2023Activity reportProject-TeamEMPENN
RNSR: 200518339S- Research center Inria Centre at Rennes University
- In partnership with:CNRS, INSERM, Université de Rennes
- Team name: Neuroimaging: methods and applications
- In collaboration with:Institut de recherche en informatique et systèmes aléatoires (IRISA)
- Domain:Digital Health, Biology and Earth
- Theme:Computational Neuroscience and Medicine
Keywords
Computer Science and Digital Science
- A3.1.2. Data management, quering and storage
- A3.1.3. Distributed data
- A3.1.7. Open data
- A3.1.8. Big data (production, storage, transfer)
- A3.2.4. Semantic Web
- A3.3.3. Big data analysis
- A3.4.1. Supervised learning
- A3.4.2. Unsupervised learning
- A3.4.3. Reinforcement learning
- A3.4.4. Optimization and learning
- A3.4.6. Neural networks
- A3.4.8. Deep learning
- A5.1.4. Brain-computer interfaces, physiological computing
- A5.2. Data visualization
- A5.3.2. Sparse modeling and image representation
- A5.3.3. Pattern recognition
- A5.3.4. Registration
- A5.4.1. Object recognition
- A5.4.6. Object localization
- A5.9.2. Estimation, modeling
- A5.9.4. Signal processing over graphs
- A6.2.3. Probabilistic methods
- A6.2.4. Statistical methods
- A6.3.3. Data processing
- A6.3.4. Model reduction
- A9.2. Machine learning
- A9.3. Signal analysis
Other Research Topics and Application Domains
- B1.2. Neuroscience and cognitive science
- B1.2.1. Understanding and simulation of the brain and the nervous system
- B1.2.2. Cognitive science
- B2.1. Well being
- B2.2.2. Nervous system and endocrinology
- B2.2.6. Neurodegenerative diseases
- B2.5.1. Sensorimotor disabilities
- B2.5.2. Cognitive disabilities
- B2.6.1. Brain imaging
1 Team members, visitors, external collaborators
Research Scientists
- Emmanuel Caruyer [CNRS, Researcher, HDR]
- Julie Coloigner [CNRS, Researcher]
- Benoit Combes [INRIA, Researcher]
- Claire Cury [INRIA, Researcher]
- Fanny Degeilh [INSERM, Researcher]
- Camille Maumet [INRIA, Researcher]
Faculty Members
- Pierre Maurel [Team leader, Université de Rennes, Professor, HDR]
- Isabelle Bonan [Université de Rennes, CHU , Professor, HDR]
- Gilles Edan [Université de Rennes, CHU , Professor, HDR]
- Jean-Christophe Ferré [Université de Rennes, CHU , Professor, HDR]
- Francesca Galassi [Université de Rennes, Associate Professor]
- Jean-Yves Gauvrit [Université de Rennes, CHU , Professor, HDR]
- Anne Kerbrat [Université de Rennes, CHU , Associate Professor, HDR]
- Gabriel Robert [Université de Rennes, CHGR, Professor, HDR]
Post-Doctoral Fellows
- Mathis Fleury [Université de Rennes, until Aug 2023]
- Agustina Fragueiro [INRIA, Post-Doctoral Fellow]
- Renaud Hedouin [INRIA, until Jan 2023]
- Burhan Rashid Hussein [INRIA, Post-Doctoral Fellow]
- Jeremy Lefort-Besnard [INRIA, Post-Doctoral Fellow]
- Cédric Meurée [INRIA]
- Camille Muller [Université de Rennes, Post-Doctoral Fellow, from Oct 2023]
- Benjamin Streichenberger [INRIA, from May 2023]
PhD Students
- Constance Bocquillon [Université de Rennes]
- Sebastien Dam [INRIA]
- Thomas Durantel [Université de Rennes]
- Carlo Ferritto [Université de Rennes, from Oct 2023]
- Elodie Germani [Université de Rennes]
- Nolwenn Jegou [INRIA, from Dec 2023]
- Carla Joud [Université de Rennes]
- Caroline Pinte [Université de Rennes]
- Marie Poirier [Université de Rennes]
- Jean-Charles Roy [INCR, until Oct 2023]
- Ricky Walsh [Université de Rennes]
Technical Staff
- Elise Bannier [CHU RENNES, Engineer, HDR]
- Boris Clenet [INRIA, Engineer]
- Isabelle Corouge [Université de Rennes]
- Pierre-Henri Dauvergne [INRIA, Engineer]
- Rene-Paul Debroize [INRIA, Engineer]
- Jean-Côme Douteau [INRIA, Engineer]
- Quentin Duché [Université de Rennes, Engineer]
- Malo Gaubert [CHU, Engineer]
- Michael Kain [Inria (SED), Engineer]
- Florent Leray [Inria (SED), Engineer]
- Julien Louis [Inria (SED), Engineer]
- Arthur Masson [INRIA, Engineer, until Sep 2023]
- Youenn Merel [INRIA, Engineer, from Mar 2023]
- Sandesh Patil [INRIA, Engineer]
Interns and Apprentices
- Thomas Agasse-Lafont [Université de Rennes, Intern, until May 2023]
- Gaelle Baudron [Université de Rennes, Intern, until Jul 2023]
- Mathys Biziere [INRIA, Intern, until May 2023]
- Louis-Gabriel Capliez [INRIA, Intern, from May 2023 until Jul 2020]
- Vivien Caron [INRIA, Intern, until Jul 2023]
- Alice Dufey [Université de Rennes, Intern, until Aug 2023]
- Léo Filoche [INRIA, Intern, from Jun 2023 until Sep 2023]
- Mathilde Liffran [INRIA, Intern, from Nov 2023]
- Lounès Meddahi [INRIA, Intern]
- Benjamin Prigent [Université de Rennes, Intern, from Mar 2023 until Aug 2023]
- Emma Redor [INRIA, Intern, from May 2023 until Jul 2023]
- Valentin Septiers [INRIA, Intern, from Mar 2023 until Sep 2023]
- Demian Vera [CIFICEN, BUENOS AIRES, Intern]
Administrative Assistant
- Armelle Mozziconacci [CNRS]
Visiting Scientists
- Erick Jorge Canales Rodríguez [EPFL]
- Olivier Coulon [CNRS]
- Tim Dyrby [Technical Universituion of Denmark, from Jan 2023 until Dec 2022]
- Elda Fischi-Gomez [EPFL]
- Laure Fournier [Université Paris Cité, UFR de Médecine, until Jan 2023]
- Rémi Gau [UNIVERSITE CATHOLIQUE LOUVAIN]
- Derek Jones [Cubric, Cardiff University]
- Villarreal Haro Juan Luis [EPFL]
- Jonathan Rafael Patino Lopez [EPFL]
- Jean-Philippe Thiran [EPFL]
2 Overall objectives
The research team Empenn ("Brain" in Breton language) ERL U1228 is co-affiliated with Inria, Inserm (National Institute for Health and Scientific Research), CNRS (INS2I institute), and the University of Rennes I. It is a team of IRISA/UMR CNRS 6074. Empenn is located in Rennes, on the medical and scientific campus. It succeeded in 2019 to the "VisAGeS" team, created in 2006 by Inria. As for "VisAGeS", Empenn holds the accreditation number U1228, renewed by Inserm in 2022 and for a period of 6 years, after an evaluation conducted by the HCERES and Inserm.
Thanks to this unique partnership, Empenn's ambition is to establish a multidisciplinary team of researchers in information sciences and medicine. Our medium and long term objective is to introduce our fundamental research into clinical practice, while maintaining the excellence of our methodological research.
Our goal is to foster research in medical imaging, neuroinformatics and population cohorts. In particular, the Empenn team aims at the detection and development of imaging biomarkers for brain diseases and focuses its efforts on transferring this research to the clinic and clinical neuroscience in general. More specifically, the objective of Empenn is to propose new statistical and computational methods, and to measure and model morphological, structural and functional states of the brain to better diagnose, monitor and treat mental, neurological and substance use disorders. We propose to combine advanced instrumental devices and novel computational models to provide advanced diagnostic, therapeutic, and neurorehabilitation solutions for some of the major developing and aging brain disorders.
Generic and challenging research topics in this broad area include finding new ways to compare models and data, aid in decision making and interpretation, and develop feedback. These activities are carried out in close collaboration with the Neurinfo imaging platform in vivo, which is an essential environment for the experimental implementation of our research on ambitious clinical research projects and the development of new clinical applications.
3 Research program
3.1 Glossary
-
Magnetic Resonance Imaging
- MR - Magnetic Resonance
- MRI - Magnetic Resonance Imaging
- fMRI - Functional Magnetic Resonance Imaging
- DWI - Diffusion-Weighted Imaging
- ASL - Arterial Spin Labeling
-
Other modalities
- PET - Positron Emission Tomography
- EEG - Eletroencephalograpy
- NIRS - Near InfraRed Spectroscopy
-
Medical terminology
- MS - Multiple Sclerosis
- TBI - Traumatic Brain Injury
-
Methodological terminology
- GLM - General Linear Model
- MCM - Multi-compartment models
- NF - Neurofeedback
3.2 Scientific Foundations
The scientific foundations of our team concern the design and development of new computational solutions for biological images, signals and measurements. Our goal is to develop a better understanding of the normal and pathological brain, at different scales.
This includes imaging brain pathologies in order to better understand pathological behavior from the organ level to the cellular level, and even to the molecular level (PET-MR imaging), and modeling of large groups of normal and pathological individuals (cohorts) from image descriptors. It also addresses the challenge of the discovery of episodic findings (i.e., rare events in large volumes of images and data), data mining and knowledge discovery from image descriptors, validation and certification of new drugs from imaging features, and, more generally, the integration of neuroimaging into neuroinformatics by promoting and supporting virtual organizations of biomedical actors using e-health technologies.
Illustration of the major scientific objectives of the Empenn team, including Imaging sources, Cells and tissue structure and function, Brain structure and function, Group study and Patient.
As shown in Figure 1, the research activities of the Empenn team closely link observations and models through the integration of clinical and multiscale data, and phenotypes (cellular, and later molecular, with structural or connectivity patterns in the first stage). Our ambition is to build personalized models of central nervous system organs and pathologies, and to compare these models with clinical research studies in order to establish a quantitative diagnosis, prevent the progression of diseases and provide new digital recovery strategies, while combining all these research areas with clinical validation. This approach is developed within a translational framework, where the data integration process to build the models is informed by specific clinical studies, and where the models are assessed regarding prospective clinical trials for diagnosis and therapy planning. All of these research activities are conducted in close collaboration with the Neurinfo platform, which benefited in 2018 from a new high-end 3T MRI system dedicated to research (3T Prisma™ system from Siemens), and through the development in the coming years of multimodal hybrid imaging (from the currently available EEG-MRI, to EEG-NIRS and PET-MRI in the future).
In this context, some of our major developments and newly arising issues and challenges include:
- The generation of new descriptors to study brain structure and function (e.g. the combination of variations in brain perfusion with and without a contrast agent; changes in brain structure in relation to normal, pathological, functional or connectivity patterns; or the modeling of brain state during cognitive stimulation using neurofeedback).
- The integration of additional spatiotemporal and hybrid imaging sequences covering a larger range of observations, from the molecular level to the organ level, via the cellular level (arterial spin labeling, diffusion MRI, MR relaxometry, MR cell labeling imaging, EEG-MRI functional imaging, EEG-NIRS-MRI).
- The creation of computational models through the data fusion of multimodal MR images, structural and functional image descriptors from group studies of normal and/or pathological subjects.
- The evaluation of these models in relation to acute pathologies, especially for the study of degenerative, psychiatric, traumatic or developmental brain diseases (primarily multiple sclerosis, stroke, traumatic brain injury (TBI) and depression, but applicable with a potential additional impact to epilepsy, Parkinson’s disease, dementia, post-traumatic stress disorder, etc.) within a translational framework.
In terms of new major methodological challenges, we address the development of models and algorithms to reconstruct, analyze and transform the images, and to manage the mass of data to store, distribute and “semanticize” (i.e. provide a logical division of the model’s components according to their meaning). As such, we expect to make methodological contributions in the fields of model inference; statistical analysis and modeling; the application of sparse representation (compressed sensing and dictionary learning) and machine learning (supervised/unsupervised classification and discrete model learning); data fusion (multimodal integration, registration, patch analysis, etc.); high-dimensional optimization; data integration; and brain-computer interfaces. As a team at the frontier between the digital sciences and clinical research in neuroscience, we do not claim to provide theoretical breakthroughs in these domains but rather to provide significant advances in using these algorithms through to the advanced applications we intend to address. In addition, we believe that by providing these significant advances using this set of algorithms, we will also contribute to exhibiting new theoretical problems that will fuel the domains of theoretical computer sciences and applied mathematics.
In summary, we expect to address the following major challenges:
- Developing new information processing methods able to detect imaging biomarkers in the context of mental, neurological, and substance use disorders.
- Providing new computational solutions for our target applications, allowing a more appropriate representation of data for image analysis and the detection of biomarkers specific to a form or grade of pathology, or specific to a population of subjects.
- Providing, for our target applications, new patient-adapted connectivity atlases for the study and characterization of diseases from quantitative MRI.
- Providing, for our target applications, new analytical models of dynamic regional perfusion, and deriving indices of dynamic brain local perfusion from normal and pathological populations.
- Investigating whether the theragnostics paradigm of rehabilitation from hybrid neurofeedback can be effective in some behavioral and disability pathologies.
These major advances are primarily developed and validated in the context of several priority applications in which we expect to play a leading role: multiple sclerosis, stroke rehabilitation, and the study and treatment of depression.
4 Application domains
Figure 2 summarizes the scientific organization of the research team through three basic research topics in information sciences (Population Imaging, Detection and Learning, and Quantitative Imaging) and three translational axes on central nervous system diseases (Behavior, Neuro-inflammation and Recovery).
