the monitoring of brain disease activity of Multiple Sclerosis patients. We also described and shared a set of imaging data dedicated to the evaluation of Multiple Sclerosis lesions segmentation algorithms. Second, automated computational approach can be used to compute brain related metrics in a robust and efficient way. This year we developed a diffeomorphic vector field approach to analyze the thickness of the hippocampus. Finally, automated methods can also provide an essential aid to process complex data. We in particular developed a machine learning method to assess the location of EEG electrodes within MRI acquisitions.
A clinically-compatible workflow for computer-aided assessment of brain disease activity in multiple sclerosis patients
Participants: Benoit Combès, Olivier Commowick, Francesca Galassi, Gilles Edan, Jean-Christophe Ferré.
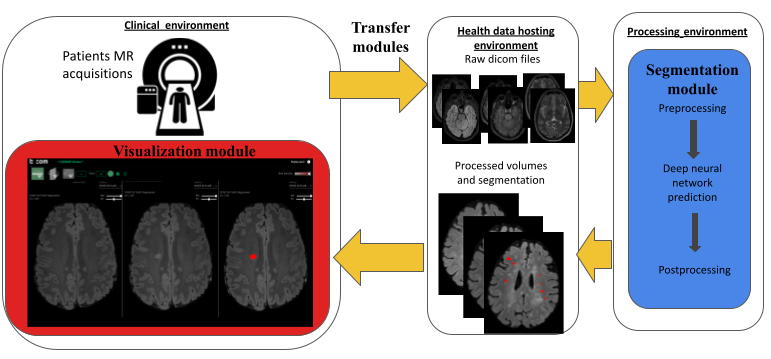
Over the last 10 years, the number of approved disease modifying drugs acting on the focal inflammatory process in Multiple Sclerosis (MS) has increased from 3 to 10. This wide choice offers the opportunity of a personalized medicine with the objective of no clinical and radiological activity for each patient. This new paradigm requires the optimization of the detection of new FLAIR lesions on longitudinal MRI. In this work, we designed a complete workflow -that we developed, implemented, deployed, and evaluated- to facilitate the monitoring of new FLAIR lesions on longitudinal MRI of MS patients. This workflow has been designed to be usable by both hospital and private neurologists and radiologists in France. It consists of three main components: (i) a software component that allows for automated and secured anonymization and transfer of MRI data from the clinical Picture Archive and Communication System (PACS) to a processing server (and vice-versa); (ii) a fully automated segmentation core that enables detection of focal longitudinal changes in patients from T1-weighted, T2-weighted and FLAIR brain MRI scans, and (iii) a dedicated web viewer that provides an intuitive visualization of new lesions to radiologists and neurologists. Then, we evaluated the workflow on 54 pairs of longitudinal MRI scans that were analyzed by 3 experts (1 neuroradiologist, 1 radiologist, and 1 neurologist) with and without the proposed workflow. We showed that our workflow provided a valuable aid to clinicians in detecting new MS lesions both in terms of accuracy (mean number of detected lesions per patient and per expert 1.8 without the workflow vs. 2.3 with the workflow, p =) and of time dedicated by the experts (mean time difference 2'45”, p = ). This increase in the number of detected lesions has implications in the classification of MS patients as stable or active, even for the most experienced neuroradiologist (mean sensitivity was 0.74 without the workflow and 0.90 with the workflow, p-value for no difference = 0.003). It therefore has potential consequences on the therapeutic management of MS patients. Associated publication: 15.
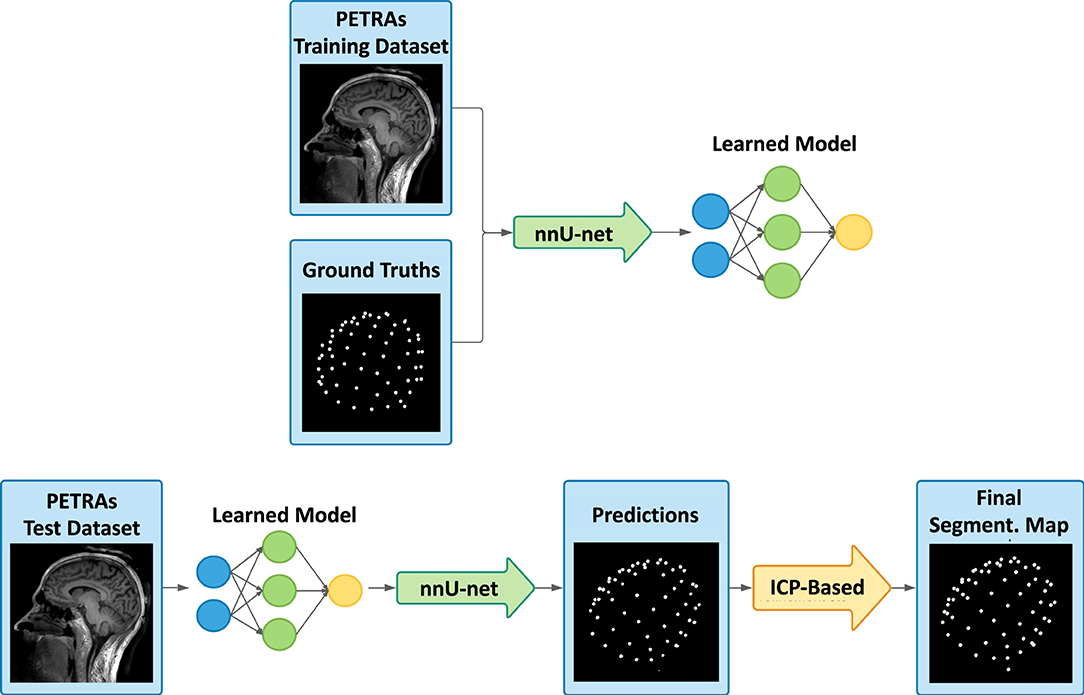
A nnUnet implementation of new lesions segmentation from serial FLAIR images of MS patients
Participants: Arthur Masson, Francesca Galassi, Benoit Combès, Gilles Edan.
In this work, we optimized an nnUNet framework for the segmentation of new multiple sclerosis lesions from FLAIR MR imaging data, acquired at different time points. Overall, our objective was to optimize detection at the lesion scale, with only little attention to lesion delineation (that can be considered as of minor importance within a clinical context). Moreover, for a similar reason, we focused on patients with few new lesions only. The method was evaluated during the MICCAI MSEG2 challenge and was ranked 2nd, out of all 24 in competition, according to the main criterion of detection. Associated publication: 49.
Multiple sclerosis lesions segmentation from multiple experts: The MICCAI 2016 challenge dataset
Participants: Olivier Commowick, Michael Kain, Jean-Christophe Ferré.
MRI plays a crucial role in multiple sclerosis diagnostic and patient follow-up. In particular, the delineation of T2-FLAIR hyperintense lesions is crucial although mostly performed manually-a tedious task. Many methods have thus been proposed to automate this task. However, sufficiently large datasets with a thorough expert manual segmentation are still lacking to evaluate these methods. We have presented a unique dataset for MS lesions segmentation evaluation. It consists of 53 patients acquired on 4 different scanners with a harmonized protocol. Hyperintense lesions on FLAIR were manually delineated on each patient by 7 experts with control on T2 sequence, and gathered in a consensus segmentation for evaluation. We provide raw and preprocessed data and a split of the dataset into training and testing data, the latter including data from a scanner not present in the training dataset. We strongly believe that this dataset will become a reference in MS lesions segmentation evaluation, allowing to evaluate many aspects: evaluation of performance on unseen scanner, comparison to individual experts performance, comparison to other challengers who already used this dataset, etc. Associated publication: 16.
Deep learning-based localization of EEG electrodes within MRI acquisitions
Participants: Caroline Pinte, Mathis Fleury, Pierre Maurel.
The simultaneous acquisition of electroencephalographic (EEG) signals and functional magnetic resonance images (fMRI) aims to measure brain activity with good spatial and temporal resolution. This bimodal neuroimaging can bring complementary and very relevant information in many cases and in particular for epilepsy. Indeed, it has been shown that it can facilitate the localization of epileptic networks. Regarding the EEG, source localization requires the resolution of a complex inverse problem that depends on several parameters, one of the most important of which is the position of the EEG electrodes on the scalp. These positions are often roughly estimated using fiducial points. In simultaneous EEG-fMRI acquisitions, specific MRI sequences can provide valuable spatial information. In this work, we proposed a new fully automatic method based on neural networks to segment an ultra-short echo-time MR volume in order to retrieve the coordinates and labels of the EEG electrodes. It consists of two steps: a segmentation of the images by a neural network, followed by the registration of an EEG template on the obtained detections. We trained the neural network using 37 MR volumes and then we tested our method on 23 new volumes. The results show an average detection accuracy of 99.7% with an average position error of 2.24 mm, as well as 100% accuracy in the labeling. Associated publication: 34.
A diffeomorphic vector field approach to analyze the thickness of the hippocampus from 7T MRI
Participants: Claire Cury.
7-Tesla MRI of the hippocampus enhances the visualization of its internal substructures. Among these substructures, the cornu Ammonis and subiculum form a contiguous folded ribbon of gray matter. Here, we propose a method to analyze local thickness measurements of this ribbon. We introduce an original approach based upon the estimation of a diffeomorphic vector field that traverses the ribbon. The method is designed to handle specificities of the hippocampus and corresponding 7-Tesla acquisitions: highly convoluted surface, non closed ribbon, incompletely defined inner/outer boundaries, anisotropic acquisitions. We furthermore propose to conduct group comparisons using a population template built from the central surfaces of individual subjects. We first assessed the robustness of our approach to anisotropy, as well as to inter-rater variability, on a post-mortem scan and on in vivo acquisitions respectively. We then conducted a group study on a dataset of in vivo MRI from temporal lobe epilepsy (TLE) patients and healthy controls. The method detected local thinning patterns in patients, predominantly ipsilaterally to the seizure focus, which is consistent with medical knowledge. This new technique allows measuring the thickness of the hippocampus from 7-Tesla MRI. It shows good robustness with respect to anisotropy and inter-rater variability and has the potential to detect local atrophy in patients. As 7-Tesla MRI is increasingly available, this new method may become a useful tool to study local alterations of the hippocampus in brain disorders. It is made freely available to the community (code, postmortem segmentation). Associated publication: 24.
8.1.3 Quantitative imaging
Quantitative imaging is essential for an accurate and specific characterisation of, among others, tissue integrity or neural activity. This year, we contributed to methods in acquisition and processing of diffusion MRI. We also explored the clinical interest of Magnetization Transfer Ratio imaging, Positron Emission Tomography, Quantitative Perfusion Mapping and Arterial Spin Labeling in various clinical applications. Finally, we made contributions in the design of phantoms to evaluate quantitative susceptibility reconstruction methods and tractrography methods.
Quantitative perfusion mapping with induced transient hypoxia using BOLD MRI
Participants: Julie Coloigner.