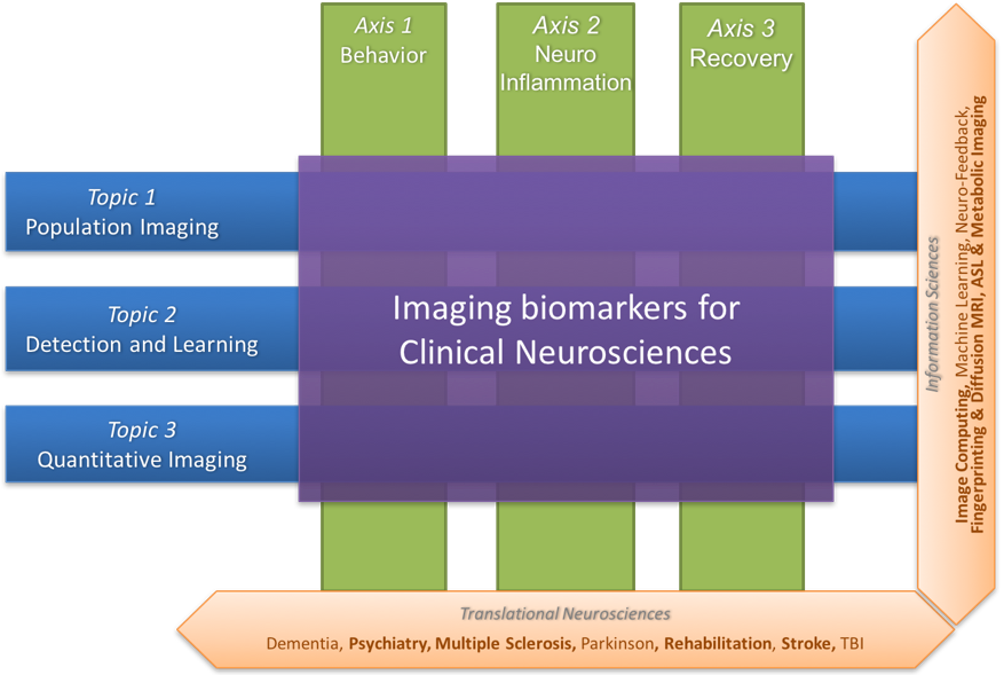
Illustration of the research topics and research axes of the Empenn team. The former (Population imaging, detection and learning and quantitative imaging) concern information sciences, and these topics intersect with the latter (Behavior, neuro-inflammation and recovery), which are translational neurosciences axes.
4.1 Basic research
4.1.1 Population imaging
One major objective of neuroimaging researchers and clinicians is to be able to stratify brain imaging data in order to derive new and more specific population models. In practice, this requires to set up large-scale experiments that, due to the lack of resources and capabilities to recruit locally subjects who meet specific inclusion criteria, motivates the need for sharing the load.
However, building and using multi-site large-scale resources pose specific challenges to deal with the huge quantity of data produced and their diversity. Empenn focuses on two challenges in particular:
- Providing computational environments for the computation and use of imaging biomarkers in the targeted brain diseases, a solution to be used by radiologists and neurologists/psychiatrists for the clinical follow-up of a large patient population.
- Modeling analytic variability of image processing pipelines to better understand and predict the behaviour of imaging biomarker detection solutions and improve reproducibility and productivity in clinical neuroimaging research.
4.1.2 Detection and learning
We intend to make significant contributions with major impacts in learning coupling models between functional recordings during neurofeedback procedures. These advances will provide a breakthrough in brain-computer interfaces for rehabilitation protocols. Our aim is to:
- Our research employs data-driven approaches, encompassing machine learning and deep learning, to enhance the detection and segmentation of abnormal patterns in medical images. Our primary focus is on multiple sclerosis (MS) and, more recently, on stroke. The findings from our studies indicate promising outcomes in automated tools for accurate disease activity assessment and lesion segmentation within large MRI databases. Special attention is given to the integration of multimodal information and the utilization of labeled and unlabeled data. As we progress, our aim is to adapt these methods to address a broader range of neurological diseases, including epilepsy, tumors, etc., in both neonate and adult brains. This research contributes to advancing diagnostic tools and methodologies in the field of medical imaging.
- Develop solutions for combining brain state measurements from multimodal sensors or sequences (e.g. fMRI, ASL, EEG, NIRS, etc.) with applications in the spatiotemporal reconstruction of brain activity from MRI-EEG or the combined detection of the endogenous hemodynamic and resting state network of the brain from ASL and NIRS. Over the longer term, the advent of new hybrid brain imaging sensors (e.g. PET-MRI) will require these methods to be extended to a larger spectrum of information combining structural, morphological, metabolic, electrophysiological and cellular/molecular information (e.g. through the use of specific ligands/nanocarriers).
4.1.3 Quantitative imaging
The Empenn research group focuses on the development of several quantitative techniques in magnetic resonance imaging of the brain. These methods allow for characterization of both the function and the structure of the brain with high precision. Arterial spin labelling (ASL) is a contrast agent-free imaging technique which labels arterial blood water as an endogenous tracer for perfusion and can measure resting-state cerebral blood flow. We are interested in estimating multiparametric hemodynamics using ASL, such as combined cerebral blood flow and arterial transit times, and derive statistical descriptors to represent significant differences between groups. In addition to quantitative perfusion parameters, our contributions on tissue compartment imaging aim at delineating neural circuits and characterize their microstructure properties, using both diffusion MRI and relaxometry. In diffusion MRI, arbitrary gradient waveforms were shown to exhibit higher sensitivity to microstructure parameters than standard pulsed gradients. We work on the optimization of sampling protocols in this domain, with the objective to propose sequences compatible with in vivo acquisition. Complementary to diffusion MRI, we develop methods for the reconstruction of myelin-bound, extra-axonal and cerebrospinal fluid water using multi-compartment modelling of the T2-relaxometry signal. We combine these techniques with tractography to identify trajectories of pathologies associated to the evolution of these microstructural parameters along specific fiber bundles in the brain white matter. Finally, we are also focusing on assessing the characteristics (repeatability, reproducibility and sensitivity) of several quantitative metrics variability (e.g. MTR, T1 relaxometry) in the spinal cord of patients with MS.
4.2 Translational research
4.2.1 Behavior
Advances in the field of in vivo imaging offer new opportunities for addressing the management of resistant affective disorders and their consequences (suicide risk and socio-professional impact), and the management of spatial cognition disorders after stroke and their consequences (postural perturbations and the loss of autonomy). Our objective, and the main challenge in this context, is to introduce medical image computing methods to the multidisciplinary field of behavioral disorders (cognitive disorders, particularly spatial and postural control disorders or anterograde memory impairment, mood disorders, notably resistant depression, schizophrenic disorders, pervasive developmental disorders, attention disorders, etc.) in order to gain a better understanding of the pathology and devise innovative therapeutic approaches.
We also expect to become a major player in the future and make important contributions with significant impacts, primarily in drug-resistant depression in young and old populations. In particular, we expect to provide new image-related metrics combining perfusion, metabolism and microstructural information regarding the brain in order to better characterize pathologies, provide prospective evolution values and potentially provide new brain stimulation targets that could be used in neurofeedback rehabilitation protocols or other types of brain stimulation procedures.
We aim to provide new imaging markers of mental diseases, especially in the context of mood disorders. The new biomarkers are derived from the metabolic (ASL and later ASL+PET) point of view as well as from the microstructural point of view (multicompartment diffusion MRI and relaxometry). Similarly, we expect to exhibit imaging biomarker regularities combining metabolic and structural information. Over the longer term, we expect these biomarkers to be the target of neurofeedback rehabilitation procedures. Also, over the longer term, we expect to supplement the MRI markers with molecular markers coming from new PET tracers, especially those associated with serotonin intake, at one time point or during a rehabilitation protocol under hybrid PET-EEG-MRI neurofeedback procedures.
4.2.2 Neuroinflammation
Some of the major ongoing research issues regarding neuroimaging of neuro-inflammatory diseases concern the definition of new biomarkers to track the development of the pathology using high- dimensional data (e.g. nD+t MRI). This includes the use of white matter-specific imaging, such as magnetization transfer MRI, relaxometry and diffusion-weighted imaging (DW-MRI). Our objective is (1) to develop information-processing tools to tag the spatiotemporal evolutions of Multiple Sclerosis patterns at the brain parenchyma and spinal cord levels from their different signatures (inflammatory cells visible with USPIO or Gd contrast agents on MRI, persistent black holes, eloquent regional atrophy and microstructure signatures); and (2) to test these new tools on new imaging cohorts. In this respect, we for instance conduct studies on brain and spinal cord imaging, continuing on from the PHRC multicentric EMISEP project (PI: G. Edan), as it is very likely that lesions in the spine will directly affect the ambulatory ability of the patient (and thereby the clinical scores). In order to extend this experiment to a larger MS population, based on our expertise from the OFSEP cohort, we also plan to improve the MS therapeutic decision process notably through the RHU PRIMUS (PRojection In MUltiple Sclerosis) project (PI: G. Edan). Our goal is to develop and assess a standardized monitoring tool that provides a robust, long-term computerized MRI follow-up that will become the gold standard in clinical practice for therapeutic decisions in MS treatment. As part of this project, Empenn will share its expertise in data management systems (Shanoir and FLI-IAM), automatic processing tools (through the medInria and Anima software repositories) to extract quantitative indices from the images and the assessment of the added-value of promising quantitative sequences.
4.2.3 Recovery
Mental and neurological disorders are the leading cause of years lived with a disability. Treatment-resistant depression affects approximately 2% of the European population. Meanwhile, in the case of brain disorders, almost 1.5 million Europeans (15 million people worldwide) suffer a stroke event each year. Current recovery methods for brain disorders and traumatic brain injuries remain limited, preventing many from achieving full recuperation. We propose to address the issue of brain recovery by introducing new advances from recent breakthroughs in computational medical imaging, data processing and human-machine interfaces, and demonstrate how these new concepts can be used, in particular for the treatment of stroke and major depressive disorders.
We ambition to combine advanced instrumental devices (hybrid EEG, NIRS and MRI platforms), with new hybrid brain computer interface paradigms and new computational models to provide neurofeedback-based therapeutic and neuro-rehabilitation paradigms in some of the major mental and neurological disorders of the developmental and aging brain.
Neurofeedback involves using a brain-computer interface that provides an individual with real-time biofeedback about his or her brain activity in the form of sensory feedback. It enables individuals to learn to better control their brain activity, which can be measured in real time using various non-invasive sensors as described above. Although EEG is currently the only modality used by clinical practitioners in that context, it lacks specificity due to its low spatial resolution. Dynamic research into fMRI-neurofeedback has held promise for treating depression, chronic pain and stroke, since it offers the prospect of real-time imagery of the activity in deep brain structures with high spatial resolution. However, the low temporal resolution and high cost of fMRI-neurofeedback has hampered the development of many applications. We believe that the future belongs to hybrid responses that combine multimodal sensors and intend to demonstrate this in the Empenn project.
5 Social and environmental responsibility
- Francesca Galassi: member of the Women in MICCAI (WiM) board - to strengthen and widen the representation of female scientists in the MICCAI community (2021-2023).
- Francesca Galassi and Elise Bannier: members of the Groupe Développement Durable de l'Inria RBA - for the assessment and mitigation of the impact of our research activities on the environment.
- Camille Maumet: co-chair of the women-men equality group at Inria Rennes / IRISA.
- Jérémy Lefort-Besnard: Member of the women-men equality group at Inria Rennes / IRISA
- Elise Bannier: member of the Matching Committee for the Inria mentoring program.
6 Highlights of the year
6.1 Awards
- Seal of Excellence from the European Commission Horizon Europe for the project proposal “Individually Optimized Neurofeedback (IONA)” submitted under the Horizon Europe Marie Sklodowska-Curie Acctions call HORIZON-MSCA-2022-PF-01-01 by Agustina Fragueiro and INRIA Rennes
6.2 Grants
- The PASTRAMI project (Patient-specific statistics for microstructure-augmented connectomics) was funded for the period 2023-2028 in the framework of the AAPG 2023 ANR call. This is a collaborative project with CHU Rennes, LMJL UMR 6629 (Nantes) and HIA Sainte-Anne (Toulon).
- The VICUNA project was funded for the period 2023-2027 in the framework of AAPG 2023 ANR call. This is a JCJC project in collaboration with Mathieu Acher (INSA Rennes). The goal is to explore the variability induced by different configurations in the neuroimaging analytical space.
- The PEPR ShareFAIR project was funded for the period 2023-2028. This is part of the PEPR Santé Nunérique with partners Université Paris-Saclay, Institut Pasteur, Université Paris-Dauphine PSL, Université Claude Bernard Lyon 1, Université de Rennes, INRIA - Université Paris Cité; CHU Rennes, CEA, INSERM and CNRS. The goal is to share reliable protocols to transform datasets into gold standards.
6.3 Habilitation à Diriger les Recherches
- Elise Bannier defended her HDR entitled "Magnetic Resonance for Brain and Spinal Cord imaging and multimodal functional imaging" in January 2023
6.4 Recruitment of permanent staff
- On January 1st, 2023, Fanny Degeilh was appointed as CRCN Inserm at ERL U1228 Empenn. Her research aims to better understand the dynamics and variability of the brain development from infancy to adolescence in typically developing children and following a traumatic brain injury (TBI).
- On January 1st, 2023, Florent Leray was recruited at the Service d'Experimentation de Developpement (SED) as an Inria IR and detached to Empenn to take technical responsibility for the development of MedinriA
7 New software, platforms, open data
7.1 New software
7.1.1 Anima
-
Keywords:
Medical imaging, Neuroimaging, Image processing
-
Scientific Description:
Anima is a set of libraries and tools developed by the team as a common repository of research algorithms. As of now, it contains tools for image registration, statistical analysis (group comparison, patient to group comparison), diffusion imaging (model estimation, tractography, etc.), quantitative MRI processing (quantitative relaxation times estimation, MR simulation), image denoising and filtering, and segmentation tools. All of these tools are based on stable libraries (ITK, VTK), making it simple to maintain.
-
Functional Description:
Anima is a set of libraries and tools in command line mode for processing and analysing medical images.
- URL:
-
Contact:
Julie Coloigner
-
Participants:
Aymeric Stamm, Fang Cao, Florent Leray, Guillaume Pasquier, Laurence Catanese, Olivier Commowick, Renaud Hedouin, René-Paul Debroize
7.1.2 MedINRIA
-
Keywords:
Visualization, DWI, Health, Segmentation, Medical imaging
-
Scientific Description:
MedInria aims at creating an easily extensible platform for the distribution of research algorithms developed at Inria for medical image processing. This project has been funded by the D2T (ADT MedInria-NT) in 2010, renewed in 2012. A fast-track ADT was awarded in 2017 to transition the software core to more recent dependencies and study the possibility of a consortium creation.The Empenn team leads this Inria national project and participates in the development of the common core architecture and features of the software as well as in the development of specific plugins for the team's algorithm.
-
Functional Description:
MedInria is a free software platform dedicated to medical data visualization and processing.
- URL:
-
Contact:
Florent Leray
-
Participants:
Maxime Sermesant, Olivier Commowick, Théodore Papadopoulo
-
Partners:
HARVARD Medical School, IHU - LIRYC, NIH
7.1.3 autoMRI
-
Keywords:
FMRI, MRI, ASL, FASL, SPM, Automation
-
Scientific Description:
This software is highly configurable in order to fit a wide range of needs. Pre-processing includes segmentation of anatomical data, as well as co-registration, spatial normalization and atlas building of all data types. The analysis pipelines perform either within-group analysis or between-group or one subject-versus-group comparison, and produce statistical maps of regions with significant differences. These pipelines can be applied to structural data to exhibit patterns of atrophy or lesions, to ASL (both pulsed or pseudo-continuous sequences) data to detect perfusion abnormalities, to functional data - either BOLD or ASL - to outline brain activations related to block or event-related paradigms. New functionalities have been implemented to facilitate the management and processing of data coming from complex projects.
-
Functional Description:
AutoMRI is based on MATLAB and the SPM12 toolbox and provides complete pipelines to pre-process and analyze various types of images (anatomical, functional, perfusion).
- URL:
-
Contact:
Isabelle Corouge
-
Participants:
Camille Maumet, Elise Bannier, Isabelle Corouge, Pierre Maurel, Quentin Duché, Julie Coloigner
7.1.4 ShanoirUploader
-
Name:
ShanoirUploader (SHAring NeurOImaging Resources Uploader)
-
Keywords:
Webservices, PACS, Medical imaging, Neuroimaging, DICOM, Health, Biology, Java, Shanoir
-
Scientific Description:
ShanoirUploader is a desktop application on base of JavaWebStart (JWS). The application can be downloaded and installed using an internet browser. It interacts with a PACS to query and retrieve the data stored on it. After this ShanoirUploader sends the data to a Shanoir server instance in order to import these data. This application bypasses the situation, that in most of the clinical network infrastructures a server to server connection is complicated to set up between the PACS and a Shanoir server instance.
-
Functional Description:
ShanoirUploader is a Java desktop application that transfers data securely between a PACS and a Shanoir server instance (e.g., within a hospital). It uses either a DICOM query/retrieve connection or a local CD/DVD access to search and access images from a local PACS or the local CD/DVD. After having retrieved the data, the DICOM files are locally anonymized and then uploaded to the Shanoir server. A possible integration of a hash creation application for patient identifiers is provided as well. The primary goals of that application are to enable mass data transfers between different remote server instances and therefore reduce the waiting time of the users, when importing data into Shanoir. Most of the time during import is spent with data transfers.
- URL:
-
Contact:
Michael Kain
-
Participants:
Christian Barillot, Inès Fakhfakh, Justine Guillaumont, Michael Kain, Yao Chi
7.1.5 Shanoir-NG
-
Name:
Shanoir-NG (SHAring iN vivO Imaging Resources - Next Generation)
-
Keyword:
Medical imaging
-
Functional Description:
Shanoir-NG (SHAring iN vivO Imaging Resources - Next Generation) is an open-source web platform designed to share, archive, search and visualize medical imaging data. It provides an user-friendly secure web access and offers an intuitive workflow to facilitate the collecting and retrieving of imaging data from multiple sources. Quality control can be applied on imported data. Mass data can be downloaded in multiple ways, via the web interface and via a Python script.
It supports the following formats: DICOM classic/enhanced (MR, CT, PT, NM), BIDS, processed datasets (NIfTI), Bruker, EEG(BrainVision/EDF).
Shanoir-NG comes along many features such as pseudonymization of data (based on DICOM standard profiles), support for multi-centric clinical studies on subjects. Shanoir-NG offers an ontology-based data organization (OntoNeuroLOG). Among other things, this facilitates the reuse of data and metadata, the integration of processed data and provides traceability trough an evolutionary approach. Shanoir-NG allows researchers, clinicians, PhD students and engineers to undertake quality research projects with an emphasis on remote collaboration. Data user agreements (DUA) can be configured by study to be accepted by each accessing users and access requests can be initated to study administrators.
-
Release Contributions:
Quality control
- URL:
-
Contact:
Michael Kain
-
Participants:
Michael Kain, Anthony Baire, Julien Louis, Jean-cÔme Douteau, Pierre-henri Dauvergne, Arthur Masson, Youenn Merel
7.1.6 LongiSeg4MS
-
Name:
Longitudinal Segmentation For Multiple Sclerosis
-
Keywords:
3D, Brain MRI, Deep learning, Detection
-
Functional Description:
LongiSeg4MS is an automatic new multiple sclerosis (MS) lesion detection tool based on longitudinal data and using deep learning. The system uses FLAIR, T1 or T2 modalities, or a combination of those. The input is 2, 4 or 6 images (2 FLAIR, 2 FLAIR and 2 T1, etc.), a set of modalities for each time point, and outputs a segmentation map describing the location of new MS lesions.
- URL:
-
Authors:
Arthur Masson, Brandon Le Bon, Benoit Combes
-
Contact:
Arthur Masson
-
Partner:
OFSEP
7.1.7 Anima medInria plugins
-
Keywords:
IRM, Medical imaging, Diffusion imaging
-
Functional Description:
Plugins for the medInria software based on the open source software Anima developed in the Visages / Empenn team. These plugins are interfaces between anima and medinria allowing to use Anima functionalities within the clinical user interface provided by medInria. The current functionalities included in the plugins are right now: image registration, denoising, quantitative imaging (relaxometry), and model estimation and visualization from diffusion imaging.
- URL:
-
Contact:
Florent Leray
-
Participants:
Olivier Commowick, Florent Leray, René-Paul Debroize, Guillaume Pasquier
7.2 New platforms
7.2.1 The Neurinfo Platform
Participants: Elise Bannier, Emmanuel Caruyer, Isabelle Corouge, Quentin Duché, Jean-Christophe Ferré, Jean-Yves Gauvrit, Nolwenn Jégou.
Empenn is the founding actor of an experimental research platform which was installed in August 2009 at the University Hospital of Rennes. The University of Rennes 1, Inria, CNRS for the academic side, and the University Hospital of Rennes and the Cancer Institute “Eugene Marquis” for the clinical side, are partners of this neuroinformatics platform called Neurinfo (Neurinfo website). Concerning the Neurinfo Platform, the activity domain is a continuum between methodological and technological research built around specific clinical research projects. On the medical field, the translational research domain mainly concerns medical imaging and more specifically the clinical neurosciences. Among them are multiple sclerosis, epilepsy, neurodegenerative, neurodevelopmental and psychiatric diseases, surgical procedures of brain lesions, neuro-oncology and radiotherapy planning. Beyond these central nervous system applications, the platform is also open to alternative applications. Neurinfo ambitions to support the emergence of research projects based on their level of innovation, their pluri-disciplinarity and their ability to foster collaborations between different actors (public and private research entities, different medical specialties, different scientific profiles). In this context, a research 3T MRI system (Siemens Verio) was acquired in summer 2009 in order to develop the clinical research in the domain of morphological, functional, structural and cellular in-vivo imaging. A new 3T Siemens Prisma MRI scanner was installed at the Neuroinfo platform in February 2018. In 2014, an equipment for simultaneous recording of EEG and MRI images was acquired from Brain Product. In 2015, a mock scanner for experimental set-up was acquired as well as a High Performance Computing environment made of one large computing cluster and a data center that is shared and operated by the Inria center and IRISA (UMR CNRS 6074). The computation cluster (480 cores) and the data center (up to 150 TB) are dedicated to host and process imaging data produced by the Neurinfo platform, but also by other research partners that share their protocols on the Neurinfo neuroinformatics system (currently more than 60 sites). In 2019, an MRI and EEG-compatible fNIRS system was acquired through a co-funding from the INS2I institute of CNRS and FEDER. At the end of 2019, GIS IBISA awarded the Neurinfo platform with a complementary funding that will be dedicated to supplement the current system with additional sensors (from 8x8 optodes to 16x16 optodes). In 2022, the Regional Council of Britanny funding was renewed to provide engineer support for another year to develop and integrate this new imaging system.
7.3 Open data
The HCP multi-pipeline dataset: an opportunity to investigate analytical variability in fMRI data analysis
Participants: Elodie Germani, Pierre Maurel, Camille Maumet.
Results of functional Magnetic Resonance Imaging (fMRI) studies can be impacted by many sources of variability including differences due to: the sampling of the participants, differences in acquisition protocols and material but also due to different analytical choices in the processing of the fMRI data. While variability across participants or across acquisition instruments have been extensively studied in the neuroimaging literature the root causes of analytical variability remain an open question. Here, we share the HCP multi-pipeline dataset, including the resulting statistic maps for 24 typical fMRI pipelines on 1,080 participants of the HCP-Young Adults dataset. We share both individual and group results - for 1,000 groups of 50 participants - over 5 motor contrasts. We hope that this large dataset covering a wide range of analysis conditions will provide new opportunities to study analytical variability in fMRI. 56. This work was done in collaboration with Prof. Elisa Fromont from the LACODAM team.
8 New results
8.1 Basic research
8.1.1 Population imaging
Population imaging is fundamental when it comes to evaluate clinical biomarkers. In this section we summarise our contributions over the last year to this theme. We studied how analytical variability can impact fMRI results and proposed recommendations and neuroinformatics models to describe the data. We also maintained our clinical interest regarding several pathologies by exploring brain function and connectivity. Also, technical recommendations regarding multicentric imaging protocols were proposed.
Uncovering communities of pipelines in the task-fMRI analytical space
Participants: Elodie Germani, Camille Maumet.
Functional magnetic resonance imaging analytical workflows are highly flexible with no definite consensus on how to choose a pipeline. While methods have been developed to explore this analytical space, there is still a lack of understanding of the relationships between the different pipelines. We use community detection algorithms to explore the pipeline space and assess its stability across different contexts. We show that there are subsets of pipelines that give similar results, especially those sharing specific parameters (e.g. number of motion regressors, software packages, etc.), with relative stability across groups of participants. By visualizing the differences between these subsets, we describe the effect of pipeline parameters and derive general relationships in the analytical space. 55. This work was done in collaboration with Prof. Elisa Fromont from the LACODAM team.
On the benefits of self-taught learning for brain decoding
Participants: Elodie Germani, Camille Maumet.
We study the benefits of using a large public neuroimaging database composed of fMRI statistic maps, in a self-taught learning framework, for improving brain decoding on new tasks. First, we leverage the NeuroVault database to train, on a selection of relevant statistic maps, a convolutional autoencoder to reconstruct these maps. Then, we use this trained encoder to initialize a supervised convolutional neural network to classify tasks or cognitive processes of unseen statistic maps from large collections of the NeuroVault database. We show that such a self-taught learning process always improves the performance of the classifiers but the magnitude of the benefits strongly depends on the number of samples available both for pre-training and finetuning the models and on the complexity of the targeted downstream task. The pre-trained model improves the classification performance and displays more generalizable features, less sensitive to individual differences 28. This work was done in collaboration with Prof. Elisa Fromont from the LACODAM team.
Representation learning for more reproducible fMRI data analyses
Participants: Elodie Germani, Camille Maumet.
Analysing functional brain MRI (fMRI) data requires a sequence of complex and specific steps, leading to what we call an analysis pipeline. Recently, numbers of studies have shown that the choice of pipeline has an impact on the final activation maps, raising questions about the validity of published results and the possibility of reusing existing data. In this context, we propose the use of generative models to convert a contrast map obtained from a source pipeline into its version obtained from a different target pipeline. We assume that different pipelines can be modelled as different styles. By analysing data from the Human Connectome Project (1000+ participants) with different conventional pipelines using SPM and FSL software, we formed pairs of contrast maps corresponding to the same raw image analysed with two different pipelines. We then trained a conditional adversarial generative network (cGAN), based on the architecture of the Pix2Pix model used in style transfer and adapted to 3D, to learn how to convert a map from a source pipeline to its version in a target pipeline. Our initial experiments show that conversion performance, evaluated using comparison metrics (correlation and voxel-to-voxel difference) between generated maps, source maps and target maps, varies according to the pairs of pipelines evaluated and the direction of the conversion. 66 This work was done in collaboration with Prof. Elisa Fromont from the LACODAM team.
NARPS open pipeline
Participants: Jeremy Lefort-Besnard, Boris Clenet, Elodie Germani, Camille Maumet.
Scientific pipelines are at the heart of modern experimental sciences. But practitioners face a highly complex pipeline landscape – different tools, algorithms, parameters – in which different pipelines can lead to contradictory research findings. Until recently, this analytic variability – i.e. the variability induced by different pipelines on the results – has typically been ignored, effectively considering that it was negligible compared to other sources of variability (e.g. as induced by participants, test-retest, measurement error, etc.). But in 2020, a landmark paper in Nature challenged this status-quo. In this paper, 70 teams were given the same dataset and tasked to answer the same yes/no research questions. Each team chose a different pipeline and – what is more worrying – those differences in pipelines also led to contradictory findings. The goal of the NARPS Open Pipelines project is thus to create a codebase reproducing the 70 pipelines of the NARPS project and share this as an open resource for the community. This article was selected due to its provision of a comprehensive set of pipelines genuinely employed within the scientific community, with nearly 200 scientists contributing to this collaborative work. Special attention has been devoted to obtaining detailed information for each of the 70 pipelines. 69
Medial positioning of the hippocampus and hippocampal fissure volume in Developmental Topographical Disorientation
Participants: Agustina Fragueiro, Claire Cury.