Gadolinium-based dynamic susceptibility contrast (DSC) is commonly used to characterize blood flow in patients with stroke and brain tumors. Unfortunately, gadolinium contrast administration has been associated with adverse reactions and longterm accumulation in tissues. In this work, we propose an alternative deoxygenation based dynamic susceptibility contrast (dDSC) method that uses a transient hypoxia gas paradigm to deliver a bolus of paramagnetic deoxygenated hemoglobin to the cerebral vasculature for perfusion imaging.Through traditional DSC tracer kinetic modeling, the MR signal change induced by this hypoxic bolus can be used to generate regional perfusion maps of cerebral blood flow, cerebral blood volume and mean transit time. This gas paradigm and BOLD-MR imaging were performed concurrently on a cohort of 66 healthy and chronically anemic subjects (age 23.5±9.7, female 64%).
Our results showed reasonable global and regional agreement between dDSC and other flow techniques like phase contrast and arterial spin labeling. In this proof-of-concept study, we demonstrated the feasibility of using transient hypoxia to generate a contrast bolus that mimics the effect of gadolinium and yields reasonable perfusion estimates. Looking forward, optimization of the hypoxia boluses and measurement of the arterial-input-function is necessary to improve the accuracy of dDSC. Additionally, a cross-validation study of dDSC and DSC in brain tumor and ischemic stroke subjects is warranted to evaluate the clinical diagnostic utility of this approach. Associated publication: 37.
Effect of distortion corrections on the tractography quality in spinal cord diffusion‐weighted imaging
Participants: Elise Bannier.
Spinal cord sagittal diffusion-weighted imaging data present major distortions. Different distortion correction (DC) method and patient geometry (sagittal balance) influence the quality of spinal cord tractography rendering according to different tractography approaches. Forty-four adults free of spinal cord diseases underwent cervical diffusion-weighted imaging. The phase-encoding direction was head→foot. Sequence with opposed polarities (foot→head) was acquired to perform DC. Eddy-current, motion effects, and susceptibility artifact correction methods were used for DC, and two deterministic and one probabilistic tractography approaches were evaluated using MRtrix and DSI Studio tractography software. Fiber length and number of fibers were extracted to evaluate the quality of the tractography rendering. For each subject, cervical lordosis was measured to assess patient geometry. The angle between the main direction of the spinal cord and the orientation of the acquisition box were computed at each spine level to assess acquisition geometry and define an angle threshold for which a tractography of good quality is no longer possible. There was a significant improvement in tractography quality after performing DC with susceptibility artifact correction using a deterministic approach based on tensor. Before DC, the angle threshold was defined at C6 (15.2°) compared with C7 (21.9°) after corrections, demonstrating the importance of spinal cord angulation for DC. In conclusion, the impact of DC on tractography quality is greatly impacted by acquisition geometry. To obtain a good-quality tractography, we propose as a future perspective to adapt the acquisition geometry to that of the patient by automatically adjusting the acquisition box. Associated publication: 17.
Interpolation and averaging of diffusion MRI multi-compartment models
Participants: Renaud Hédouin, Olivier Commowick.
Multi-compartment models (MCM) are increasingly used to characterize the brain white matter microstructure from diffusion-weighted imaging (DWI). Their use in clinical studies is however limited by the inability to resample an MCM image towards a common reference frame, or to construct atlases from such brain microstructure models. We proposed to solve this problem by first identifying that these two tasks amount to the same problem. We proposed to tackle it by viewing it as a simplification problem, solved thanks to spectral clustering and the definition of semi-metrics between several usual compartments encountered in the MCM literature. This generic framework was evaluated for two models: the multi-tensor model where individual fibers are modeled as individual tensors and the diffusion direction imaging (DDI) model that differentiates intra- and extra-axonal components of each fiber. Results on simulated data, simulated transformations and real data showed the ability of our method to well interpolate MCM images of these types. We finally presented as an application an MCM template of normal controls constructed using our approach. Associated publication: 25.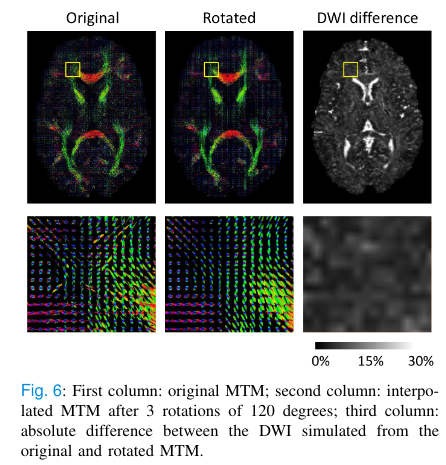
Multiparametric analysis of cerebral development in preterm infants using magnetic resonance imaging
Participants: Isabelle Corouge, Olivier Commowick, Jean-Christophe Ferré.
The severity of neurocognitive impairment increases with prematurity. However, its mechanisms remain poorly understood. Our aim was firstly to identify multiparametric magnetic resonance imaging (MRI) markers that differ according to the degree of prematurity, and secondly to evaluate the impact of clinical complications on these markers. We prospectively enrolled preterm infants who were divided into two groups according to their degree of prematurity: extremely preterm (>28 weeks’ gestational age) and very preterm (28–32 weeks’ gestational age). They underwent a multiparametric brain MRI scan at term-equivalent age including morphological, diffusion tensor and arterial spin labeling (ASL) perfusion sequences. We quantified overall and regional volumes, diffusion parameters, and cerebral blood flow (CBF). We then compared the parameters for the two groups. We also assessed the effects of clinical data and potential MRI morphological abnormalities on those parameters. Thirty-four preterm infants were included. Extremely preterm infants had significantly higher frontal relative volumes , frontal GM relative volumes, and regional CBF than very preterm infants, but they had lower brainstem and insular relative volumes. Preterm infants with WM lesions on MRI had significantly lower overall GM CBF (13.3 ± 2 ml/100 g/min versus 17.7 ± 2.5, < ml/100 g/min). Associated publication: 20.
Decoding the microstructural properties of white matter using realistic models
Participants: Renaud Hédouin.
Multi-echo gradient echo (ME-GRE) magnetic resonance signal evolution in white matter has a strong dependence on the orientation of myelinated axons with respect to the main static field. Although analytical solutions have been able to predict some of the white matter (WM) signal behaviour of the hollow cylinder model, it has been shown that realistic models of WM offer a better description of the signal behaviour observed.
In this work, we presented a pipeline to (i) generate realistic 2D WM models with their microstructure based on real axon morphology with adjustable fiber volume fraction (FVF) and g-ratio. We (ii) simulated their interaction with the static magnetic field to be able to simulate their MR signal. For the first time, we (iii) demonstrated that realistic 2D WM models can be used to simulate a MR signal that provides a good approximation of the signal obtained from a real 3D WM model derived from electron microscopy. We then (iv) demonstrated in silico that 2D WM models can be used to predict microstructural parameters in a robust way if ME-GRE multi-orientation data is available and the main fiber orientation in each pixel is known using DTI. A deep learning network was trained and characterized in its ability to recover the desired microstructural parameters such as FVF, g-ratio, free and bound water transverse relaxation and magnetic susceptibility. Finally, the network was trained to recover these micro-structural parameters from an ex vivo dataset acquired in 9 orientations with respect to the magnetic field and 12 echo times. We demonstrated that this is an overdetermined problem and that as few as 3 orientations can already provide comparable results for some of the decoded metrics. Associated publication: 26
QSM reconstruction challenge 2.0: A realistic in silico head phantom for MRI data simulation and evaluation of susceptibility mapping procedures
Participants: Renaud Hédouin.
The purpose of the challenge was to create a realistic in silico head phantom for the second QSM reconstruction challenge and for future evaluations of processing algorithms for QSM. We created a digital whole-head tissue property phantom by segmenting and postprocessing high-resolution (0.64 mm isotropic), multiparametric MRI data acquired at 7 T from a healthy volunteer. We simulated the steady-state magnetization at 7 T using a Bloch simulator and mimicked a Cartesian sampling scheme through Fourier-based processing. Computer code for generating the phantom and performing the MR simulation was designed to facilitate flexible modifications of the phantom in the future, such as the inclusion of pathologies as well as the simulation of a wide range of acquisition protocols. Specifically, the following parameters and effects were implemented: TR and TE, voxel size, background fields, and RF phase biases. Diffusion-weighted imaging phantom data are provided, allowing future investigations of tissue-microstructure effects in phase and QSM algorithms. The brain part of the phantom featured realistic morphology with spatial variations in relaxation and susceptibility values similar to the in vivo setting. We demonstrated some of the phantom’s properties, including the possibility of generating phase data with nonlinear evolution over TE due to partial-volume effects or complex distributions of frequency shifts within the voxel. The presented phantom and computer programs are publicly available and may serve as a ground truth in future assessments of the faithfulness of quantitative susceptibility reconstruction algorithms. Associated publication: 33.The diffusion-simulated connectivity (DiSCo) dataset
Participants: Emmanuel Caruyer, Raphaël Truffet.
The methodological development in the mapping of the brain structural connectome from diffusion-weighted magnetic resonance imaging (DW-MRI) has raised many hopes in the neuroscientific community. Indeed, the knowledge of the connections between different brain regions is fundamental to study brain anatomy and function. The reliability of the structural connectome is therefore of paramount importance. In the search for accuracy, researchers have given particular attention to linking white matter tractography methods – used for estimating the connectome – with information about the microstructure of the nervous tissue. The creation and validation of methods in this context were hampered by a lack of practical numerical phantoms. To achieve this, we created a numerical phantom that mimics complex anatomical fibre pathway trajectories while also accounting for microstructural features such as axonal diameter distribution, myelin presence, and variable packing densities. The substrate has a micrometric resolution and an unprecedented size of 1 cubic millimetre to mimic an image acquisition matrix of voxels. DW-MRI images were obtained from Monte Carlo simulations of spin dynamics to enable the validation of quantitative tractography. The phantom is composed of 12,196 synthetic tubular fibres with diameters ranging from 1.4 µm to 4.2 µm, interconnecting sixteen regions of interest. The simulated images capture the microscopic properties of the tissue (e.g. fibre diameter, water diffusing within and around fibres, free water compartment), while also having desirable macroscopic properties resembling the anatomy, such as the smoothness of the fibre trajectories. While previous phantoms were used to validate either tractography or microstructure, this phantom can enable a better assessment of the connectome estimation’s reliability on the one side, and its adherence to the actual microstructure of the nervous tissue on the other. Associated publications: 35, 48.

8.2 Translational research
8.2.1 Behavior
Our objective is also to provide new computational solutions for our target clinical applications (Alzheimer's disease, psychiatry, neurology or public health issues), allowing a more appropriate representation of data for image analysis and the detection of biomarkers specific to a form or grade of pathology, or specific to a population of subjects. In this section, we present our contributions in different clinical applications.
Building memories on prior knowledge: behavioral and fMRI evidence of impairment in early Alzheimer's disease
Participants: Pierre-Yves Jonin, Quentin Duché, Elise Bannier, Isabelle Corouge, Jean-Christophe Ferré.
Impaired memory is a hallmark of prodromal Alzheimer's disease (AD). Prior knowledge associated with the memoranda improves memory in healthy individuals, but we ignore whether the same occurs in early AD. We used functional MRI to investigate whether prior knowledge enhances memory encoding in early AD, and whether the nature of this prior knowledge matters. Patients with early AD and Controls underwent a task-based fMRI experiment where they learned face-scene associations. Famous faces carried pre-experimental knowledge (PEK), while unknown faces with which participants were familiarized prior to learning carried experimental knowledge (EK). Surprisingly, PEK strongly enhanced subsequent memory in healthy controls, but importantly not in patients. Partly nonoverlapping brain networks supported PEK vs. EK associative encoding in healthy controls. No such networks were identified in patients. In addition, patients displayed impaired activation in a right subhippocampal region where activity predicted successful associative memory formation for PEK stimuli. Despite the limited sample sizes of this study, these findings suggest that the role prior knowledge in new learning might have been so far overlooked and underestimated in AD patients. Prior knowledge may drive critical differences in the way healthy elderly and early AD patients learn novel associations. Associated publication: 28.
Multimodal brain imaging connectivity analyses of emotional and motivational deficits in depression among women
Participants: Elise Bannier, Isabelle Corouge, Jean-Christophe Ferré.
Major depressive disorder (MDD) is characterized by impaired cortical–subcortical functional connectivity. Apathy adds to functional impairment, but its cerebral basis in MDD remains unknown. Our objective was to describe impairments in functional connectivity during emotional processing in MDD (with varying levels of congruency and attention), and to determine their correlation with apathy. We used the Variable Attention Affective Task during functional MRI, followed by diffusion-weighted MRI, to assess 55 right-handed women (30 with MDD and 25 healthy controls) between September 2012 and February 2015. We estimated functional connectivity using generalized psychophysiologic interaction and anatomic connectivity with tract-based spatial statistics. We measured apathy using the Apathy Evaluation Scale. We found decreased functional connectivity between the left amygdala and the left anterior cingulate cortex (ACC) during negative stimuli in participants with MDD (t54 = 4.2; p = 0.035, family-wise error [FWE]–corrected). During high-attention stimuli, participants with MDD showed reduced functional connectivity between the right dorsolateral prefrontal cortex (dlPFC) and the right ACC (t54 = 4.06, pFWE = 0.02), but greater functional connectivity between the right dlPFC and the right amygdala (t54 = 3.35, p = 0.048). Apathy was associated with increased functional connectivity between the right dlPFC and the right ACC during high-attention stimuli (t28 = 5.2, p = 0.01) and increased fractional anisotropy in the right posterior cerebellum, the anterior and posterior cingulum and the bilateral internal capsule (all pFWE < 0.05). Limitations included a moderate sample size, concomitant antidepressant therapy and no directed connectivity. We found that MDD was associated with impairments in cortical–subcortical functional connectivity during negative stimuli that might alter the recruitment of networks engaged in attention. Apathy-related features suggested networks similar to those observed in degenerative disorders, but possible different mechanisms. Associated publication: 36.
Multimodal MRI cerebral correlates of verbal fluency switching and its impairment in women with depression
Participants: Isabelle Corouge, Elise Bannier, Jean-Christophe Ferré.
The search of biomarkers in the field of depression requires easy implementable tests that are biologically rooted. Qualitative analysis of verbal fluency tests (VFT) are good candidates, but its cerebral correlates are unknown. We collected qualitative semantic and phonemic VFT scores along with grey and white matter anatomical MRI of depressed (n = 26) and healthy controls (HC, n = 25) women. Qualitative VFT variables are the “clustering score” (i.e. the ability to produce words within subcategories) and the “switching score” (i.e. the ability to switch between clusters). The clustering and switching scores were automatically calculated using a data-driven approach. Brain measures were cortical thickness (CT) and fractional anisotropy (FA). We tested for associations between CT, FA and qualitative VFT variables within each group. Patients had reduced switching VFT scores compared to HC. Thicker cortex was associated with better switching score in semantic VFT bilaterally in the frontal (superior, rostral middle and inferior gyri), parietal (inferior parietal lobule including the supramarginal gyri), temporal (transverse and fusiform gyri) and occipital (lingual gyri) lobes in the depressed group. Positive association between FA and the switching score in semantic VFT was retrieved in depressed patients within the corpus callosum, right inferior fronto-occipital fasciculus, right superior longitudinal fasciculus extending to the anterior thalamic radiation (all p < 0.05, corrected). Together, these results suggest that automatic qualitative VFT scores are associated with brain anatomy and reinforce its potential use as a surrogate for depression cerebral bases. Associated publication: 19.
8.2.2 Neuro-inflammation
This year, we consolidated our expertise regarding the relevance of imaging the spinal cord to investigate early biomarkers for MS. Moreover, we provided an atlas-based framework to analyse the optic radiations of MS patients with acute optic neuritis.
Prognostic value of spinal cord MRI in multiple sclerosis patients
Participants: Benoit Combès, Elise Bannier.
Multiple sclerosis (MS) is a common inflammatory, demyelinating and neurodegenerative disease of the central nervous system that affects both the brain and the spinal cord. In clinical practice, spinal cord MRI is performed far less frequently than brain MRI, mainly owing to technical limitations and time constraints. However, improvements of acquisition techniques, combined with a strong diagnosis and prognostic value, suggest an increasing use of spinal cord MRI in the near future. We provided a review of the current data from the literature on the prognostic value of spinal cord MRI in MS patients in the early and later stages of their disease. Both conventional and quantitative MRI techniques are discussed. The prognostic value of spinal cord lesions is clearly established at the onset of disease, underlining the interest of spinal cord conventional MRI at this stage. However, studies are currently lacking to affirm the prognostic role of spinal cord lesions later in the disease, and therefore the added value of regular follow-up with spinal cord MRI in addition to brain MRI. Besides, spinal cord atrophy, as measured by the loss of cervical spinal cord area, is also associated with disability progression, independently of other clinical and MRI factors including spinal cord lesions. Although potentially interesting, this measurement is not currently performed as a routine clinical procedure. Finally, other measures extracted from quantitative MRI have been established as valuable for a better understanding of the physiopathology of MS, but still remain a field of research. This work was done in collaboration with Anne Kerbrat from the Neurology Department, University Hospital of Rennes. Associated publication: 39.
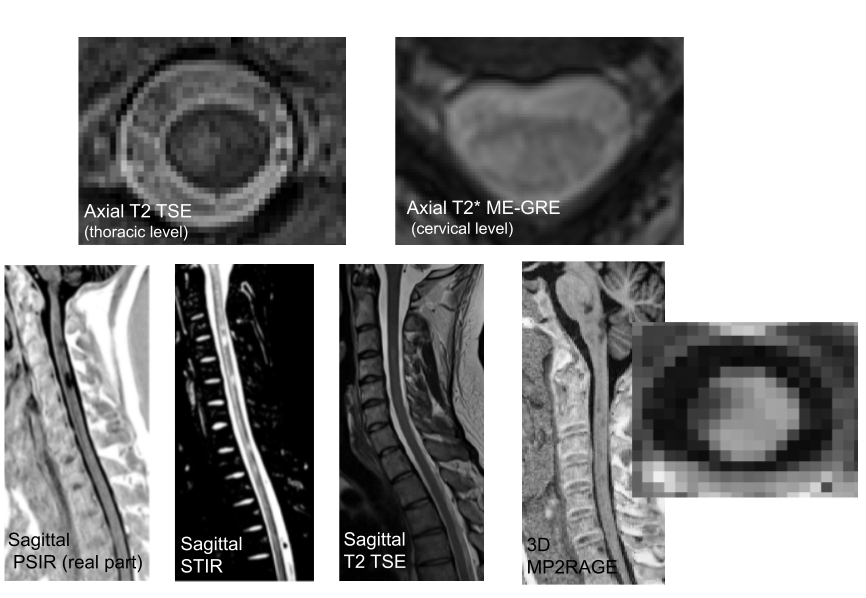
Asymmetry of magnetization transfer ratio (MTR) measurements in the cervical spinal cord of MS patients
Participants: Benoit Combès, Elise Bannier.
Spinal cord (SC) is often affected in Multiple Sclerosis (MS) and its microstructure can be assessed using magnetization transfer (MT) imaging. However, the variability of the MT ratio (MTR) measurements between subjects and between acquisitions makes it difficult to use in clinical practice. In this work, we proposed to study the left-to-right asymmetry of spinal cord MTR measurements. Such a metric would have the advantage to be insensitive to inter-session and inter-subject variations, thus offering a lower natural variability than those of the raw MTR mean values. Moreover, this measure could allow us to explore the asymmetry of sensory and motor impairment frequently observed in MS patients. For this purpose, we characterised the left-right asymmetry of cervical spinal cord MTR in control subjects and MS patients and assess its association with lesion load and clinical data in 66 early relapsing remitting MS patients (< 1 year). 44 controls were also included. For each cervical level, MTR left-to-right asymmetries were assessed for three regions (whole cervical cord, white matter and grey matter), compared between subgroups of patients and correlated to lesion load measured on T2-gradient echo images and clinical status evaluated with the disability score Expanded Disability Status Scale (EDSS) at baseline and two years. This study allowed us to observe no evidence of significant MTR asymmetry over the different cervical cord levels. Moreover, values of metrics based on MTR asymmetry were significantly higher in patients than in controls (typically for level C4 in whole cord: patient-to-control difference=0.60pu, p value for no mean difference=). Finally, overall for patients, these metrics were associated with the global and the lateralized cervical lesion load (, and , ) and with the two-years EDSS score (r=0.30, p=0.02). Associated publication: 52.
Patient specific tracts-based analysis of diffusion compartment models: application to multiple sclerosis patients with acute optic neuritis
Participants: Olivier Commowick, Renaud Hédouin, Jean-Christophe Ferré.
Multiple sclerosis is a complex disease where voxel-based, group-based statistics of the brain microstructure have shown their limits in explaining patient evolution. This is first due to too simple diffusion models, mixing information. Voxel-based studies also lack knowledge on brain structural connectivity. Finally, group-based analysis does not describe well the specific patient status (a crucial point for clinicians). We have proposed an atlas-based framework, combined with advanced diffusion compartment models, for patient specific analysis of microstructural disease burden on major fiber bundles. We applied our framework to the analysis of optic radiations of MS patients with acute optic neuritis. Associated publication: 40.
Combining F-18-DOPA PET and MRI with perfusion-weighted imaging improves delineation of high-grade subregions in enhancing and non-enhancing gliomas prior treatment: a biopsy-controlled study
Participants: Elise Bannier.