Developmental Topographical Disorientation (DTD) refers to the lifelong inability to orient by means of cognitive maps in familiar surroundings despite otherwise well-preserved general cognitive functions, and the absence of any acquired brain injury or neurological condition. While reduced functional connectivity between the hippocampus and other brain regions has been reported in DTD individuals, no structural differences in grey matter tissue for the whole brain neither for the hippocampus were detected. Considering that the human hippocampus is the main structure associated with cognitive map-based navigation, we investigated differences in morphological and morphometric hippocampal features between individuals affected by DTD (N=20) and healthy controls (N=238). Specifically, we focused on a developmental anomaly of the hippocampus that is characterized by the incomplete infolding of hippocampal subfields during foetal development, giving the hippocampus a more round or pyramidal shape, called Incomplete Hippocampal Inversion (IHI). We rated IHI according to standard criteria and extracted hippocampal subfield volumes after FreeSurfer’s automatic segmentation. We observed similar IHI prevalence in the group of individuals with DTD with respect to the control population. Neither differences in whole hippocampal nor major hippocampal subfield volumes have been observed between groups. However, when assessing the IHI independent criteria, we observed that the hippocampus in the DTD group is more medially positioned comparing to the control group. In addition, we observed bigger hippocampal fissure volume for the DTD comparing to the control group. Both of these findings were stronger for the right hippocampus comparing to the left. Our results provide new insights regarding the hippocampal morphology of individuals affected by DTD, highlighting the role of structural anomalies during early prenatal development in line with the developmental nature of the spatial disorientation deficit. [Papier: Medial positioning of the hippocampus and hippocampal fissure volume in Developmental Topographical Disorientation, HIPPOCAMPUS] 26 [Presentation in conference: Shift in hippocampal medial position and increased fissure volumes in individuals affected by Developmental Topographical Disorientation, 8th Scientific Meeting of the Federation of European Societies of Neuropsychology (FESN)] 63
Incomplete Hippocampal Inversion and Hippocampal Subfield Volumes: Implementation and Inter-Reliability of Automatic Segmentation
Participants: Agustina Fragueiro, Claire Cury.
The incomplete hippocampal inversion (IHI) is an atypical anatomical pattern of the hippocampus. However, the hippocampus is not a homogeneous structure, as it consists of segregated subfields with specific characteristics. While IHI is not related to whole hippocampal volume, higher IHI scores have been associated to smaller CA1 in aging. Although the segmentation of hippocampal subfields is challenging due to their small size, there are algorithms allowing their automatic segmentation. By using a Human Connectome Project dataset of healthy young adults, we first tested the inter-reliability of two methods for automatic segmentation of hippocampal subfields, and secondly, we explored the relationship between IHI and subfield volumes. Results evidenced strong correlations between volumes obtained thorough both segmentation methods. Furthermore, higher IHI scores were associated to bigger subiculum and smaller CA1 volumes. Here, we provided new insights regarding IHI subfields volumetry, and we offer support for automatic segmentation inter-method reliability. [Papier: Incomplete Hippocampal Inversion and Hippocampal Subfield Volumes: Implementation and Inter-Reliability of Automatic Segmentation, 2023 IEEE 20th International Symposium on Biomedical Imaging (ISBI)] 54 [Poster: Incomplete Hippocampal Inversion and Hippocampal Subfield Volumes: Implementation and Inter-Reliability of Automatic Segmentation, 20th IEEE-International Symposium on Biomedical Imaging (ISBI)] 43
Temporo-basal sulcal connections: a manual annotation protocol and an investigation of sexual dimorphism and heritability
Participants: Claire Cury.
This study about the temporo-basal region of the human brain - composed of the collateral, the occipito- temporal, and the rhinal sulci - has bee published in Brain Structure and Function. We manually rated (using a novel protocol) the connections between rhinal/collateral (RS-CS), collateral/occipito-temporal (CS-OTS) and rhinal/occipito-temporal (RS-OTS) sulci, using the MRI of nearly 3,400 individuals including around 1000 twins. We reported both the associations between sulcal polymorphisms as well with a wide range of demographics (e.g. age, sex, handedness). Finally, we also estimated the heritability, and the genetic correlation between sulcal connections. We reported the frequency of the sulcal connections in the general population, which were hemisphere dependent. We found a sexual dimorphism of the connections, especially marked in the right hemisphere, with a CS-OTS connection more frequent in females (approximately 35-40% versus 20-25% in males) and an RS-CS connection more common in males (approximately 40-45% versus 25-30% in females). We confirmed associations between sulcal connections and characteristics of incomplete hippocampal inversion (IHI). We estimated the broad sense heritability to be 0.28-0.45 for RS-CS and CS-OTS connections, with hints of dominant contribution for the RS-CS connection. The connections appeared to share some of their genetic causing factors as indicated by strong genetic correlations. Heritability appeared much smaller for the (rarer) RS-OTS connection. 35
Reproducibility of motor task-based fNIRS and comparison with functionalMRI in healthy adults
Participants: Nolwenn Jegou, Elise Bannier, Emmanuel Caruyer, Isabelle Corouge.
NIRS is an optical imaging technique that estimates cerebral hemodynamic variations and thus indirectly reflects brain activity 25. In 2023, we focused on automating NIRS data processing. Our data analysis pipeline has been refined at individual level, and group-level analysis has been implemented with code migration to gitlab. We took part in the international "FRESH challenge: fNIRS REproducibility Study Hub" () to study the variability and impact of the different analysis techniques used used by the NIRS community, the results of which we are awaiting. Besides, we studied the ability and reproducibility of fNIRS to map the cortical motor areas. Simultaneously acquired fMRI was used as a reference and functional maps of both modalities were obtained from GLM analysis. NIRS results shows satisfactory reproducibility but partial agreement with fMRI. This work led to a publication in a national conference 68. Last, we welcomed Demian Vera, PhD student at Tandil University, Argentina, for a 3 month stay, to collaborate on the development of multi-layer models.
Successful reproduction of a large EEG study across software packages
Participants: Camille Maumet, Nina Forde.
As an active field of research and with the development of state-of-the-art algorithms to analyze EEG datasets, the parametrization of Electroencephalography (EEG) analysis workflows has become increasingly flexible and complex, with a great variety of methodological options and tools to be selected at each step. This high analytical flexibility can be problematic as it can yield to variability in research outcomes. Therefore, growing attention has been recently paid to understand the potential impact of different methodological decisions on the reproducibility of results. In this paper, we aim to examine how sensitive the results of EEG analyses are to variations in preprocessing with different software tools. We reanalyzed the shared EEG data (N=500) from (Williams et al. 2021) using three of the most commonly used EEG software tools: EEGLAB, Brainstorm and FieldTrip. After reproducing the same original preprocessing workflow in each software, the resulting evoked-related potentials (ERPs) were qualitatively and quantitatively compared in order to examine the degree of consistency/discrepancy between softwares. Our findings show a good degree of convergence in terms of the general profile of ERP waveforms, peak latencies and effect size estimates related to specific signal features. However, considerable variability was also observed in the magnitude of the absolute voltage observed with each software package as reflected by the similarity values and observed statistical differences at particular channels and time instants. In conclusion, we believe that this study provides valuable clues to better understand the impact of the software tool on the analysis of EEG results. This work was led by Aya Kabbara in a project co-supervised by Mahmoud Hasssan and Camille Maumet 32.
8.1.2 Detection and learning
Can we accurately assess disease activity using automated methods in large real-life MRI databases? Insights from the OFSEP HD database
Participants: Arthur Masson, Benoit Combès, Alice Dufey, Anne Kerbrat.
Large real-life databases (DB) of MS patients usually consist of clinical data, including limited imaging metrics. While the systematic collection of MRI is rare, the possibility of re-analyzing images to extract a wide range of metrics is now possible through the use of AI based methods. The presence of new lesions on longitudinal MRIs for exemple is used to assess the effectiveness of treatments in real-life studies. However, the automated tools that currently identify these new lesions are designed as an aid for radiologists, potentially generating false positives. The possibility of transferring these methods to analyze DB without supervision should be assessed. In this work 70, our objectives were to compare the performance of an automated method to classify MS patients as “active” or “inactive” based on new lesions on FLAIR images in a large real-life multicentric DB with respect to the data provided in the clinical DB. For that purpose, we included 1412 pairs of brain MRI scans from 868 MS patients with both FLAIR images available in the French OFSEP HD cohort imaging DB at 2 time points, and the radiological comparison captured in the clinical DB. An automated tool based on a fully convolutional neural network (trained on 159 patients) was used to detect new lesions between the corresponding longitudinal FLAIR images. Then, 160 pairs of brain MRI scans for which the automated method output and the corresponding clinical DB comparison disagree were randomly selected and their MRI were reviewed by 2 experts to constitute a ground truth. Differences in sensitivity, specificity and accuracy between the automated method and the clinical DB were assessed. Overall, 222 out of 1412 (16%) intervals were considered active from the clinical DB, compared to 467 (35%) from the automated method. Over the 160 cases of disagreement included in the ground truth, the automated method correctly classified patients in 66% of the cases and the clinical DB in 34%. More specifically, the automated method was more sensitive than the clinical DB (p<0.001), but the clinical DB was more specific (p<0.001). Under simplified assumptions, we extrapolate from these results a sensitivity, specificity and accuracy of about 0.95, 0.99 and 0.92 for the clinical DB and 0.99, 0.69 and 0.96 for the automated method. In conclusion, the automated analysis of images collected in large real-life databases allows to correctly classify MS patients as active or inactive in a large majority of cases, and offers the possibility to extract other metrics such as lesion number or volume to analyze the efficacy of treatments in real-life.
Expert variability and deep learning performance in spinal cord lesion segmentation for multiple sclerosis patients.
Participants: Ricky Walsh, Cédric Meurée, Anne Kerbrat, Arthur Masson, Burhan Rashid Hussein, Malo Gaubert, Francesca Galassi, Benoît Combès.
Multiple sclerosis (MS) patients often present with lesions in spinal cord magnetic resonance (MR) volumes. However, accurately detecting these lesions is challenging and prone to inter-and intra-rater variability. Deep learning-based methods have the potential to aid clinicians in detecting and segmenting MS lesions, but can also be affected by rater variability. In this work 50, we assessed the inter-and intra-rater variability in manual segmentation of spinal cord lesions, and evaluated raters and a state-of-the-art nnU-Net model against a ground truth (GT) segmentation of a senior expert. Four experts segmented twelve spinal cord MR volumes from six patients twice, at a time distance of two weeks. Considerable inter-and intra-rater variability were observed, with the total number of detected lesions ranging from 28 to 60, depending on the rater. An example of lesion is depicted in Figure 3. Moreover, the segmented volumes of individual lesions varied substantially between raters. All raters and the model achieved high precision when evaluated against the senior expert GT, but sensitivity was notably lower. These results motivate the need for more sensitive automated methods to aid clinicians in lesion detection, and suggest that consideration should be given to inter-rater variability when training and evaluating automated methods.
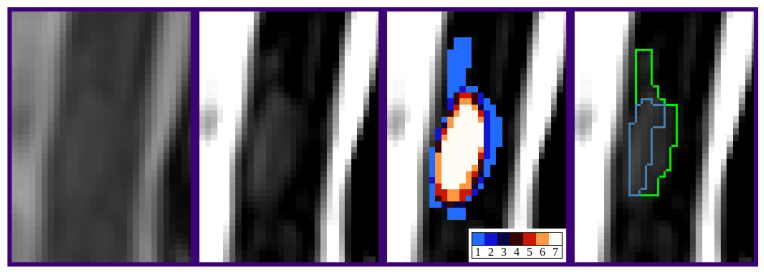
Illustration of the low contrast of MS spinal cord lesions and of the resulting discrepancy between experts annotations
Example of a lesion in the dataset (at vertebra T6). The bright vertical bands are cerebrospinal fluid surrounding the spinal cord. Left: subsection of sagittal T2-w volume. Centre Left: contrast adjusted to improve lesion visibility. Centre Right: voxel-wise agreement between the eight annotations (four raters with two sessions each). One of the raters did not detect this lesion in one of the sessions. Right: predicted segmentation of nnU-Net model (blue) and adjudicated segmentation by senior expert (green).
A study on loss functions and decision thresholds for the segmentation of multiple sclerosis lesions on spinal cord MRI.
Participants: Burhan Rashid Hussein, Cédric Meurée, Malo Gaubert, Arthur Masson, Anne Kerbrat, Benoît Combès, Francesca Galassi.
Multiple sclerosis (MS) patients often present hyper-intense T2-w lesions in the spinal cord. The severe imbalance between background and lesion classes poses a major challenge to Deep Learning segmentation approaches, requiring for ad hoc strategies. Careful selection of the loss function and adjustment of the conventional 0.5-thresholding may help mitigating this issue. In this work 47, 67, we showed the performance advantages of loss functions based on the Tversky Index and the benefits of threshold tuning over more standard settings and the state-of-the-art model for MS lesion segmentation on spinal cord MRI (Figure 4).
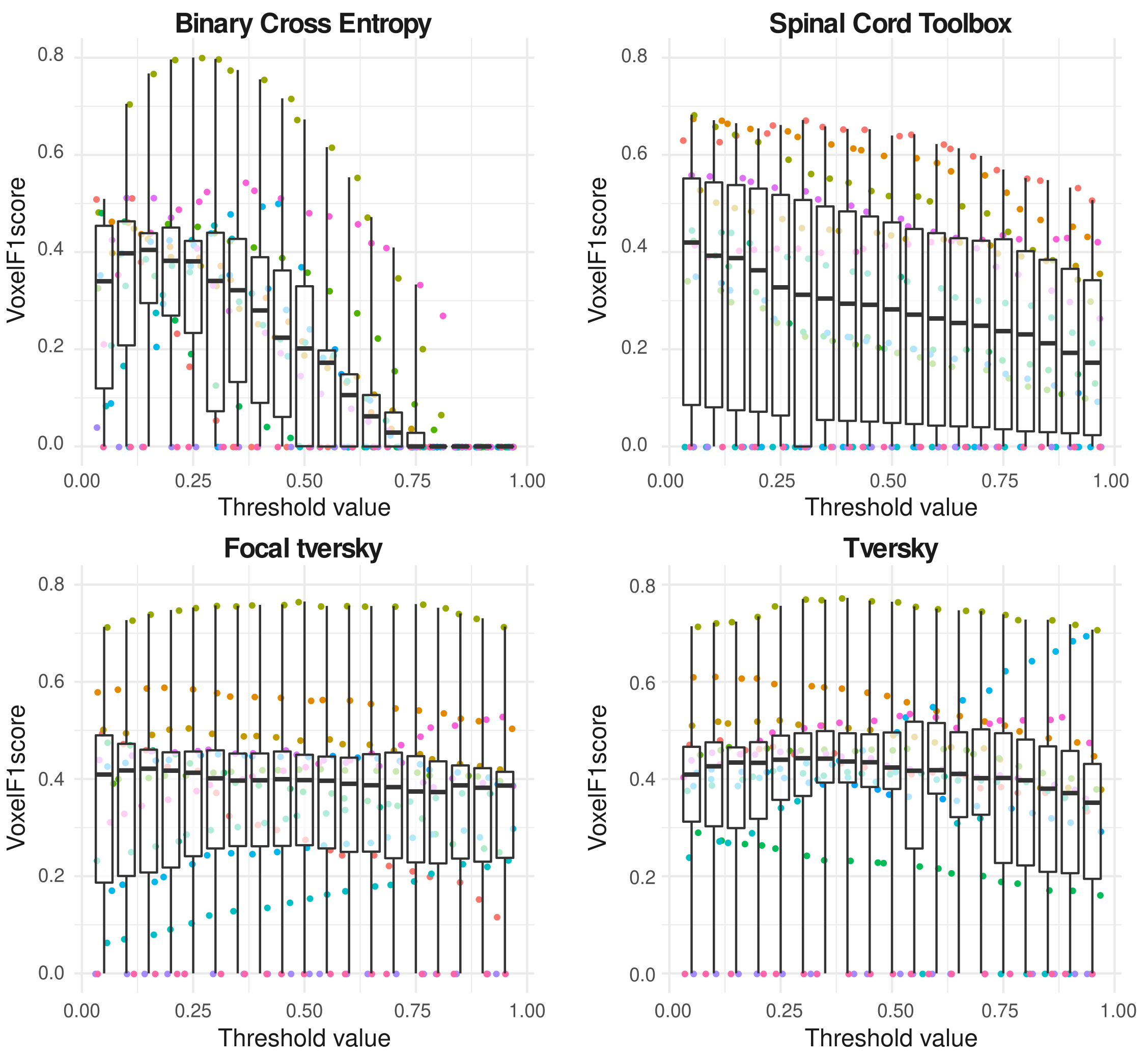
Boxplots illustrating the stablility of the F1 score when changing the softmax threshold values for the Tversky-based losses.