We aimed to compare spatial extent of high-grade subregions detected with combined [F-18]-dihydroxyphenylalanine (F-18-DOPA) PET and MRI to the one provided by advanced multimodal MRI alone including Contrast-Enhanced (CE) and Perfusion-Weighted Imaging (PWI). Then, we compared the accuracy between imaging modalities, in a per biopsy analysis. Participants with suspected diffuse glioma were prospectively included between June 2018 and September 2019. Volumes of high-grade subregions were delineated respectively on F-18-DOPA PET and MRI (CE and PWI). Up to three per-surgical neuronavigation-guided biopsies were performed per patient. Thirty-eight biopsy samples from sixteen participants were analyzed. Six participants had grade IV IDH wild-type glioblastoma, six had grade III IDH-mutated astrocytoma and four had grade II IDH-mutated gliomas. Three patients had intratumoral heterogeneity with coexisting high- and low-grade tumor subregions. High-grade volumes determined with combined F-18-DOPA PET/MRI (median of 1.7 [interquartile range (IQR) 0.0, 19.1] mL) were larger than with multimodal MRI alone (median 1.3 [IQR 0.0, 12.8] mL) with low overlap (median Dice's coefficient 0.24 [IQR 0.08, 0.59]). Delineation volumes were substantially increased in five patients. In a per biopsy analysis, combined F-18-DOPA PET/MRI detected high-grade subregions with an accuracy of 58 percent compared to 42 percent (p = 0.03) with CE MRI alone and 50 percent (p = 0.25) using multimodal MRI (CE + PWI). The addition of F-18-DOPA PET to multimodal MRI (CE and PWI) enlarged the delineation volumes and enhanced overall accuracy for detection of high-grade subregions. Thus, combining F-18-DOPA with advanced MRI may improve treatment planning in newly diagnosed gliomas. Associated publication: 23.
8.2.3 Recovery
This axis aims at developing and evaluating new rehabilitation protocols involving imaging. In particular, the first work presented hybrid EEG-MRI bimodal neurofeedback protocols that were carried out and evaluated on the effective connectivity of chronic stroke patients. The second work evaluated on healthy subjects two rehabilitation strategies for post-stroke patients with spatial cognition disorders.
The impact of Neurofeedback on effective connectivity networks in chronic stroke patients: an exploratory study
Participants: Julie Coloigner, Quentin Duché, Elise Bannier, Isabelle Bonan.
In this work, we assessed the impact of EEG-fMRI Neurofeedback (NF) training on connectivity strength and direction in bilateral motor cortices in chronic stroke patients. Most of the studies using NF or brain computer interfaces for stroke rehabilitation have assessed treatment effects focusing on successful activation of targeted cortical regions. However, given the crucial role of brain network reorganization for stroke recovery, our broader aim was to assess connectivity changes after a NF training protocol targeting localised motor areas.
We considered changes in fMRI connectivity after a multisession EEG-fMRI NF training targeting ipsilesional motor areas in nine stroke patients. We applied the Dynamic Causal Modeling and Parametric Empirical Bayes frameworks for the estimation of effective connectivity changes. We considered a motor network including both ipsilesional and contralesional premotor, supplementary and primary motor areas.
Our results indicated that NF upregulation of targeted areas (ipsilesional supplementary and primary motor areas) not only modulated activation patterns, but also had a more widespread impact on fMRI bilateral motor networks. In particular, inter-hemispheric connectivity between premotor and primary motor regions decreased, and ipsilesional self-inhibitory connections were reduced in strength, indicating an increase in activation during the NF motor task.
To the best of our knowledge, this is the first work that investigates fMRI connectivity changes elicited by training of localized motor targets in stroke. Our results open new perspectives in the understanding of large-scale effects of NF training and the design of more effective NF strategies, based on the pathophysiology underlying stroke-induced deficits. Associated publication: 32.
The neural bases of egocentric spatial representation for extracorporeal and corporeal tasks: an fMRI study
Participants: Stéphanie Leplaideur, Quentin Duché, Jean-Christophe Ferré, Elise Bannier, Isabelle Bonan.
Humans use reference frames to elaborate the spatial representations needed for all space-oriented behaviors such as postural control, walking or grasping. We investigated the neural bases of two egocentric tasks: the extracorporeal subjective straight-ahead task (SSA) and the corporeal subjective longitudinal body plane task (SLB) in healthy participants using functional magnetic resonance imaging (fMRI). This work was an ancillary part of a study involving stroke patients. Seventeen healthy participants underwent a 3T fMRI examination. During the SSA, participants had to divide the extracorporeal space into two equal parts. During the SLB, they had to divide their body along the midsagittal plane. Both tasks elicited a parieto-occipital network encompassing the superior and inferior parietal lobules and lateral occipital cortex, with a right hemispheric dominance. Additionally, the SLB > SSA contrast revealed activations of the left angular and premotor cortices. These areas, involved in attention and motor imagery suggest a greater complexity of corporeal processes engaging body representation. This was the first fMRI study to explore the SLB-related activity and its complementarity with the SSA. Our results pave the way for the exploration of spatial cognitive impairment in patients. Associated publication: 30.
8.3 Contributions to Open Science
Additionally to our main research axes, we participated to large scale transversal collaborative works devoted to the improvement and democratization of open science practices.
Centering inclusivity in the design of online conferences - An OHBM - Open Science perspective
Participants: Camille Maumet.
As the global health crisis unfolded, many academic conferences moved online in 2020. This move has been hailed as a positive step towards inclusivity in its attenuation of economic, physical, and legal barriers and effectively enabled many individuals from groups that have traditionally been underrepresented to join and participate. A number of studies have outlined how moving online made it possible to gather a more global community and has increased opportunities for individuals with various constraints, e.g., caregiving responsibilities. Yet, the mere existence of online conferences is no guarantee that everyone can attend and participate meaningfully. In fact, many elements of an online conference are still significant barriers to truly diverse participation: the tools used can be inaccessible for some individuals; the scheduling choices can favour some geographical locations; the set-up of the conference can provide more visibility to well-established researchers and reduce opportunities for early-career researchers. While acknowledging the benefits of an online setting, especially for individuals who have traditionally been underrepresented or excluded, we recognize that fostering social justice requires inclusivity to actively be centered in every aspect of online conference design. In this project, we drew from the literature and from our own experiences to identify practices that purposefully encourage a diverse community to attend, participate in, and lead online conferences. Reflecting on how to design more inclusive online events is especially important as multiple scientific organizations have announced that they will continue offering an online version of their event when in-person conferences can resume. Associated publication: 31.
This work was done as part of an international collaboration with 100
Brainhack: Developing a culture of open, inclusive, community-driven neuroscience
Participants: Elise Bannier, Claire Cury.
Brainhack is an innovative meeting format that promotes scientific collaboration and education in an open, inclusive environment. In this project, we described the myriad benefits for participants and the research community and how Brainhacks complement conventional formats to augment scientific progress. This work was done as part of an international collaboration with 200
9 Bilateral contracts and grants with industry
9.1 Bilateral contracts with industry
9.1.1 Siemens
Participants: Elise Bannier, Emmanuel Caruyer, Olivier Commowick, Isabelle Corouge, Jean-Christophe Ferré, Jean-Yves Gauvrit.
In the context of the Neurinfo imaging platform, a master research agreement between Siemens SAS - Healthcare and University of Rennes 1 defines the terms of the collaboration between Siemens, Empenn and the Neurinfo platform. Relying on this research agreement contract, Neurinfo has received work in progress (WIP) sequences from Siemens in the form of object code for evaluation in the context of clinical research. The Neurinfo platform has also received source code of selected MRI sequences. As an example, the diffusion sequence code was modified to load arbitrary diffusion gradient waveforms for the FastMicroDiff project led by E. Caruyer. This is crucial in the collaboration since it enables the development of MRI sequences on site. The MR Diffusion pulse sequence source code was modified in collaboration with our Siemens clinical scientist as part of our Master Research Agreement, Marc Lapert, in order to play arbitrary gradient waveforms. This was done on the Syngo VB17 software version and again VE11C. Acquisitions on healthy controls have started to evaluate several sets of waveforms.10 Partnerships and cooperations
10.1 International initiatives
10.1.1 Associate Teams in the framework of the EPFL-Inria International Lab
MMINCARAV
Participants: Élise Bannier, Emmanuel Caruyer, Julie Coloigner, Olivier Commowick, Thomas Durantel, Renaud Hédouin, Raphaël Truffet.
The objectives of the MMINCARAV (Multimodal Microstructure-Informed Connectivity: Acquisition, Reconstruction, Analysis and Validation) associate team is to address new scientific challenges related to the use of multimodal magnetic resonance imaging (MRI) to derive microstructure indices and apply them to the measure of brain connectivity. We focus on 4 aspects of this: first we will develop novel sampling techniques, with the objective to reduce acquisition time for the accurate reconstruction of microstructure indices using diffusion MRI; next we will propose joint T2 relaxometry and diffusion models for the description of microstructure, to take advantage of the complementarity of both modalities in the estimation of microstructure indices; in continuation, we will propose new statistical and network analysis methods using the microstructure-informed connectome, and evaluate its potential to reduce bias and false positives; last we will develop a realistic simulation tool combining a fine macroscopic description of fiber bundles, with a fast and realistic simulator at the mesoscopic scale developed by LTS5.
We organized regular meetings virtually throughout the year; Jean-Philippe Thiran was part of Raphaël Truffet's PhD committee. The DiSCo challenge, organized at MICCAI'2021, is a result of this collaboration.
10.1.2 Informal international partners
Camille Maumet collaborates with
- Prof. Thomas Nichols and his group, NISOx at the Oxford Big Data Institute on neuroimaging statistics,
- Prof. Jean-Baptiste Poline and his group at McGill University on neuroimaging data sharing,
- Prof. Satrajit Ghosh and his group at MIT,
- International members of the INCF on neuroimaging data sharing.
Claire Cury is associated researcher at the ICM, Brain and Spine Institute, Paris. Working with the ARAMIS team on the IHI project.
Pierre Maurel collaborates with Dr Noorzadeh at the Institute of medical science and technologies (Shahid Beheshti university, Iran) on combining EEG (Electroencephalogram) and fMRI (functional Magnetic Resonance Imaging).
10.2 International research visitors
10.2.1 Visits of international scientists
-
Name
Agustina Fragueiro
-
Status
PhD
-
Institution of origin:
University of Studies G. d’Annunzio Chieti Pescara
-
Country:
Italy
-
Dates:
November 2021 - February 2022
-
Context of the visit:
Shape analysis of the hippocampus and the intra-parietal fissure to initiate a collaboration with Claire Cury.
-
Mobility program/type of mobility:
PhD mobility grant from University of Studies G. d’Annunzio Chieti Pescara
10.3 European initiatives
10.3.1 European COST Action GLIMR
Participants: Camille Maumet, Elise Bannier.