Boxplots of voxel-wise F1 score for BCE, SCT, Focal Tversky and Tversky losses at each decision threshold value. One point corresponds to one subject.
An automatic solution for chronic stroke lesion segmentation in brain MRI.
Participants: Lounès Meddahi, Arthur Masson, Elise Bannier, Stéphanie Leplaideur, Francesca Galassi.
Stroke is one of the leading causes of long-term adult disability worldwide. Post-stroke rehabilitation is crucial for long-term patient recovery. Determining the volume and location of lesions caused by stroke is essential to guide treatment and provide effective rehabilitation. Currently, the gold standard for chronic stroke lesion segmentation is manual tracing, a procedure that requires knowledge, is time consuming and prone to inter‐rater variability. Automatic segmentation algorithms have the potential to overcome these limitations. While a large number of solutions have been proposed for the automatic segmentation of lesions in the acute phase, tools for chronic stroke lesion segmentation are underdeveloped. Methods for acute stroke are not readily applicable to chronic stroke due to the different characteristics of the imaging protocol and of the lesion itself. We have developed a pipeline that outperforms state-of-the-art methods and demonstrates the advantages of incorporating a second modality in terms of segmentation accuracy. Results have been submitted to the World Congress for Neurorehabilitation 2024, and a research paper is currently under preparation.
Improving portability of bimodal neurofeedback: predicting NF-fMRI scores from EEG signals
Participants: Caroline Pinte, Claire Cury, Pierre Maurel.
Neurofeedback (NF) is a method that allows a subject to learn how to regulate his or her brain activity. During a training session, the subject will see real-time feedback from his or her brain activity and can use it to perform a task such as motor imagery. To measure brain activity, simultaneous acquisitions with EEG and fMRI provide more effective NF training due to their temporal and spatial complementarity 84. However, using MRI is expensive and can be draining for the subject. Therefore, we would like to reduce its use and thus improve the portability of EEG-fMRI neurofeedback. Following the work of 83, we propose a method based on a convolutional neural network (CNN). This method consists in learning a model from simultaneous EEG-fMRI acquisitions to predict NF-fMRI scores with EEG signals alone 80, 79.
Pilot study: eye-tracking and skin conductance to monitor task engagement during bimodal neurofeedback
Participants: Agustina Fragueiro, Rene-Paul Debroize, Elise Bannier, Claire Cury.
NF consists in providing real-time neural activation feedback to self-regulate brain activity. It is a promising brain rehabilitation technique as it can trigger brain plasticity. EEG based NF is widely used, but while it has excellent temporal resolution, it has limited spatial resolution. On the other hand, NF based on fMRI, offers a better spatial resolution, but has slow dynamics. Current studies are showing the high potential of combining EEG and fMRI in bimodal NF. However, a significant percentage of people undergoing NF training, fails. Motivational and attentional factors have been identified as predictors of NF learning, as poor performances can lead to disengagement with the task and a label of “non-responder”. We proposed to use ET and SC to monitor participants’ task engagement during bimodal NF sessions. We aimed at: 1) synchronizing all devices (ET, SC, EEG, fMRI), 2) identifying ET and SC features to detect changes in task engagement. In this pilot study, we synchronized all devices an tested the set up in 2 participants (s1 and s2). We acquired structural and functional data in a 3T scanner, while simultaneously recording EEG activity using a 64-channel MR-compatible cap. A MR-compatible eye-tracking camera system was used to register ocular movements from the dominant eye at 60Hz. View Point was used to compute the number of saccades and fixation durations (saccades velocity threshold [normalized gaze position change/ms]=0.20). A MR-compatible Brain Vision Galvanic Skin Response set was used to acquire electrodermal activity from the index and middle fingers. Quantity of SC responses (SCR) and their amplitude were obtained through a Continuous Decomposition Analysis using Ledalab. Cognitive workload was stimulated using a color-word interference Stroop task ( 4min) followed by a two-minute rest in which a video encouraging heart coherence was presented. We simultaneously recorded and time-stamped fMRI, EEG, ET and SC signals, while the participant was completing the cognitive task. The rate of SCR (quantity/secs, relative difference: s1=0.23, s2=0.69) and their amplitude (Z-scores difference: s1=0.56, s2=0.36) were higher for the task compared to the rest block in both participants. The rate of saccades (quantity/secs) was higher for the task compared to the rest (relative difference: s1=0.81, s2=0.74), while fixations duration was longer for the rest block (seconds difference: s1=-3.2, s2=-0.12). EEG and fMRI signals were simultaneously recorded as proof of the set-up feasibility for subsequent analysis during NF trainings. While the Stroop task allow us to observe differences in workload-related arousal, in the forthcoming acquisitions (N=20) we are including a task to monitor attentional focus. ET and SC differences between conditions, will be used to identify different task-engagement states during a NF session, so NF targets may be adapted to keep the participant focused. NF procedures may be personalized. [Presentation in conference: Pilot study: eye-tracking and skin conductance to monitor task engagement during bimodal neurofeedback, 20th IEEE-International Symposium on Biomedical Imaging (ISBI)] 62
Interpretable automatic detection of incomplete hippocampal inversions using anatomical criteria
Participants: Claire Cury.
Incomplete Hippocampal Inversion (IHI) is an atypical anatomical pattern of the hippocampus that has been associated with several brain disorders (epilepsy, schizophrenia). IHI can be visually detected on coronal T1 weighted MRI images. IHI can be absent, partial or complete (no IHI, partial IHI, IHI). However, visual evaluation can be long and tedious, justifying the need for an automatic method. In this paper, we propose, to the best of our knowledge, the first automatic IHI detection method from T1-weighted MRI. The originality of our approach is that, instead of directly detecting IHI, we propose to predict several anatomical criteria, which each characterize a particular anatomical feature of IHI, and that can ultimately be combined for IHI detection. Such individual criteria have the advantage of providing interpretable anatomical information regarding the morphological aspect of a given hippocampus. We relied on a large population of 2,008 participants from the IMAGEN study. The approach is general and can be used with different machine learning models. We explored two different backbone models for the prediction: a linear method (ridge regression) and a deep convolutional neural network. We demonstrated that the interpretable, anatomical based prediction was at least as good as when predicting directly the presence of IHI, while providing interpretable information to the clinician or neuroscientist. This approach may be applied to other diagnostic tasks which can be characterized radiologically by several anatomical features. 46
Systematic review and evaluation of meta-analysis methods for neuroimaging same-date meta-analysis
Participants: Jeremy Lefort-Besnard, Camille Maumet.
Researchers using task-fMRI data have access to a wide range of analysis tools to model brain activity. This diversity of analytical approaches has been shown to have substantial effects on neuroimaging results. Combined with selective reporting, this analytical flexibility can lead to an inflated rate of false positives and contributes to the irreproducibility of neuroimaging findings. Multiverse analyses are a way to systematically explore and integrate pipeline variation on a given dataset. We focused on the setting where multiple statistic maps are produced as an output of a set of analyses originating from a single datset. Meta-analysis is a natural approach to extract consensus inferences from these maps, yet the traditional assumption of independence amongst input datasets does not hold. We thus considered a suite of methods to conduct meta-analysis in the multiverse setting, accounting for inter-pipeline dependence among the results. The validity of these methods were assessed in a set of simulations and evaluated on a real world dataset from "NARPS", a multiverse analysis with 70 different statistic maps originating from the same data, and a multiverse analysis originating form the same HCP data. Our findings demonstrated the validity of our proposed same-data meta-analysis (SDMA) models under inter-pipeline dependence, and provided an array of options for the analysis multiverse data. This work was done in collaboration with Thomas Nichols from Oxford University 51
8.1.3 Quantitative imaging
Quantitative imaging methods can provide access to imaging metrics which can help characterize tissue integrity or neural activity. These methods can be used to assess tissue impairment, lesion severity and follow disease evolution. We investigated the potential of T1 relaxometry as well as diffusion and functional imaging methods.
Simultaneous brain and cervical spinal cord MP2RAGE for T1 measurement: robustness and sensitivity for tissue modification assessment in multiple sclerosis in a multicenter context
Participants: Malo Gaubert, Benoit Combès, Jean-Christophe Ferré, Alice Dufey, Anne Kerbrat, Elise Bannier.
Recent optimisations of T1 quantification through magnetization‐prepared two rapid acquisition gradient echoes (MP2RAGE) allow to perform both brain and cervical spinal cord acquisitions simultaneously with good trade-off between acquisition time, robustness and accuracy. This sequence is of particular interest to investigate tissue microstructural modifications in pathologies such as multiple sclerosis (MS). In order to spread out the use of the MP2RAGE sequence, we evaluated 45 the reproducibility and variability in two different centres. Six healthy controls (HC) were scanned 3 times each (separated sessions), in two different centres both equipped with 3T Siemens scanners. Additionally 26 HC (centre 1/2: 20/6) were scanned one time. The same acquisition protocol was performed in both centres and included MP2RAGE and B1 map acquisitions covering both brain and cervical spinal cord (cSC). After B1 correction, mean T1 values were extracted in different regions including brain white matter (bWM), deep grey matter (dGM) and cortical grey matter (cGM; all computed using CAT12) and all cSC segments (computed using the SCT toolbox). We evaluated the variability between centres and subjects using linear mixed-effects models with subject as random effect and centre as fixed effect. The coefficients of variation (CV) and the intraclass correlations (ICC) of between-session and between-participant variabilities were computed. In order to interpret these results with respect to potential application in MS pathology, we also reported exploratory analyses based on the extraction of T1 values in the same regions for 5 MS patients (centre 1/2: 3/2, same acquisition protocol and image processing) without cSC lesions. For the whole dataset collected in HC, the mean (and standard deviation) T1 values in the brain were 1281.5 (28.8), 1176.5 (20) and 823.9 (21.1) ms for cGM,dGM, and bWM, respectively and were ranging from 921 (22.6) to 954 (30.5) ms over the 7 cSC segments. For the brain, we observed evidence of centre differences for the three regions (all p<.01). Nevertheless, the estimated differences between centres were low, ranging from 4.71 (bWM) to 25.31 (dGM) ms (ie. 0.57 to 1.98% of the mean). Between-participant CV were 2.1, 1.7 and 1.8%, and between-session CV were 0.2, 2.2 and 0.5% for bWM cGM and dGM, respectively. Between-session ICC were .01, .61 and .06 for the same regions. For the SC, we observed evidence of centre differences for all vertebrae (all p<.05), except C4, C5 and C7 (p=.149, .163, .062, resp.). The estimated mean differences were also low, ranging from 9.6 (C5) to 20.2 (C1) ms (ie. 1.03 to 2.15%). To simplify the results, T1 values from C3 to C5 levels were averaged. In this region, between-participant and between-session CV were 1.5 and 1.6%, while between-session ICC was .53. MS patients showed a mean T1 value increase ranging from 19.5 (cGM) to 44.2 (dGM) ms for the brain, and from 14 (C7) to 122.7 (C3) ms for the cSC compared to the mean in all HC. To sum up, even if differences exist between the two centres, the variability is low, especially for bWM (0.57%) and central cSC segments (1.03%). Moreover, the T1 variability is primarily explained by between-participant variability for the brain and by both session- and participant-variabilities for cSC. The differences between scanners were found to be less important than the differences observed between HC and MS patients with no cSC lesions. Overall, the simultaneous brain and cervical spinal cord acquisition is robust to multicentre. This sequence has an interesting potential for further applications in multicenter MS studies to assess regional tissue impairment.
A Riemannian framework for incorporating white matter bundle priors in ODF-based tractography algorithms.
Participants: Thomas Durantel, Julie Coloigner.
Diffusion magnetic resonance imaging (dMRI) tractography is a powerful approach to study brain structural connectivity. However, its reliability in a clinical context is still highly debated. Recent studies have shown that most classical algorithms achieve to recover the majority of existing true bundles. However, the generated tractograms contain many invalid bundles. This is due to the crossing fibers and bottleneck problems which increase the number of false positive fibers. In this work, we proposed to overpass this limitation with a novel method to guide the algorithms in those challenging regions with prior knowledge of the anatomy. In this work 52, we developed a method to create a combination of anatomical prior applicable to any orientation distribution function (ODF)-based tractography algorithms. The proposed method captures the track orientation distribution (TOD) from an atlas of segmented fiber bundles and incorporates it during the tracking process, using a Riemannian framework. We tested the prior incorporation method on two ODF-based state-of-the-art algorithms, iFOD2 and Trekker PTT, on the diffusion-simulated connectivity (DiSCo) dataset and on the Human Connectome Project (HCP) data. We also compared our method with two bundles priors generated by the bundle specific tractography (BST) method. We showed that our method improves the overall spatial coverage and connectivity of a tractogram on the two datasets, especially in crossing fiber regions. Moreover, the fiber reconstruction may be improved on clinical data, informed by prior extracted on high quality data, and therefore could help in the study of brain anatomy and function.
Brain BOLD and NIRS response to hyperoxic challenge in sickle cell disease and chronic anemias.
Participants: Julie Coloigner.
Congenital anemias, including sickle cell anemia and thalassemia, are associated with cerebral tissue hypoxia and heightened stroke risks. Recent works in sickle cell disease mouse models have suggested that hyperoxia respiratory challenges can identify regions of the brain having chronic tissue hypoxia. Therefore, this work 41 investigated differences in hyperoxic response and regional cerebral oxygenation between anemic and healthy subjects. A cohort of 38 sickle cell disease subjects (age 22 8 years, female 39%), 25 nonsickle anemic subjects (age 25 11 years, female 52%), and 31 healthy controls (age 25 10 years, female 68%) were examined. A hyperoxic gas challenge was performed with concurrent acquisition of blood oxygen level-dependent (BOLD) MRI and near-infrared spectroscopy (NIRS). In addition to hyperoxia-induced changes in BOLD and NIRS, global measurements of cerebral blood flow, oxygen delivery, and cerebral metabolic rate of oxygen were obtained and compared between the three groups. Regional BOLD changes were not able to identify brain regions of flow limitation in chronically anemic patients. Higher blood oxygen content and tissue oxygenation were observed during hyperoxia gas challenge. Both control and anemic groups demonstrated lower blood flow, oxygen delivery, and metabolic rate compared to baseline, but the oxygen metabolism in anemic subjects were abnormally low during hyperoxic exposure. These results indicated that hyperoxic respiratory challenge could not be used to identify chronically ischemic brain. Furthermore, the low hyperoxia-induced metabolic rate suggested potential negative effects of prolonged oxygen therapy and required further studies to evaluate the risk for hyperoxia-induced oxygen toxicity and cerebral dysfunction.
Study of the effect of noise on the estimation of microscopic anisotropy in diffusion MRI
Participants: Constance Bocquillon, Élise Bannier, Isabelle Corouge, Emmanuel Caruyer.
B-tensor acquisitions in diffusion MRI enables the measurement of additional microstructure parameters such as microscopic anisotropy (μFA), in order to better describe the heterogeneity of diffusion properties in each voxel, compared to more conventional measurements using the diffusion tensor. In this work, we wanted to evaluate the effect of the noise and the selected acquisition scheme on the estimation of such parameters. We generated diffusion tensors distributions (DTD) with a known target μFA, then we simulated the signal for different acquisition schemes including b-tensors (bmax = 6000s/mm2) in order to study the impact of the choice of b-tensors on the estimation of the μFA, then corrupted by Rician noise (SNR between 30 and 100, calculated on the non-diffusion-weighted image). The mean diffusion tensors and the covariance of DTD are estimated from the signal and can then be used to compute the μFA. A diffusion sequence was developed to play these arbitrary gradients and an acquisition featuring b-tensors on a healthy volunteer was carried out on the 3T Magnetom Prisma MRI (Siemens Healthineers, Erlangen) (VE11C) of the Neurinfo platform (CPP OSS-IRM). At low SNR (SNR = 30), there is a significant bias in the estimation of μFA; this bias decreases when the SNR increases (SNR = 100 and without noise). For low values of μFA, there is a strong overestimation of the value, while high values are slightly underestimated. Even if the trends remain the same, the acquisition pattern has an impact on the accuracy of the measurement. The images on healthy subjects show a homogeneous μFA in the white matter, unlike the FA whose value decreases in the crossing regions. We highlighted the errors in the estimation of μFA; this suggests that the bias introduced by Rician noise must be taken into account when estimating the parameters of the tensor distribution 60.
Tractography passes the test: Results from the diffusion-simulated connectivity (disco) challenge
Participants: Emmanuel Caruyer.
Estimating structural connectivity from diffusion-weighted magnetic resonance imaging is a challenging task, partly due to the presence of false-positive connections and the misestimation of connection weights. Building on previous efforts, the MICCAI-CDMRI Diffusion-Simulated Connectivity (DiSCo) challenge was carried out to evaluate state-of-the-art connectivity methods using novel large-scale numerical phantoms. The diffusion signal for the phantoms was obtained from Monte Carlo simulations. The results of the challenge suggest that methods selected by the 14 teams participating in the challenge can provide high correlations between estimated and ground-truth connectivity weights, in complex numerical environments. Additionally, the methods used by the participating teams were able to accurately identify the binary connectivity of the numerical dataset. However, specific false positive and false negative connections were consistently estimated across all methods. Although the challenge dataset doesn’t capture the complexity of a real brain, it provided unique data with known macrostructure and microstructure ground-truth properties to facilitate the development of connectivity estimation methods 29.
8.2 Translational research
Our goal is also to provide new computational solutions for our target clinical applications (Alzheimer's disease, psychiatry, neurology or public health issues), allowing a more appropriate representation of the data for image analysis and detection of specific biomarkers. In this section, we present the contributions of the last year in the clinical applications of behavior and neuro-inflammation.
8.2.1 Behavior
Structural Brain Connectivity and Treatment Improvement in Mood Disorder
Participants: Sebastien Dam, Pierre Maurel, Julie Coloigner.
Mood depressive disorder (MDD) affects the emotional state as expressed as a persistent feeling of sadness and loss of interest. Antidepressant medications are first line treatment for depression. In this work, we propose to identify patterns of MDD via a cross-sectional cohort, with the assumption that alterations in brain connectivity may constitute a sensitive biomarker of depression and more specifically of poor outcome of a mood depressive episode. Using diffusion magnetic resonance imaging, we performed structural connectivity analyses using graph theory approach on a cohort of depressed patients and healthy volunteers. In order to study illness improvement, the MDD patients went through two clinical interviews at baseline and at 6 months follow-up, thus allowing us to classify them into “responders” (R) or “non-responders” (NR) based on the Clinical Global Impression-Improvement score. First, the threshold-free network-based statistics (TFNBS) was conducted to highlight the graph modifications between the different groups. Second, we performed a statistical analysis of topological metrics tests between depressed patients versus healthy controls and between R versus NR.
Connectivity patterns of the core resting-state networks associated with apathy in late-life depression.
Participants: Jean-Charles Roy, Julie Coloigner, Gabriel Robert.
Apathy is associated with reduced antidepressant response and dementia in late-life depression (LLD). However, the functional cerebral basis of apathy is understudied in LLD. In this work 38, we investigated the functional connectivity of 5 resting-state networks (RSN) hypothesized to underlie apathy in LLD. Resting-state functional MRI data were collected from individuals with LLD who did not have dementia as well as healthy older adults between October 2019 and April 2022. Apathy was evaluated using the diagnostic criteria for apathy (DCA), the Apathy Evaluation Scale (AES) and the Apathy Motivation Index (AMI). Subnetworks whose connectivity was significantly associated with each apathy measure were identified via the threshold-free network-based statistics. Regions that were consistently associated with apathy across the measures were reported as robust findings. Our sample included 39 individuals with LLD who did not have dementia and 26 healthy older adults. Compared with healthy controls, individuals with LLD had an altered intraRSN and inter-RNS connectivity in the default mode, the cingulo-opercular and the frontoparietal networks. All 3 apathy measurements showed associations with modified intra-RSN connectivity in these networks, except for the DCA in the cingulo-opercular network 5. The AMI scores showed stronger associations with the cingulo-opercular and frontoparietal networks, whereas the AES had stronger associations with the default mode network and the goal-oriented behaviour network. The study was limited by the small number of participants without apathy according to the DCA, which may have reduced the statistical power of between-group comparisons. Additionally, the reliance on specific apathy measures may have influenced the observed overlap in brain regions. Conclusion: Our findings indicate that apathy in LLD is consistently associated with changes in both intra-RSN and inter-RSN connectivity of brain regions implicated in goal-oriented behaviours. These results corroborate previous findings of altered functional RSN connectivity in severe LLD.
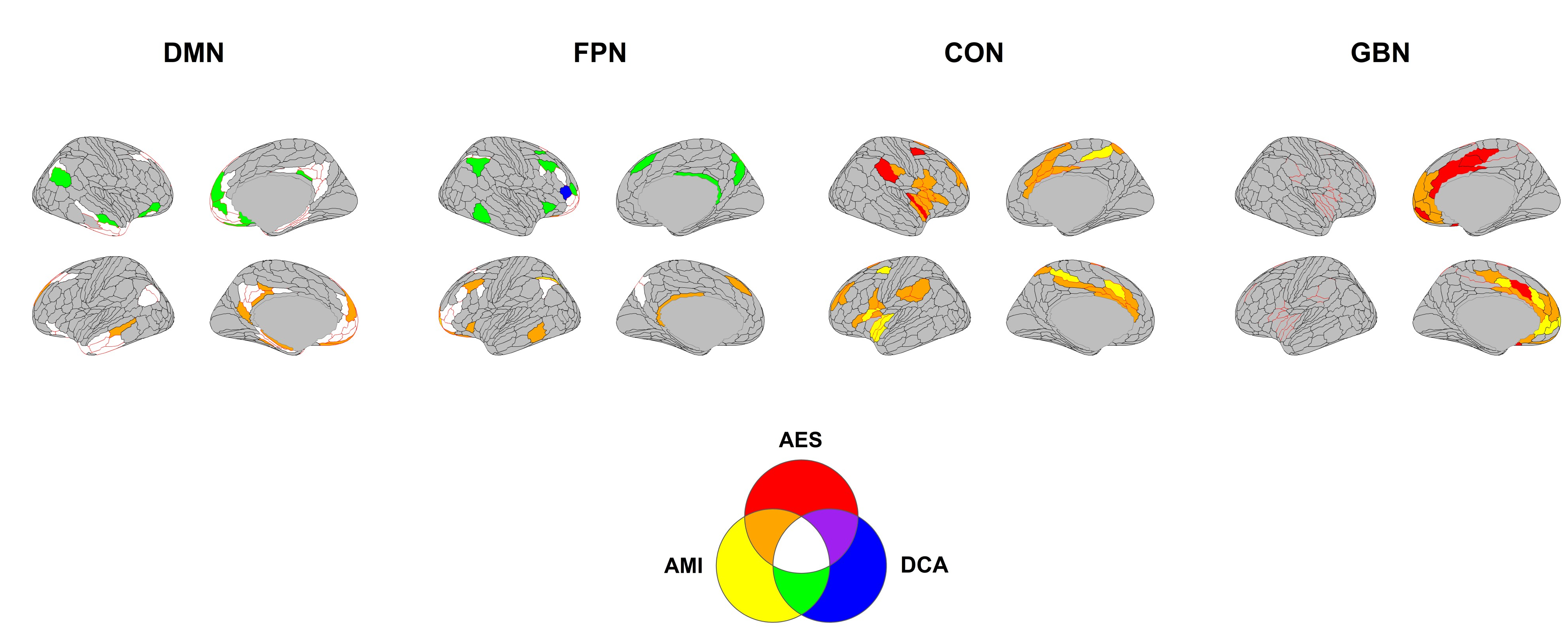
Representation of the brain regions whose functional connectivity are correlated with the apathy measures (AES, AMI and DCA).
Brain regions common to the different apathy measures. The brain regions identified as being associated with apathy across the measures are represented in colour. AES = Apathy Evaluation Scale; AMI = Apathy Motivation Index; CON = cingulo-opercular network; DCA = diagnostic criteria for apathy; DMN = default mode network; FPN = frontoparietal network; GBN = regions associated with goal-directed behaviours.
Apathy in depression: An arterial spin labeling perfusion MRI study.
Participants: Jean-Marie Batail, Isabelle Corouge, Benoit Combès, Jean-Yves Gauvrit, Gabriel Robert, Christian Barillot, Jean-Christophe Ferre.
Apathy, as defined as a deficit in goal-directed behaviors, is a critical clinical dimension in depression associated with chronic impairment. Little is known about its cerebral perfusion specificities in depression. To explore neurovascular mechanisms underpinning apathy in depression by pseudo-continuous arterial spin labeling (pCASL) magnetic resonance imaging (MRI). Perfusion imaging analysis was performed on 90 depressed patients included in a prospective study between November 2014 and February 2017. Imaging data included anatomical 3D T1-weighted and perfusion pCASL sequences. A multiple regression analysis relating the quantified cerebral blood flow (CBF) in different regions of interest defined from the FreeSurfer atlas, to the Apathy Evaluation Scale (AES) total score was conducted. After confound adjustment (demographics, disease and clinical characteristics) and correction for multiple comparisons, we observed a strong negative relationship between the CBF in the left anterior cingulate cortex (ACC) and the AES score (standardized = -0.74, corrected p value = 0.0008). Our results emphasized the left ACC as a key region involved in apathy severity in a population of depressed participants. Perfusion correlates of apathy in depression evidenced in this study may contribute to characterize different phenotypes of depression 13.
Alcohol consumption intentions of French young people exposed to two alcohol warning formats displayed on ads
Participants: Quentin Duché, Élise Bannier.
The study investigated the brain activity and self-report alcohol consumption intentions of French young people exposed to two alcohol warning formats displayed on ads: Alcohol ads with small Text-only Warning (ATW) currently used in many countries vs Alcohol ads with larger text-and-Picture Warning (APW). Seventy-four eligible male drinkers aged 18-25 completed pre-scan, face-to-face individual visit with a practitioner and an fMRI scanning session. They viewed 288 stimuli (96 alcohol ads with ATW, the same 96 ads with APW, 96 water ads -control group-; viewed 3 seconds each). Participants reported if the ad makes them want to consume the product. Whole-brain analysis and complementary region-of-interest analyses were performed. Whole brain BOLD fMRI highlighted contrasting effects: APW, compared to ATW, increased activations in the precuneus, the angular gyrus, the occipital, frontal and temporal areas, while the nucleus accumbens, the ventral tegmental areas, the putamen were less activated with APW. The region-of-interest analysis confirmed reduced activations in the reward circuit when presenting APW as compared to ATW. Regarding self-report responses, the tested ads elicited less desire to consume the promoted alcohol product when APW were displayed compared to ATW. Our findings suggest that stronger and text-and-picture warnings display in ads reduce the activity of key regions of the reward system and may influence the desire to consume alcohol products. These results provide advice for governments interesting in developing more effective labelling measures to target young people.