The GLIMR COST Action (PI: Esther Warnert, Erasmus MC, Netherlands) aims to build a pan-European and multidisciplinary network of international experts in glioma research, patient organisations, data scientists, and MR imaging scientists by uniting the glioma imaging community within Europe and progressing the development and application of advanced MR imaging for improved decision making in diagnosis, patient monitoring, and assessment of treatment response in clinical trials and clinical practice. Camille Maumet leads the work package "WG2 - Multi-site data integration" with Cyril Pernet (University of Edinburgh, UK). GliMR’s first grant period ran from September 2019 to April 2020, during which several meetings were held and projects were initiated, such as reviewing the current knowledge on advanced MRI; developing a General Data Protection Regulation (GDPR) compliant consent form; and setting up the website. A publication led by Patricia Clement (Ghent University, Belgium) was published in 2021 describing the results of this GliMR’s first grant period 14. The Action overcomes the pre-existing limitations of glioma research and is funded until September 2023. New members will be accepted during its entire duration.
10.4 National initiatives
10.4.1 EyeSkin-NF : Eye-tracking and skin conductance measures for neurofeedback analysis and validation
Participants: Claire Cury, Elise Bannier, Pierre Maurel, Hachim Bani.
Funding : Action exploratoire Inria - Duration: 2021 - 2024.
Neurofeedback techniques (NF) or restorative brain computer interfaces (BCI) consist in providing a subject with real-time feedback about its own brain activity, in order to learn self-regulate specific brain regions during NF training. Brain activity can be measured by various techniques such as EEG and/or fMRI. However, analysis of NF sessions is limited due to the difficulty at identifying the origin of failed training. To enhance and monitor participant's motivation in real-time during EEG-fMRI recording, bio-signal can be measured via eye-tracking (ET) or skin conductance (SC) devices. For a precise evaluation of the motivation mental states of interest such as focus, arousal, mind wandering or mental load can be analysed. The main objective of this project is to investigate measures from eye-tracking and skin conductance signals to evaluate in real-time subject’s motivation during NF training.
10.4.2 Rapid Neocortical Declarative Learning in normal aging and memory disorders
Participants: Pierre-Yves Jonin, Julie Coloigner.
Funding: Fondation de l'Avenir - Duration: 2020-2022 - Budget: 40k€
Our project aims at making the case for the existence of a rapid declarative learning system largely independent from the extended hippocampal system and characterizing its neural bases, by use of experimental psychology, cognitive neuropsychology and neuroimaging methods. This project is led in collaboration with Dr Audrey Noël, Assistant Prof., University of Rennes 2, France, with Dr Gabriel Besson, Associate Researcher, University of Coimbra, Portugal, with Dr Ann-Kathrin Zaiser, Associate Researcher, University of Heidelberg, Germany, with Dr Serge Belliard, PhD, MD, Rennes University Hospital, Neurology Dept., France, and with Dr Anca Pasnicu, MD, Rennes University Hospital, Neurology Dept., France.
10.4.3 Effect of prenatal exposures to neurotoxicants on the developing brain: an MRI study (PERINE)
Participants: Élise Bannier, Isabelle Corouge, Julie Coloigner, Jean-Christophe Ferré.
Funding: Fondation de France - Duration: 2015-2021 - Budget: 100k€
The PELAGIE cohort evaluates the effect of prenatal exposure to neurotoxicants on child development. Following previous studies, the PERINE study focuses on the assessment of brain developement at 10-12 years old using MRI (ASL, Diffusion imaging, working memory as well as motor inhibition BOLD fMRI together with neuropsychological tests). A total of 101 children were included. A PhD of Anne-Claire Binter was defended in December 2019 linking epidemiology with functional imaging during a GoNoGo task and neuropsychological scores and two publications were co-authored. This work is done in collaboration with Fabienne Pelé and Cécile Chevrier (IRSET).
10.4.4 Connectivity of the amygdala in depression
Participants: Olivier Commowick, Emmanuel Caruyer, Julie Coloigner, Claire Cury.
Funding: Fondation de France - Duration: 2019-2021 - Budget: 200k€
The onset of depression in teenagers and young adults increases the risk to develop a drug-resistant depression in the adulthood. This project aims at evaluating the role of early changes in the microstructure and connectivity of the amygdala. Using a cohort of drug-resistant patients (N=30), non drug-resistant patients (N=30) and controls (N=30), the aim is to identify imaging biomarkers of the pathology and to compare these with emotional and cognitive phenotypes in this population, searching for early differences in the development of the amygdala connectivity. Inclusions are ongoing.
This is a colloborative project with M.-L. Paillère Martinot from Paris-Descartes University, as Principal Investigator.
10.4.5 Multimodal Imaging of the Limbic Amygdala for the Prognosis of Depression (IMpAirED)
Participants: Julie Coloigner, Olivier Commowick, Élise Bannier, Emmanuel Caruyer.
Funding: CNRS-Inserm Défi Santé numérique AAP 2019 - Start: 2019 - Budget: 19k€
This grant is an extension of the Projet Fondation de France: Connectivity of the amygdala in depression.
In order to identify early features of this depression disease, the aim of this project is to develop multimodal modeling of the limb amygdala and its network from MR imaging combining activation and rest functional imaging and MR brain microstructure quantitative imaging (diffusion and relaxometry). The development of this model will allow us to define three imaging biotypes corresponding to depressed adult patients responding to antidepressant treatments, depressed resistant patients and controls. These multimodal imaging biomarkers will be used to stratify a large longitudinal cohort of young adults into three sub-groups, in order to retrospectively identify early differences in development trajectories of amygdala.
10.4.6 Brain modeling from multi-scale, multimodal and dynamic graphs and development of statistical prediction models
Participants: Julie Coloigner.
Funding: Rennes Métropole, Allocation d'installation scientifique - Duration: 2020-2023 - Budget: 10k€
The human brain is organized into a complex network of billions of neurons, each connected to 100 000 others through axons constituting the bundles of white matter fibers. Over the last decade, some research projects such as the human connectome project have proposed to map the connections between neural pathways that underlie brain function and behavior, in order to improve our understanding of brain function. Mapping these neural connections is crucial to investigate the neuronal underpinnings of the healthy as well as pathological brain. In addition, mathematical modeling from graph theory provides an extremely powerful approach in the study of brain networks. In this project, we propose to develop a new, more robust statistical prediction model based on a graph-based connectivity model that predicts the progression of a new patient’s disease as well as its possible evolution. This model will stratify different states of the diseases in order to reduce heterogeneity and move away from longitudinal cohorts. The developed approach will be tested on local and online cohorts of patients. Moreover, an innovative multimodal cohort of healthy volunteers was funded by this grant.
10.4.7 ANR "MAIA" Multiphysics image-based AnalysIs for premature brAin development understanding
Participants: Pierre Maurel, Antoine Legouhy, Olivier Commowick, Isabelle Corouge, Jean-Christophe Ferré.
Funding: ANR, generic projects program - Duration: 2016-2021 - Budget: 150k€ - PI: F. Rousseau, IMT Atlantique, Brest
Each year in France, 55 000 children are born prematurely, i.e., before the 37th week of gestation. Long-term studies of the outcome of prematurely born infants have clearly documented that the majority of such infants may have significant motor, cognitive, and behavioral deficits. However, there is a limited understanding of the nature of the cerebral abnormality underlying these adverse neurologic outcomes. In this context, the emergence of new modalities of 3D functional MRI, e.g., Arterial Spin Labeling(ASL) or optical imaging technologies, e.g., Near InfraRed Spectroscopy (NIRS), brings new perspectives for extracting cognitive information, via metabolic activity measures. Other classical techniques devoted to cerebral signal measurement, such as ElectroEncephaloGraphy (EEG), provide cognitive information at the cortical level. Each of these various non-invasive imaging technologies brings substantial and specific information for the understanding of newborn brain development.
This project is developing innovative approaches for multi-image / multi-signal analysis, in order to improve neurodevelopment understanding methods. From a fundamental point of view, mathematics and computer science have to be considered in association with imaging physics and medicine, to deal with open issues of signal and image analysis from heterogeneous data (image, signal), considered in the multiphysics contexts related to data acquisition (magnetic, optic, electric signals) and biophysics modeling of the newborn brain. A sustained synergy between all these scientific domains is then necessary. Finally, the sine qua non condition to reach a better understanding of the coupled morphological cognitive development of premature newborns, is the development of effective software tools, and their distribution to the whole medical community. The very target of this project is the design of such software tools for medical image / signal analysis, actually operational in clinical routine, and freely available. Academic researchers and industrial partners are working in close collaboration to reach that ambitious goal.
10.4.8 Hybrid EEG/MRI Neurofeedback for rehabilitation of brain pathologies
Participants: Élise Bannier, Isabelle Bonan, Isabelle Corouge, Jean-Christophe Ferré, Jean-Yves Gauvrit, Pierre Maurel, Mathis Fleury, Giulia Lioi.
Funding: Fondation pour la recherche médicale (FRM) - Duration: 2017-2021 - Budget: 370k€
This project is a continuation of the HEMISFER project ("Hybrid Eeg-MrI and Simultaneous neuro-FEedback for brain Rehabilitation") conducted at Inria Rennes with the support of the Labex "CominLabs".
The goal of this project is to make full use of neurofeedback (NF) paradigm in the context of brain rehabilitation. The major breakthrough will come from the coupling associating functional and metabolic information from Magnetic Resonance Imaging (fMRI) to Electro-encephalography (EEG) to “optimize” the neurofeedback protocol. We propose to combine advanced instrumental devices (Hybrid EEG and MRI platforms), with new hybrid Brain computer interface (BCI) paradigms and new computational models to provide novel therapeutic and neuro-rehabilitation paradigms in some of the major mental and neurological disorders of the developmental and the aging brain (e.g. stroke, language disorders, Mood Depressive Disorder (MDD)). Though the concept of using neurofeedback paradigms for brain therapy has somehow been experimented recently (mostly through case studies), performing neurofeedback through simultaneous fMRI and EEG has almost never been done before so far (two teams in the world including us within the HEMISFER CominLabs project). This project will be conducted through a very complementary set of competences over the different involved teams: Empenn U1228, HYBRID and PANAMA Teams from Inria/Irisa Rennes and EA 4712 team from University of Rennes 1.
10.4.9 Knowledge addition through Neuroimaging of Alcohol consumption in healthy young VOlunteers, causes or consequences
Participants: Elise Bannier, Quentin Duché, Gabriel Robert.
Funding: INCR - Duration: 2021-2022 - Budget: 45k€ - PI: E.Bannier
Alcohol consumption is responsible for 3 million annual deaths worldwide (5.1 percent of the global burden of disease). It causes disease (liver cirrhosis, cancers, etc.) and other social costs (injuries, road accidents, alcohol dependence, etc.). Excessive alcohol consumption grows through adolescence. This type of behavior has also been shown to have subtle but significant deleterious effects on cognitive function in adolescents. Advances in the field of neuroimaging make it possible to characterize anatomical changes and the evolution of neuropsychological deficits. Besides, focusing on the societal causes of alcohol abuse, a large body of studies show that exposure to alcohol advertising through media bootstraps early consumption initiation, greater desire to drink, increased alcohol use and binge drinking patterns among young people, especially minors. We aim to combine the analysis of the locally acquired IMAJ dataset (PI Karine Gallopel-Morvan, INCA Funding) and data from the european consortium IMAGEN datasets to determine whether there are functional characteristics and external factors that can explain behavior towards alcohol and to extract biomarkers capable of predicting excessive behavior. Relying on the IMAJ dataset, we will analyze whether, depending on warning formats displayed on ads (small and text-only vs. larger, shock-inducing and pictorial), health messages can influence brain activity by decreasing the effect of attractive alcohol content ads on the reward system area and on behavioral responses. Relying on the already effective collaboration of Dr Robert with Prof Schumann, we will explore the longitudinal anatomical and functional data from the IMAGEN cohort to extract biomarkers of alcohol consumption evolution and complement the analysis with the results obtained from the IMAJ dataset.
10.4.10 PHRC EMISEP: Evaluation of early spinal cord injury and late physical disability in Relapsing Remitting Multiple Sclerosis
Participants: Élise Bannier, Emmanuel Caruyer, Benoit Combès, Olivier Commowick, Gilles Edan, Jean-Christophe Ferré.
Funding: PHRC - Duration: 2016-2021 - Budget: 200k€
Multiple Sclerosis (MS) is the most frequent acquired neurological disease affecting young adults (1 over 1000 inhabitants in France) and leading to impairment. Early and well adapted treatment is essential for patients presenting aggressive forms of MS. This PHRC (Programme hospitalier de recherche clinique) project focuses on physical impairment and especially on the ability to walk. Several studies, whether epidemiologic or based on brain MRI, have shown that several factors are likely to announce aggressive development of the disease, such as age, number of focal lesions on baseline MRI, clinical activity. However, these factors only partially explain physical impairment progression, preventing their use at the individual level. Spinal cord is often affected in MS, as demonstrated in postmortem or imaging studies. Yet, early radiological depiction of spinal cord lesions is not always correlated with clinical symptoms. Preliminary data, on reduced number of patients, and only investigating the cervical spinal cord, have shown that diffuse spinal cord injury, observed via diffusion or magnetisation transfer imaging, would be correlated with physical impairment as evaluated by the (EDSS) Expanded Disability Status Scale score. Besides, the role of early spinal cord affection (first two years) in the evolution of physical impairment remains unknown.
In this project, we propose to address these different issues and perform a longitudinal study on Relapsing Remitting Multiple Sclerosis (RRMS) patients, recruited in the first year of the disease. Our goal is to show that diffuse and focal lesions detected spinal cord MRI in the first two years can be used to predict disease evolution and physical impairment at 5 years. Twelve centers are involved in the study to include 80 patients.
To date, all subjects have been included. The EMISEP data consists of brain and spinal cord structural and quantitative MR images of early MS patients followed over 5 years. From November 2016 to August 2020, B. Combès processed and analyzed the data corresponding to the two first years of follow-up. Four papers have been published so far and three additional papers are under submission or in preparation. Haykel Snoussi defended his PhD Thesis on diffusion imaging in the spinal cord starting with distortion correction.
10.4.11 Estimating the impact of multiple sclerosis lesions in motor and proprioceptive tracts, from the brain to the thoracic spinal cord, on their functions, assessed from clinical tests (MS-TRACTS and MAP-MS)
Participants: Élise Bannier, Benoit Combès, Malo Gaubert.
Funding: ARSEP, COREC and INCR - Duration: 2020-2022 - Budget: 200k
Previous studies, whether epidemiologic or based on brain MRI, have shown that several factors were likely to announce aggressive development of the disease, such as age, clinical relapses, number of focal lesions on baseline MRI. However, these factors only partially explain physical disability progression, preventing their use at the individual level. We hypothesize that a fine assessment of damage on specific networks, from the brain to the thoracic cord, offers a relevant biomarker of disability progression in MS. Such damage assessments must take into account both lesion location, assessed on structural brain and cord MR images and lesion severity, assessed using advanced brain and cord imaging through quantitative MRI.We propose to test this hypothesis by combining assessments of lesion location and severity on corticospinal and proprioceptive tracts from the brain to the thoracic cord with clinical and () electrophysiological measurements. The MS-TRACTS study involves two French centers (Rennes,Marseille) and includes a total of 60 relapsing remitting MS patients. The expected outcome is to obtain early biomarkers of physical impairment evolution in RRMS patients, first treated with immunomodulatory treatment. The long-term goal is to provide the clinician with biomarkers able to anticipate therapeutic decisions and support the switch to alternative more aggressive treatment. Inclusions are ongoing. The MAP-MS study involves the same two French centers and will nclude 40 progressive MS patients. The investigation will focus on motor asymetry in these more advanced patients.
This study includes two French centers (Rennes, Marseille) and includes a total of 60 patients. The expected outcome is to obtain early biomarkers of physical impairment evolution in RRMS patients, first treated with immunomodulatory treatment. The long-term goal is to provide the clinician with biomarkers able to anticipate therapeutic decisions and support the switch to alternative more aggressive treatment. Inclusions are ongoing.
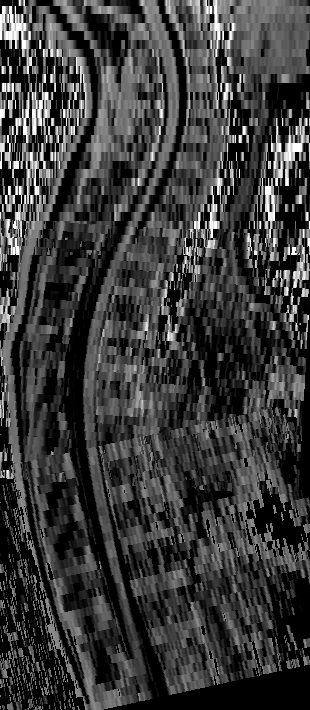

10.4.12 PIA projects
France Life Imaging (FLI)
Participants: Olivier Commowick.
Funding: FLI - Duration: 2012-2023 - Total budget: 2000k€ (phase 1) + 1200k€ (phase 2) + 800k€ (phase 3)
France Life Imaging (FLI) is a large-scale research infrastructure project to establish a coordinated and harmonized network of biomedical imaging in France. This project was selected by the call “Investissements d’Avenir - Infrastructure en Biologie et Santé”. One node of this project is the node Information Analysis and Management (IAM), a transversal node built by a consortium of teams that contribute to the construction of a network for data storage and information processing. Instead of building yet other dedicated facilities, the IAM node use already existing data storage and information processing facilities (LaTIM Brest; CREATIS Lyon; CIC-IT Nancy; Empenn U1228 Inria Rennes; CATI CEA Saclay; ICube Strasbourg) that increase their capacities for the FLI infrastructure. Inter-connections and access to services are achieved through a dedicated software platform that is developed based on the expertise gained through successful existing developments. The IAM node has several goals. It is building a versatile facility for data management that inter-connects the data production sites and data processing for which state-of-the-art solutions, hardware and software, are available to infrastructure users. Modular solutions are preferred to accommodate the large variety of modalities acquisitions, scientific problems, data size, and to be adapted for future challenges. Second, it offers the latest development that are made available to image processing research teams. The team Empenn fulfills multiple roles in this nation-wide project. Olivier Commowick is participating in the working group workflow and image processing and Michael Kain is the technical manager. Apart from the team members, software solutions like MedInria and Shanoir are part of the software platform.
OFSEP: French Multiple Sclerosis Observatory
Participants: Élise Bannier, Olivier Commowick, Gilles Edan, Jean-Christophe Ferré, Francesca Galassi, Arthur Masson, Benoît Combès, Brandon Le Bon.
Funding: ANR-PIA - Duration: 2017-2021 - Budget: 175k€
The French Observatory of Multiple Sclerosis (OFSEP) is one of ten projects selected in January 2011 in response to the call for proposal in the “Investissements d’Avenir - Cohorts 2010” program launched by the French Government. It allows support from the National Agency for Research (ANR) of approximately 10 million € for 10 years. It is coordinated by the Department of Neurology at the Neurological Hospital Pierre Wertheimer in Lyon (Professor Christian Confavreux), and it is supported by the EDMUS Foundation against multiple sclerosis, the University Claude Bernard Lyon 1 and the Hospices Civils de Lyon. OFSEP is based on a network of neurologists and radiologists distributed throughout the French territory and linked to 61 centers. OFSEP national cohort includes more than 50,000 people with Multiple Sclerosis, approximately half of the patients residing in France. The generalization of longitudinal monitoring and systematic association of clinical data and neuroimaging data is one of the objectives of OFSEP in order to improve the quality, efficiency and safety of care and promote clinical, basic and translational research in MS. For the concern of data management, the Shanoir platform of Inria has been retained to manage the imaging data of the National OFSEP cohort in multiple sclerosis.
One long term objective of the OFSEP project is to identify prognostic factors of the evolution of Multiple Sclerosis. The HD Cohort is an enhanced cohort specifically designed for this purpose in which some patients are followed-up on a yearly basis. Additional clinical, quality of life and other patient-reported data is also collected. This study aims at developing personalized predictive tools to improve patient care management, and help in making decision to start, maintain or adapt medical care. Collected data will be processed to extract valuable information enabling to determine specific biomarkers of the evolution of the disease. Multiple Sclerosis brain lesions are of particular interest, hence the need for a careful comparison of lesion segmentation methods. A litterature review enabled to gather most promising cross-sectionnal methods, designed to identify and localize lesions with precise measurement of the lesion load at one particular point in time ; and longitudinal methods which gives more insight on the evolution of those lesions over the different time points. Those later methods are particularly interesting for clinicians for whom the type of lesion evolution is of foremost importance. A ground truth has been carefully designed with images taken from a group of 100 patients selected from the HD Cohort. Those images were meticulously segmented by four experts and the obtained semgentations where reviewed by another experienced radiologist. Four cross-sectionnal methods and one longitudinal methods where trained and evaluated to select the ones which will be used to analyze the entire HD Cohort dataset.
10.5 Regional initiatives
10.5.1 Integration of a MRI- and EEG-compatible NIRS system for the exploration of brain function
Participants: Isabelle Corouge, Elise Bannier, Emmanuel Caruyer, Nolwenn Jégou.
Funding : Conseil Régional de Bretagne via Biogenouest - Duration: 2021 - 2023.
Here, we aim to develop our "multimodal imaging" activity, by jointly measuring electrophysiological (EEG) and hemodynamic (NIRS) brain activity, using functional imaging protocols combining functional MRI (fMRI), EEG and NIRS. The integration of these different types of imaging will allow us to access high spatial and temporal resolution imaging, and thus increase our ability to explore brain activity and identify biomarkers that are more sensitive to physiopathological changes. This funding enabled the recruitment of a research engineer who will develop the experimental NIRS-MRI platform as well as the software suite for the processing of NIRS data alone in a first time and of joint NIRS-MRI or NIRS-EEG data in a second time.