Subjective feeling of control during fNIRS-based neurofeedback targeting the DL-PFC is related to neural activation determined with short-channel correction
Participants: Élise Bannier.
Neurofeedback (NF) training is a promising preventive and therapeutic approach for brain and behavioral impairments, the dorsolateral prefrontal cortex (DL-PFC) being a relevant region of interest. As part of a collaboration with the EAT group at Inrae and the PhD of Ambre Godet, functional near-infrared spectroscopy (NIRS) has recently been applied in NF training of the dorsolateral prefrontal cortex. However, this approach is highly sensitive to extra-cerebral vascularization, which could bias measurements of cortical activity. Here, we examined the feasibility of a NF training targeting the DL-PFC and its specificity by assessing the impact of physiological confounds on NF success via short-channel offline correction under different signal filtering conditions. We also explored whether the individual mental strategies affect the NF success. Thirty volunteers participated in a single 15-trial NF session in which they had to increase the oxy-hemoglobin (HbO2) level of their bilateral DL-PFC. We found that 0.01-0.09 Hz band-pass filtering was more suited than the 0.01-0.2 Hz band-pass filter to highlight brain activation restricted to the NF channels in the DL-PFC. Retaining the 10 out of 15 best trials, we found that 18 participants (60%) managed to control their DL-PFC. This number dropped to 13 (43%) with short-channel correction. Half of the participants reported a positive subjective feeling of control, and the "cheering" strategy appeared to be more effective in men (p<0.05). Our results showed successful DL-PFC fNIRS-NF in a single session and highlighted the value of accounting for extra cortical signals, which can profoundly affect the success and specificity of NF training. 30
8.2.2 Neuro-inflammation
This year, we pursued our investigations regarding the relevance of imaging the spinal cord to target early biomarkers for MS.
Spinal cord microstructural damage measured in recently diagnosed Relapsing Remitting MS patients: prognostic value at 5-year
Participants: Vivien Caron, Benoit Combès, Malo Gaubert, Gilles Edan, Elise Bannier, Anne Kerbrat.
Early spinal cord (SC) lesions in patients with relapsing-remitting MS (RRMS) are associated with an increased risk of disability in the medium term. However, accurate quantification of these lesions on conventional MRI is difficult and imperfectly reflects the severity of SC damage. In this study, we assessed the added value of different metrics extracted from quantitative spinal cord MRI and reflecting microstructure to predict patient disability at 5 years. More specifically, we assessed the relationship between baseline SC fractional anisotropy (FA) and magnetization transfer ratio (MTR), the occurrence of atrophy and disability at 5-year in early RRMS patients and their added value compared to initial brain and SC lesion load. After IRB approval (NCT02117375), 76 RRMS patients (disease duration <1 year; mean EDSS=0.78) were included in a multicenter study and scanned at baseline and 5 years. For each subject, we measured 1) SC microstructural damage using magnetization transfer ratio (MTR) and DTI fractional anisotropy (FA) averaged over C4C6; 2) SC atrophy using cross sectional area (CSA) averaged over C2C3; 3) SC lesion load manually segmented on axial cervical T2*w; 4) brain lesion load automatically segmented on 3D FLAIR. Partial correlations between each quantitative metric and disability score or atrophy measurement at 5-year were calculated, with age and gender, and in a second model with SC lesion load as additional covariates. Overall, the 5-year EDSS was associated with the baseline SC lesion load (r=.29, p=.049), but not with the brain lesion load. The 5-year pyramidal sub-score was not associated with either cervical or brain lesion load. Concerning the microstructural components, both the baseline SC FA and MTR values were associated with the EDSS (r=-.32, p=.02; r=-.31, p=.04, resp.) and the pyramidal sub-score at 5-year (r=-.38, p=.01; r=-.42, p=.005, resp.). These associations were confirmed independently of cervical lesion load for cervical FA value and 5-year EDSS and for FA and MTR values and 5-year pyramidal sub-score. By contrast, we found no significant association between initial brain or SC lesion load or initial microstructural damage and evolution of CSA at 5-year. In conclusion, we highlighted the dominant role of initial SC involvement in the subsequent development of disability in early RRMS patients. In particular, initial spinal cord MTR and FA values may offer a reliable complement to lesion volume, able to capture lesion severity and non focal structural changes in this key structure.
Myelin dynamics in the spinal cord predict cord atrophy and disability progression at 5-years in early relapsing-remitting MS
Participants: Malo Gaubert, Benoit Combès, Elise Bannier, Arthur Masson, Vivien Caron, Gaëlle Baudron, Jean-Christophe Ferré, Anne Kerbrat.
Despite the major prognostic value of early spinal cord (SC) damage in MS, the processes of demyelination and remyelination in this structure and their clinical relevance remain to be evaluated. Thus, magnetization transfer ratio (MTR) changes in the cervical SC were used to generate patient-specific profiles of myelin content change and investigated their clinical relevance. In this work 44, our objectives were twofold: i) to characterise myelin content changes in the SC over a period of 1 year in early relapsing-remitting MS patients (RRMS); ii) to investigate the association between SC myelin content changes with disability and cross-sectional area (CSA) at 5-year. Thirty-seven RRMS patients (disease duration<1 year; mean EDSS=0.6 at baseline [BL]) underwent a cervical SC MRI at BL, 1 year and 5 years, and 19 healthy controls (HC) at BL only. SC lesions were manually segmented on T2*w cervical axial images at BL. CSA in C2C3 was computed at BL and 5 years on T1w brain images. Based on MTR maps of HC, SC MTR z-maps were computed for each MS patient at BL and 1 year and binarized at -2.58 (p=.01) as a proxy of demyelination. A global index of myelin content change (GIMCC) was calculated as the proportion of voxels classified as normal at BL and identified as demyelinated after 1 year (demyelination over time) minus the proportion of voxels classified as demyelinated at BL but not at 1 year (remyelination over time). Partial correlations between GIMCC and disability scores or CSA at BL and 5 years were calculated, with age and gender as covariates. A wide variability of GIMCC (from -13% to 28%) was observed, with 18 patients showing a predominance of demyelination over 1 year (GIMCC>0) and 18 a predominance of remyelination (GIMCC<0). Greater GIMCC, reflecting a predominant process of demyelination over remyelination in the SC, was associated with more severe disability at 5 years (EDSS p=.007) and with greater disability progression over the follow-up (EDSS change over 5 years p=.012). Greater GIMCC was associated with reduced CSA at 5 years (p=.032) and BL (p=.023), but not with lesion volume at BL (p=.36). To sum-up, patients with early RRMS exhibit heterogeneous profiles of SC myelin loss and repair measured over a period of one year. Greater SC remyelination during the first year was significantly associated with lower disability progression and greater SC volume 5 years later. These results highlight the potential of myelin repair in the SC to prevent neurodegeneration and clinical progression in patients with MS.
Impact of automatic tools for detecting new lesions on therapeutic strategies offered to patients with MS by neurologists
Participants: Blandine Merkel, Arthur Masson, Gilles Edan, Benoit Combès, Anne Kerbrat.
Automatic tools for detecting new lesions in patients with MS between two MRI scans are now available to clinicians. They have been assessed from the radiologist's point of view, but their impact on the therapeutic strategies that neurologists offer their patients has not yet been documented. In this study we aimed at comparing neurologist's decisions according to whether a lesion detection support system had been used and describe variability between neurologists on decision-making for the same clinical cases. We submitted 28 clinical cases associated with pairs of MRI images and radiological reports (produced by the same radiologist without vs. with the help of a system to detect new lesions) to 10 neurologists who regularly follow patients with MS. They examined each clinical case twice (without vs. with support system) in two sessions several weeks apart, and their patient management decisions were recorded. There was considerable variability between neurologists on decision-making (both with and without support system). When the support system had been used, neurologists more often made changes to patient management (75% vs. 68% of cases, p = 0.01) and spent significantly less time analyzing the clinical cases (249 s vs. 216 s, p = 3.10-4). In conclusion, the use of a lesion detection support system has an impact not only on radiologists' reports, but also on neurologists' subsequent decision-making. This observation constitutes another strong argument for promoting the wider use of such systems in clinical routine. However, despite their use, there is still considerable variability in decision-making across neurologists, which should encourage us to refine the guidelines.
8.2.3 Recovery
Improving outcome after paediatric concussion: challenges and possibilities
Participants: Fanny Dégeilh.
The term concussion has permeated mainstream media and household vocabulary mainly due to awareness regarding the risks of concussion in professional contact sports, yet it occurs across a variety of settings and ages. Concussion is prevalent in infants, preschoolers, children, and adolescents, and is a common presentation or reason for referral to primary care providers, emergency departments, and specialised trauma clinics. Its broad range of symptoms and sequelae vary according to multiple individual, environmental, and clinical factors and can lead to health and economic burden. More than 20 years of research into risk factors and consequences of paediatric concussion has revealed as many questions as answers, and scientific work and clinical cases continue to expose its complexity and heterogeneity. In this Review, we present empirical evidence for improving outcome after paediatric concussion. We consider work pertaining to both sports and other injury mechanisms to provide a perspective that should be viewed as complementary to publications focused specifically on sports concussion. Contemporary challenges in prevention, diagnosis, prognosis, and intervention are discussed alongside pathways and future directions for improving outcome. 14
Behavioral-play familiarization for non-sedated magnetic resonance imaging in young children with mild traumatic brain injury
Participants: Fanny Dégeilh.
Background: Mild traumatic brain injury (mTBI) sustained in early childhood affects the brain at a peak developmental period and may disrupt sensitive stages of skill acquisition, thereby compromising child functioning. However, due to the challenges of collecting non-sedated neuroimaging data in young children the consequences of mTBI on young children’s brains have not been systematically studied. In typically developing preschool children (TDC, 3-5 years), brief a behavioral-play familiarization provides an effective alternative to sedation for acquiring awake magnetic resonance imaging (MRI) in a time- and resource-efficient manner. To date, no study has applied such an approach for acquiring non-sedated MRI in preschool children with mTBI who may present with additional MRI acquisition challenges such as agitation or anxiety. Objective: The present study aimed to compare the effectiveness of a brief behavioral-play familiarization for acquiring non-sedated MRI for research purposes between young children with and without mTBI, and to identify factors associated with successful MRI acquisition. Materials and methods: Preschool children with mTBI (n=13) and TDC (n=24) underwent a 15-minute behavioral-play MRI familiarization followed by a 35-minute non-sedated MRI protocol. Success rate was compared between groups, MRI quality was assessed quantitatively, and factors predicting success were documented. Results: Among the 37 participants, 15 TDC (63%) and 10 mTBI (77%) reached the MRI acquisition success criteria (i.e., completing the two first sequences). The success rate was not significantly different between groups (p=.48; 95% CI [-0.36 14.08]; Cramer’s V=.15). The images acquired were of high-quality in 100% (for both groups) of the structural images, and 60% (for both groups) of the diffusion images. Factors associated with success included older child age (B=0.73, p=.007, exp(B)=3.11, 95% CI [1.36 7.08]) and fewer parental concerns (B=-1.56, p=.02, exp(B)=0.21, 95% CI [0.05 0.82]) about the MRI procedure. Conclusion: Using brief behavioral-play familiarization allows acquisition of high-quality non-sedated MRI in young children with mTBI with success rates comparable to those of non-injured peers. 21 From 2024, this protocol will be used to acquire non-sedated MRI data in young children with a mTBI at the Neurinfo platform.
Social problems and brain structure development following childhood mild traumatic brain injury
Participants: Fanny Dégeilh.
Childhood mild traumatic brain injury (mTBI) is associated with elevated risk of developing social problems, which may be underpinned by changes in the structural developmental trajectory of the social brain, a network of cortical regions supporting social cognition and behavior. However, limited sample sizes and cross-sectional designs generally used in neuroimaging studies of pediatric TBI have prevented explorations of this hypothesis. This longitudinal retrospective study examined the development of parent-reported social problems and cortical thickness in social brain regions following childhood mTBI using data from the large population-based Adolescent Brain Cognitive Development (ABCD) Study. Two-group latent change score models revealed different developmental trajectories from ages 10 to 12 years in social problems between children with (n=345) and without (n=7,089) mTBI. Children with mTBI showed higher levels of social problems than controls at age 10. Then, social problems decreased over 2 years, but still remained higher than in controls in which they stayed stable. Both groups showed similar decreases in social brain cortical thickness between ages 10 and 12 years. Further studies providing detailed information on the injury mechanism and acute symptoms are needed to better understand individual differences in social impairment and brain development in pediatric TBI. 23
EEG-fMRI Neurofeedback versus Motor Imagery after stroke, a Randomized Controlled Trial
Participants: Elise Bannier, Isabelle Bonan, Quentin Duché, Mathis Fleury, Pierre Maurel.
Bimodal EEG-fMRI neurofeedback (NF) is a guided brain activity self-regulation technique which allows target brain regions to learn and regulate their activity. In the case of affected by the stroke it can be used to try to reduce the motor impairment. In this study we investigated whether chronic stroke survivors were able to improve their motor performances better with NF than with a motor imagery (MI) training without feedback. We carried out a randomized controlled trial in which 30 chronic stroke patients with upper-limb partial motricity and preserved corticospinal tract followed a five-week rehabilitation protocol. They either performed a bimodal EEG-fMRI NF training on ipsilesional motor areas (M1 and SMA) (n=15) or a MI training (n=15). Clinical and brain assessments were performed at the beginning and at the end of the protocol and an extra clinical assessment was performed one month after the training. The primary outcome measure was the Fugl-Meyer Assessment Upper Extremity score (FMA-UE). All NF patients completed the training and succeeded in modulating their brain activity in the target regions. In terms of clinical outcomes, we found that FMA-UE scores increased in the NF group only (p=0.003 vs p=0.633 for MI) and maintained this improvement one month after the protocol (p=0.029). The FMA-UE improvement was higher in the NF group (p=0.048). Overall, 8/15 patients in the NF group and 3/15 in the MI group were clinical responders.. Concerning brain activity results, the NF group showed an increase in ipsilesional M1 (t=1.2, p=0.23) and SMA (t=0.7, p=0.47) BOLD activation. MRI laterality index (LI) in M1 increased in a non-significant way for both groups. A significant difference emerged between the groups for unimodal EEG LI, with greater lateralization in the NF group (t=-3.56, p=0.0004). We demonstrate that chronic stroke patients can follow a personalized bimodal EEG-fMRI NF training to self-regulate their brain activity. This training was more efficient in improving motor recovery than the MI training.