10.5.2 Region Bretagne: project VARANASI
Participants: Camille Maumet, Pierre Maurel, Xavier Rolland.
Funding : Budget: 0.5 PhD thesis
Thanks to the development of open science practices, more and more public datasets are available to the research community. In the field of brain imaging, these data, combined, bring a critical increase in sample size, necessary to build robust models of the typical and atypical brain. However, in order to build valid inferences on these data, we need to take into account their heterogeneity. Variability can arise due to multiple factors such as: differences in imaging instruments, in acquisitions protocols and even, in post-processing pipelines. In particular, the expansion of open source machine learning workflows creates a multitude of possible outputs out of the same dataset. The variations induced by this methodological plurality can be referred to as ‘analytic variability’ which will be the focus of the thesis funded in half by region Bretagne. The thesis of Xavier Rolland (2018-2021) addresses two challenges: 1) How to combine neuroimaging data generated by different analysis pipelines? 2) How to publish neuroimages with an adequate level of metadata to enable their reuse? Methodological developments will combine machine learning techniques with methods from knowledge representation.
11 Dissemination
11.1 Promoting scientific activities
11.1.1 Scientific events: organisation
General chair, scientific chair
- Emmanuel Caruyer and Raphaël Truffet co-organized the "Diffusion-Simulated Connectivity Challenge (DiSCo)" at MICCAI 2021 (virtual).
- Olivier Commowick was General Chair of the “New Multiple Sclerosis Lesions Segmentation Challenge”, at MICCAI 2021.
Member of the organizing committees
- Elise Bannier organized two virtual meetings (March 18th and September 29th) on MRI in clinical research - methodological and pratical aspects in the context of the REMI Network.
11.1.2 Scientific events: selection
Chair of conference program committees
- Pierre-Yves Jonin was Chair of the KeyNote Lecture by Prof. Jacques Grégoire, 4ème Congrès de Neuropsychologie Clinique, Rennes, October, 7th.
Member of the conference program committees
- Pierre-Yves Jonin was m.ember of the Scientific Committee, 4ème Congrès de Neuropsychologie Clinique, Rennes, October, 7th.
- Elise Bannier is board member of the REMI Network and of the SFRMBM society.
Reviewer
- Information Processing in Medical Imaging - IPMI (Olivier Commowick, Claire Cury)
- IEEE International Symposium on Biomedical Imaging - ISBI (Olivier Commowick, Francesca Galassi)
- Medical image computing and Computer assisted intervention - MICCAI (Olivier Commowick, Claire Cury)
- Workshop on Biomedical Image Registration - WBIR (Olivier Commowick)
- Annual congress of the Organisation of Human Brain Mapping - OHBM (Camille Maumet)
- Société Française de Résonance Magnétique en Biologie et Médecine - SFRMBM (Elise Bannier)
- European Society for Magnetic Resonance in Medicine and Biology - ESMRMB(Elise Bannier)
11.1.3 Journal
Member of the editorial boards
- Camille Maumet is member of the Editorial Board of Neuroinformatics.
- Benoit Combès and Olivier Commowick are guest associate editors of the Frontiers in neuroscience research topic on new MS lesions segmentation.
Reviewer - reviewing activities
- Medical Image Analysis (2 papers, Emmanuel Caruyer, Olivier Commowick)
- Magnetic Resonance in Medicine (2 papers, Emmanuel Caruyer)
- IEEE Transactions in Medical Imaging (1 paper, Francesca Galassi)
- Neuroimage (3 papers, Emmanuel Caruyer, Olivier Commowick, Camille Maumet)
- Brain Sciences, MDPI (1 paper, Claire Cury)
- Frontiers in Human Neuroscience, Frontiers (1 paper, Claire Cury)
- Frontiers in Neuroscience, Frontiers (2 papers, Claire Cury)
- Frontiers in Neurology (1 paper, Emmanuel Caruyer)
- Journal of Neuroscience Methods, Elsevier (2 papers, Claire Cury)
- Journal of Machine Learning in Biomedical Imaging (MELBA) (1 paper, Claire Cury)
- Memory (1 paper, Pierre-Yves Jonin)
- Revue de Neuropsychologie (1 paper, Pierre-Yves Jonin)
- Neurocase (4 papers, Pierre-Yves Jonin)
- Nature Communications (1 paper, Camille Maumet)
11.1.4 Invited talks
- Camille Maumet, "Données ouvertes : comment permettre leur réutilisation ?" Rencontres du Réseau d’Entraide Multicentrique en IRM (REMI), Lyon, France, Online, March 2021 45.
- Camille Maumet, "EU Cost Action GliMR — European multi-site data integration & large dataset creation for glioma diagnostics" Reunion du EORTC imaging group, Online, March 2021 46.
- Camille Maumet, "Effort involved in truly FAIR neuroimaging: Towards community-driven research" INCF neuroinformatics Assembly, Online, April 2021 47.
- Benoit Combès, "Apports de l’IA pour la prise en charge des patients ayant une SEP" Journée Francaise de Radiologie, October 2021.
- Benoit Combès, "Outils d'aide à la détection de nouvelles lésions dans un contexte clinique" Journée ARSEP-IRM, February 20.
- Pierre-Yves Jonin, "Statistiques du cas unique en neuropsychologie", Association des Neuropsychologues de Lorraine, Nancy, Nov, 25th.
- Pierre-Yves Jonin, "Superior explicit memory after extensive damage to the medial temporal lobe", Hippocampus Subfield Group Webinar, May, 26th.
- Pierre-Yves Jonin, "Neuroplasticité et mémoire déclarative", Symposium des Journées de Neurologie de Langue Française, May, 26th.
- Pierre-Yves Jonin, "Ultra-short periods of wakeful rest promotes viewpoint invariance in familiarity for visual objects", Journées du GDR Mémoire CNRS, October, 13th.
- Pierre-Yves Jonin, "Developmental Topographical Disorientation Syndrome: neural substrates", Journées du GDR Mémoire CNRS, October, 14th.
- Elise Bannier, Claire Cury and Pierre Maurel, "Bimodal Neurofeedback", University of Geneva on May 18th 2021.
- Elise Bannier "Lecture on Clinical research multicenter imaging specificities - data analysis and sharing", French Clinical Research Infrastructure Network F-CRIN in June 2021.
11.1.5 Leadership within the scientific community
- Camille Maumet is past chair of the Open Science special interest group of the international Organization of Human Brain Mapping. This group is known for the organization of the Open Science Room and the OHBM Brainhack, two international events for the open neuroscience community.
- Camille Maumet is member (by selection) of the national committee on Open Science, Working group "open software" led by Roberto Di Cosmo and François Pellegrini.
11.1.6 Scientific expertise
- Claire Cury was external reviewer for ANR proposal.
- Claire Cury was external reviewer for PhD grant funded by the ANR, University of Lyon.
11.2 Teaching - Supervision - Juries
11.2.1 Teaching
- ESIR, École Supérieure d’Ingénieur de Rennes:
- Pierre Maurel, General image processing (30h),
- Pierre Maurel, Algorithmics and complexity (30h),
- Pierre Maurel, Medical imaging (30h).
- Julie Coloigner, Analyse Multimodale de Signaux Biomédicaux (12h).
- Claire Cury, Traitement avancé des images (Plenary : 12h).
- Claire Cury, Statistiques (TD : 8h).
- Claire Cury, Base de données (TP : 32h).
- Francesca Galassi, SI, M1, "IA en mode projet" (Responsable, 20h).
- Francesca Galassi, SI, M1, "Convolutional Neural Networks" (Plenary: 4h).
- Francesca Galassi, SI, M1, "Data Mining" (TP: 30h).
- Francesca Galassi, IN, M2, "Imagerie Medicale" (TP: 15h).
- Francesca Galassi, "Algorithmique des graphs" (TD: 12h TP: 12h).
- Francesca Galassi, "Algorithmique et Complexité " (TP: 12h).
- Francesca Galassi, "Industrial Internship supervision".
- Caroline Pinte, "Mathématiques appliquées au traitement d'images (MATI)" (TP: 18h).
- Xavier Rolland, "Statistiques" (TD/TP: 10h).
- Master SIBM, M2, University of Angers-Brest-Rennes:
- Jean-Christophe Ferré is head of the master.
- Benoît Combès is co-head of the UE “Modélisation et Apprentissage Automatique pour le Traitement des Images Médicales”.
- Emmanuel Caruyer, "Méthodes d'analyse d'IRM de diffusion" (Plenary: 3h).
- Julie Coloigner, "Méthode d'analyse de la connectivité cérébrale" (Plenary: 3h).
- Benoit Combès, "Méthodes de segmentation pour l'imagerie médicale" (Plenary: 3h).
- Benoit Combès, “Méthodes de recalage linéaire et non-linéaires des images médicales” (Plenary: 6h).
- Benoit Combès, “Applications des méthodes de traitement des images médicales” (Plenary: 3h).
- Benoit Combès, “Eléments de statistiques pour l’induction scientifique” (Plenary: 4.5h).
- Benoit Combès, Camille Maumet, “Soutenance de présentations critiques d'articles scientifiques” (TD: 3h).
- Isabelle Corouge, "IRM de perfusion par Arterial Spin Labeling (ASL)" (Plenary: 3h).
- Camille Maumet, "Workflows de traitements d'images" (Plenary: 3h).
- Elise Bannier, "Imagerie fonctionnelle cérébrale" (Plenary: 1h).
- Quentin Duché, "Traitement des données d'IRM fonctionnelle" (Plenary: 1h).
- Elise Bannier, “Utilisation et réutilisation des données d'imagerie” (Plenary: 1h).
- L3 SIF, L3, ENS Rennes/University of Rennes 1:
- Emmanuel Caruyer, "Méthodes numériques pour le traitement d'images" (Plenary: 20h).
- Licence BECV (Biologie, Environnement, Chimie du Vivant), University of Rennes 1: Elodie Germani, Practical work to learn Python in Mathematics for Biologist (TP: 18h).
- Master mention Informatique, parcours Science informatique (SIF), M2, University of Rennes: Julie Coloigner, Computer Vision (Plenary: 10h).
- Bachelor for speech therapy, L3, University of Rennes: Elise Bannier, "Imagerie fonctionnelle cérébrale du langage" (Plenary: 2h).
- Diplome Universitaire MERC (Manipulateur en Recherche Clinique), University of Montpellier: Elise Bannier, "Spécificités de la recherche clinique en imagerie" (Plenary: 7h).
- ENS Rennes: Pierre Maurel, Introduction to image processing (20h).
- Francesca Galassi, Inria Summer School in AI, "The Problem of Domain Shift" (Plenary: 2h).
- Master Neuropsychologie, M2, University of Savoie: Pierre-Yves Jonin, "Limites méthodologiques du bilan neuropsychologique à visée diagnostique" (Plenary: 3h30).
- Master Psychologie et neuropsychologie de l’enfant et de l’adulte : langage, cognition et apprentissage, M2, University of Poitiers: Pierre-Yves Jonin, "Méthodologie de l'étude de cas" (Plenary: 3h).
- Master Biologie et Santé, M1, University of Bretagne Occidentale: Pierre-Yves Jonin, "Explorations neuropsychologiques des maladies neurologiques et psychiatriques" (Plenary: 4h).
- Master Psychologie Clinique, Psychopathologie et Psychologie de la Santé, M2, University of Rennes 2:
- Pierre-Yves Jonin, "Neuropsychologie clinique des pathologies neurodégénératives" (Plenary: 4h).
- Pierre-Yves Jonin, "Méthodologie de l'étude de cas" (Plenary: 4h).
- Licence Psychologie, L3, University of Rennes 2:
- Pierre-Yves Jonin, "Les syndromes neuropsychologiques" (TD: 16h).
- Pierre-Yves Jonin, "Approche neuropsychologique du handicap" (TD: 4h).
- Master Neurosciences Cliniques, M2, University of Rennes 1: Pierre-Yves Jonin, "Neurosciences cognitives et cliniques de la mémoire humaine" (Plenary: 3h).
11.2.2 Supervision
PhD
- PhD in progress: Thomas Durantel, "Diffusion MRI tractography with prior knowledge", from Sept 2021, O. Commowick and J. Coloigner.
- PhD in progress: Jean-Charles Roy, "Apathy in Late Life Depression: New Biomarkers Using Actimetry and Magnetic Resonance Imaging", from Nov 2020, G. Robert and J. Coloigner.
- PhD in progress: Caroline Pinte, “Methodology for enhanced and adapted Neurofeedback training”, Univ. Rennes, from Oct 2021, Claire Cury and Pierre Maurel.
- PhD in progress: Lisa Hemforth, "Methodology for automatic scoring of Incomplete Hippocampal Inversion", Sorbonne University, from Oct. 2021, Claire Cury, Baptiste Couvy-Duchesne and Olivier Colliot.
- PhD in progress: Xavier Rolland, “Modeling analytic variability in brain imaging”, CNRS, from Oct 2018, Camille Maumet and Pierre Maurel.
- PhD in progress: Elodie Germani, “Mapping the fMRI pipeline-space towards more robust pipelines”, University of Rennes 1, from Oct 2021, Camille Maumet and Elisa Fromont.
- PhD in progress: Stéphanie Leplaideur, “The neural basis of egocentric spatial representation and neck muscle vibration effects on spatial cognition, in healthy subjects and patients who had a right stroke”, from Oct 2018 (Medical Doctor, on part time), Isabelle Bonan and Elise Bannier
- PhD in progress: Ambre Godet "fNIRS neurofeedback implementation in the context of food addiction - brain (fNIRS/MRI) and behavioral caracterisation", from Oct. 2021, Elise Bannier, Nicolas Coquery, David Val-Laillet.
Other supervisions
- L3 SIF, ENS Rennes/UR1: Étienne Objois, "Estimation des paramètres en IRM de diffusion multi-dimensionnelle", supervised by Emmanuel Caruyer.
- Co-supervised project: Alexander Bowring, "Exploring what part of the fMRI pipeline has the strongest impact on Task fMRI Results", Camille Maumet in collaboration with Thomas Nichols (University of Oxford).
- M2 SIBM, UR1: Quentin Frecelle, "Méthodes de quantification des slowly evolving lesions", supervised by Benoit Combès and Arthur Masson.
- M2 SIBM, UR1: Nora El Graoui , "Exploration multimodale de l’activité cérébrale motrice par NIRS et IRM BOLD", supervised by Elise Bannier.
- L3 Physique Médicale, UR1: Samia Djaiji , "Automatisation des procédures de contrôle qualité IRM de l’appareil d’IRM de recherche Neurinfo ", supervised by Elise Bannier.
- M2 SIBM, UR1: Ismail Abaakil, "Exploration comparative bimodale de lctivit crbrale fonctionnelle au repos par IRM fonctionnelle BOLD et par spectroscopie du proche infrarouge fonctionnelle NIRSf ", supervised by Isabelle Corouge.
- M2 ESIR, University of Rennes: Caroline Pinte, "Bi-modal neurofeedback: Predicting fMRI neurofeedback scores from EEG signals alone using automatic learning techniques", supervised by Claire Cury and Pierre Maurel.
- M1 University of Rennes: Tristan Calas, "The use of eye-tracking and skin conductance measurements to measure different mental states of a subject.", supervised by Claire Cury.
- M2 University Paris Saclay: Alexandre Martin, "Deep learning pour la cotation d’un variant atypique de l’hippocampe", supervised by Olivier Colliot, Claire Cury and Baptiste Couvy-Duchesne.
- M2 Sorbonne University: Kevin De Matos, "Assessment of temporal lobe sulcal patterns", supervised by Olivier Colliot, Claire Cury and Baptiste Couvy-Duchesne.
- M2 ESIR, UR1: Chloé Mercier, "IRM de diffusion et relaxométrie: vers une utilisation combinée" , supervised by Olivier Commowick.
- M2 Bioinformatique, UR1: Elodie Germani, "Varibilité analytique en IRMf: combien de participants pour une mesure robuste ?", supervised by Camille Maumet.
- M1 Neurosciences Cliniques, UR1: Mélissa Brossais, "The role of visual expertise and prior knowledge in familiarity for faces", supervised by Pierre-Yves Jonin.
11.2.3 Juries
- Claire Cury, reviewer and jury for Inria Grants Moyen incitatifs 2021 (PhD, postdoc and ADT), Inria Rennes.
- Camille Maumet. PhD committee: Elina Thibeau-Sutre, Sorbonne Université, France, December 2021.
- Olivier Commowick. PhD referee: J. Benzakoun, Université de Paris, France, December 2021.
- Olivier Commowick. HDR Jury: Gabriel Robert, Université de Rennes 1.
11.3 Popularization
11.3.1 Internal or external Inria responsibilities
- Claire Cury, Scientific mediation Officer of the Inria Rennes Scientific mediation team.
- Camille Maumet is a co-organizer of the local version of the program "L codent L créent", an outreach program to send PhD students to teach Python to middle school students in 8 sessions of 45 minutes. It was initiated in Lille, with Anne-Cecile Orgerie and Tassadit Bouadi. The program is currently supported by: Alstom, Fondation Blaise Pascal, ED MathSTIC, Inria and Fondation Rennes 1.
11.3.2 Articles and contents
- Camille Maumet, Elise Bannier, chapter "L'imagerie au service de la santé : questions éthiques et sociétales" in the book "Le corps en images : Les nouvelles imagerie pour la santé", in press, chapter written in collaboration with Anne Hespel (CHU Rennes).
- Pierre-Yves Jonin, "Quand la mémoire flanche", Magazine Sciences Ouest, October 2021
- Pierre-Yves Jonin, "La mémoire, comment ça marche?", Podcast cognitif, December 2021
- Pierre-Yves Jonin, "Des troubles du langage à la maladie d'Alzheimer", Le Magazine des chercheurs en santé du CHU de Rennes, June 2021
11.3.3 Education
- Elodie Germani, Caroline Pinte, Camille Maumet. L codent L créent - Teaching middle schoolers at Collège Le Landry, Rennes. Animation of practical session that aim to initiate young girls to programming and to show them that careers in computer science are accessible.
- Elodie Germani, Isabelle Corouge: "J'peux pas, j'ai informatique". Animation of workshops dedicated to "computing without computers" for teachers in middle schools.
11.3.4 Interventions
- Camille Maumet, Panelist in the panel discussion "Les données de santé : un trésor convoité" Salon Innovation Santé CITY HEALTHCARE, Nancy, October 2021.
- Elise Bannier, Benoit Combès, Olivier Commowick, Malo Gaubert: Journée ARSEP des patients, CHU de Rennes, November 2021 : day to present current research on medical imaging in MS to patients.
- Benoit Combès, "Améliorer la détection des nouvelles lésions SEP". Journée ARSEP des patients, CHU de Rennes, November 2021.
- Benoit Combès, "MSTracts : Vers un meilleur pronostic dans la sclérose en plaques par une exploration complète du système nerveux central". Une présentation parmi une série d'autres présentée lors des Mardis de l'Espaces des Sciences, Rennes, October 2021.
- Quentin Duché, "KENAVO : Apports de la neuroimagerie pour étudier le comportement vis à vis de l’alcool des jeunes adultes, causes ou conséquences ?". Une présentation parmi une série d'autres présentée lors des Mardis de l'Espaces des Sciences, Rennes, 5th October 2021.
- Claire Cury, organisation committee of the Journée Science et Musique, 9th of October 2021, Rennes.
- Pierre-Yves Jonin, "Tour d'horizon de la mémoire humaine", Conférence, Salon de l'innovation de Bains sur Oust, December.
12 Scientific production
12.1 Major publications
- 1 articleUnsupervised Domain Adaptation With Optimal Transport in Multi-Site Segmentation of Multiple Sclerosis Lesions From MRI Data.Frontiers in Computational Neuroscience14March 2020, 1-13
- 2 articleIsolating the Sources of Pipeline-Variability in Group-Level Task-fMRI results.Human Brain MappingNovember 2021
- 3 articleFocal and diffuse cervical spinal cord damage in patients with early relapsing--remitting MS: A multicentre magnetisation transfer ratio study.Multiple Sclerosis Journal258February 2019, 1113-1123
- 4 articleObjective Evaluation of Multiple Sclerosis Lesion Segmentation using a Data Management and Processing Infrastructure.Scientific Reports81December 2018, 13650
- 5 articleA sparse EEG-informed fMRI model for hybrid EEG-fMRI neurofeedback prediction.Frontiers in Neuroscience13January 2020
- 6 articleMultiple sclerosis lesions in motor tracts from brain to cervical cord: spatial distribution and correlation with disability.Brain - A Journal of Neurology 1437July 2020, 2089-2105
- 7 articleThe impact of Neurofeedback on effective connectivity networks in chronic stroke patients: an exploratory study.Journal of Neural Engineering185September 2021, 056052
- 8 articlePatch-Based Super-Resolution of Arterial Spin Labeling Magnetic Resonance Images.NeuroImage189January 2019, 85-94
- 9 articleAssociation of Gray Matter and Personality Development With Increased Drunkenness Frequency During Adolescence.JAMA Psychiatry774April 2020, 409-419
12.2 Publications of the year
International journals
National journals
International peer-reviewed conferences
Conferences without proceedings
Edition (books, proceedings, special issue of a journal)
Doctoral dissertations and habilitation theses
Other scientific publications
12.3 Other
Scientific popularization