Two is better ? Combining EEG and fMRI for BCI and Neurofeedback : A systematic review
Participants: Mathis Fleury.
Electroencephalography (EEG) and functional magnetic resonance imaging (fMRI) are two commonly used non-invasive techniques for measuring brain activity in neuroscience and braincomputer interfaces (BCI). While EEG has high temporal resolution and low spatial resolution, fMRI has high spatial resolution and low temporal resolution. In this review, we focus on the use of EEG and fMRI in neurofeedback (NF) and discuss the challenges of combining the two modalities in order to improve understanding of brain activity and achieve more effective clinical outcomes. Advanced technologies have been developed to simultaneously record EEG and fMRI signals in order to better understand the relationship between the two modalities. However, the complexity of brain processes and the heterogeneous nature of EEG and fMRI present challenges in extracting useful information from the combined data. We will survey existing EEG-fMRI combinations and recent studies that exploit EEG-fMRI in NF, highlighting the experimental and technical challenges. We will also identify remaining challenges in this field 53.
9 Bilateral contracts and grants with industry
9.1 Bilateral contracts with industry
9.1.1 Siemens
Participants: Elise Bannier, Emmanuel Caruyer, Isabelle Corouge, Jean-Christophe Ferré, Jean-Yves Gauvrit.
A collaboration between Siemens, Empenn and the Neurinfo platform is in place and formalized by a research contract. Thanks to this agreement, the Neurinfo platform has received the object code of MRI sequences under development at Siemens for evaluation in clinical research. In addition, the Neurinfo platform has received the source code of selected MRI sequences. As a result, MRI sequences can be developed on site by our team. For example, an MRI diffusion sequence was modified to load arbitrarly diffusion gradient waveforms for the FastMicroDiff project (led by E. Caruyer).
10 Partnerships and cooperations
10.1 International research visitors
10.1.1 Visits of national and international scientists
Demian Vera
-
Status
PhD Candidate
-
Institution of origin:
Institution of origin: National University of Central Buenos Aires
-
Country:
Argentina
-
Dates:
Since Oct 2022 until January 2023
-
Context of the visit:
Implementation of a multilayer fNIRS model and experimentation at the Neurinfo platform
Laure Fournier
-
Status
PU-PH
-
Institution of origin:
AP-HP
-
Country:
France
-
Dates:
January 10th
-
Context of the visit:
HDR of Elise Bannier
Michel Dojat
-
Status
Senior Researcher
-
Institution of origin:
GIN, Grenoble
-
Country:
France
-
Dates:
January 10th
-
Context of the visit:
HDR of Elise Bannier
Sophie Achard
-
Status
Senior researcher
-
Institution of origin:
Laboratoire Jean Kuntzmann
-
Country:
France
-
Dates:
November 20-22
-
Context of the visit:
PhD of Jean-Charles Roy
Olivier Coulon
-
Status
Senior researcher
-
Institution of origin:
Institut des Neurosciences de la Timone
-
Country:
France
-
Dates:
December 15
-
Context of the visit:
PhD of Thomas Durantel
10.1.2 Visits to international teams
Research stays abroad
- Caroline Pinte, international mobility to Advanced Telecommunications Research Institute International (ATR) Kyoto, Japan, September to October 2023. Project: The DecNef Lab at ATR plays a significant role in the neurofeedback field, being the originator of a methodology that has paved the way for a new branch termed "Decoded Neurofeedback". This approach involves learning a pattern of brain activity without providing any re-education instructions to the patient. During my stay at the DecNef Lab, my focus was on comprehending the methodology of decoded neurofeedback and its connections with our conventional neurofeedback protocol. Presenting my work during an invited talk also initiated a collaboration to enhance technical insights into the measurement and processing of EEG signals through Domain Adaptation (DA) methods, including Euclidean space alignment.
- Elodie Germani, international mobility to Big Data for NeuroInformatics lab (Concordia University) and NeuroDataScience - ORIGAMI lab (McGill University), Montreal, Canada, August to November 2023. Project: The LivingPark project aims to improve the generalizability and robustness of MRI-derived biomarkers of Parkinson's Disease through analytical and data variability evaluations. During this mobility, we extended the project to functional MRI-derived biomarkers. We reproduce (same data, same method) and replicate (different data or method) the models used in a study to predict individual's Parkinson's disease current state and progression using demographic, clinical and neuroimaging features (fALFF and ReHo extracted from resting-state fMRI). Using the reproduction workflow, we managed to obtain better than chance performance for all our models, but this performance remained very different from the ones reported in the original study. The challenges encountered while reproducing and replicating the original work are likely explained by the complexity of neuroimaging studies, in particular in clinical settings.
Visit to international teams for collaboration
- Fanny Dégeilh, visit to ABCs (PI: Miriam Beauchamp) and Grandir ensemble (PI: Annie Bernier) labs, University of Montreal, Montreal, Canada, July 2023 (1 week). This visit took place in the context of several pediatric neuroimaging projects within the ABCs and Grandir Ensemble laboratories with which Fanny Dégeilh collaborates. The aim of the working week was 1) to plan the continuation of work on which our laboratories are collaborating, both on healthy and clinical populations (e.g. pediatric traumatic brain injury) and 2) to identify common axes and projects that could be carried out in collaboration between our laboratories, in particular with a view to strengthening scientific links between the Quebec teams and Empenn.
10.2 National initiatives
10.2.1 ANR-20-THIA-0018: programme Contrats doctoraux en intelligence artificielle
Participants: Francesca Galassi, Ricky Walsh, Benoît Combès.
-
Funding:
Co-funding for a PhD thesis in AI - Duration: 2022-2025.
-
Summary:
Co-funding (50% with Univ. Rennes) for a doctoral program in Artificial Intelligence. The PhD concerns the automatic segmentation of MS lesions in spinal cord MRI by means of AI-based solutions.
10.2.2 ANR-20-THIA-0018: programme Contrats doctoraux en intelligence artificielle
Participants: Camille Maumet, Elodie Germani.
-
Funding:
Co-funding for a PhD thesis in AI - Duration: 2021-2024.
-
Summary:
Co-funding (50% with Univ. Rennes) for a doctoral program in Artificial Intelligence. The PhD concerns representation learning for reproducible neuroimaging.
10.2.3 RHU PRIMUS: Transforming the care of patients with Multiple Sclerosis using a multidimensional data-driven clinical decision support system
Participants: Elise Bannier, Benoît Combès, Gilles Edan, Jean-Christophe Ferré, Francesca Galassi, Anne Kerbrat.
-
Funding:
RHU - Duration: 2022-2026 - Budget: 8272kC
-
Partners:
Observatoire Français de la Sclérose en Plaques (OFSEP), France Life Imaging (FLI), Pixyl.
-
Summary:
The overall objective of PRIMUS is to develop and validate a CE-marked data-driven clinical decision support system (CDSS) for multiple sclerosis (MS). The CDSS will support clinical decision-making by providing easily interpretable information on treatment options. MS is a complex disease, with different phenotypes and heterogeneous progression patterns. Over the past two decades, MS practice has been flooded with data and the number of available treatments has considerably increased. Although clinical, biological and imaging information is now being generated on a massive scale, it contributes to clinical decision-making in a rather haphazard, siloed and non-standardised fashion, so that selecting the most appropriate therapeutic option remains hard. PRIMUS contributes to data-driven homogenization of shared decision practices with and for patients with MS. To achieve this goal, the project will develop advanced artificial intelligence solutions, for a patient- and physician-centred CDSS.
10.2.4 EyeSkin-NF : Eye-tracking and skin conductance measures for neurofeedback analysis and validation
Participants: Claire Cury, Elise Bannier, Pierre Maurel, Hachim Bani, Rene-Paul Debroize, Agustina Fragueido.
-
Funding:
Exploratory action Inria - Duration: 2021 - 2024.
-
Summary:
Neurofeedback techniques (NF) or restorative brain computer interfaces (BCI) consist in providing a subject with real-time feedback about its own brain activity, in order to learn self-regulate specific brain regions during NF training. Brain activity can be measured by various techniques such as EEG and/or fMRI. However, analysis of NF sessions is limited due to the difficulty at identifying the origin of failed training. To enhance and monitor participant's motivation in real-time during EEG-fMRI recording, bio-signal can be measured via eye-tracking (ET) or skin conductance (SC) devices. For a precise evaluation of the motivation mental states of interest such as focus, arousal, mind wandering or mental load can be analysed. The main objective of this project is to investigate measures from eye-tracking and skin conductance signals to evaluate in real-time subject’s motivation during NF training.
10.2.5 GRASP: Generalizing Results Across Scientific Pipelines
Participants: Camille Maumet, Boris Clenet, Jeremy Lefort-Besnard.
-
Funding:
Exploratory action Inria - Duration: 2022 - 2024.
-
Summary:
Scientific pipelines are at the heart of modern experimental sciences. But practionners face a highly complex pipeline landscape – different tools, algorithms, parameters – in which different pipelines can lead to contradictory research findings. GRASP will model pipeline-induced variability to derive valid and generalizable results in the field of brain imaging.
10.2.6 ANR-NODAL: Identification de biomarqueurs de maladies neurodégénératives par l'analyse de la connectivité multimodale.
Participants: Julie Coloigner, Carlo Ferritto.
-
Funding:
Appel à projets générique 2022 - Duration: 2022 - 2026.
-
Summary:
The neurodegenerative diseases like Alzheimer’s (AD) and Parkinson’s (PD) disease are the consequences of pathological processes that begin decades before the onset of the typical clinical symptoms. However, current diagnosis comes quite late in the course of the disease, while evidences underline the multiple benefits that would be associated with earlier diagnosis. An outstanding challenge for clinical neurosciences is therefore to provide reliable, non-invasive, affordable and easy-to-track biomarkers able to improve both the early detection and the monitoring of neurodegenerative diseases. Recent advances in non-invasive connectome mapping techniques offer great hope for significant progress in taking up this challenge by investigating cerebral organization. Indeed, it is well acknowledged that AD and PD display a progressive multifactorial disruption of functional and structural cerebral networks, all along the course of the diseases. A recent framework called Graph Signal Processing (GSP) is particular promising to shed new light on the complex interplay between brain function and structure. For the first time, GSP will be extended to the development of more sensitive metric of AD and PD progression, taking into account the cerebral functional-structural coupling, contrary to the classical biomarkers using a single-modality data or clinical assessment. In the PRESCO project, we will develop a new multimodal and multi-stage approach using innovative machine learning methods, adapted for GSP-based features, to provide non-invasive, reliable and easy-to-track candidate biomarkers for each stage of AD and PD diseases. We will apply this approach on two large patients’ cohorts. Then, we will assess the effectiveness of candidate disease-specific biomarkers on a new innovative local multimodal cohorts including patients with and without cognitive impairment, at various stages of AD and PD. At the end of 2023, we began the acquisition of the cohort including the MRI data and neurocognitve assessment. Carlo Ferritto, a PhD student, works on this project from October 2023. This is a collaborative project with Pierre-Yves Jonin, CHU Rennes and Giulia Lioi, researcher, IMT, Brest.
10.2.7 ANR-PASTRAMI: Patient-specific statistics for microstructure-augmented connectomics
Participants: Élise Bannier, Emmanuel Caruyer, Julie Coloigner, Claire Cury, Marie Poirier.
-
Funding:
Appel à projets générique 2023 - Duration: 2023 - 2028.
-
Summary:
The PASTRAMI project proposes to promote the use of diffusion magnetic resonance imaging (MRI) to derive biomarkers of axonal injury along white matter (WM) fascicles as prognostic factors of functional recovery after severe traumatic brain injury (TBI). We propose to develop statistical methods for patient-specific localization of abnormalities in microstructure and/or structural connectivity, along specific WM fascicles and/or on the full connectome. In a clinical study, the objective will be to assess the predictive accuracy of the proposed model evaluated in the 10 days period following TBI to predict unfavourable outcome at 1-year after the first injury in patients admitted in intensive care for severe TBI. This project is a collaboration with the Laboratoire de mathématiques Jean Leray (Nantes), CHU Rennes and the HIA Sainte-Anne (Toulon).
10.2.8 ANR-JCJC-VICUNA: Exploring the variabiIity induced by different configurations in the neuroimaging analytical space
Participants: Camille Maumet.
-
Funding:
Appel à projets générique 2022 - Duration: 2022 - 2026.
-
Summary:
Using the same data to answer the same scientific question, researchers may reach contradictory conclusions depending on the analytical pipeline they choose. For many years this problem has been rampant in experimental sciences and recent studies stemming from many fields have brought scientific evidence of this issue. Overall this phenomenon has reduced confidence in research findings and is effectively an important remaining driver of the reproducibility crisis. Software is central to modern scientific research and with the development of data science and its subfields (such as bioinformatics or neuroinformatics) the different tools and approaches available to study a dataset have multiplied. Those software have been very valuable to practitioners and brought the capacity to process more data in a shorter amount of time. But overall, they also provide a large number of possible analysis paths that can be used in order to address a scientific question. With VICUNA, we will provide a proof-of-concept explorat analytical variability in brain imaging. We will navigate in the pipeline space at large and understand which parts of the space are effectively in-use. We will explore and look into how results vary in 3 large open datasets (NARPS, UK Biobank and HCP). This is a collaborative project with Mathieu Acher, INSA Rennes.
10.2.9 Connectivity of the amygdala in depression
Participants: Emmanuel Caruyer, Julie Coloigner, Claire Cury.
-
Funding:
Fondation de France + INCR – Institut des Neurosciences Cliniques de Rennes - Duration: 2019-2023 - Budget: 250k€
-
Summary:
The onset of depression in teenagers and young adults increases the risk to develop a drug-resistant depression in the adulthood. This project aims at evaluating the role of early changes in the microstructure and connectivity of the amygdala. Using a cohort of drug-resistant patients (N=30), non drug-resistant patients (N=30) and controls (N=30), the aim is to identify imaging biomarkers of the pathology and to compare these with emotional and cognitive phenotypes in this population, searching for early differences in the development of the amygdala connectivity. Inclusions are ongoing. This is a colloborative project with M.-L. Paillère Martinot from Paris-Descartes University, as Principal Investigator.
10.2.10 Knowledge addition through Neuroimaging of Alcohol consumption in healthy young Volunteers, causes or consequences
Participants: Elise Bannier, Quentin Duché, Gabriel Robert.
-
Funding:
Funding: INCR - Duration: 2020-2023 - Budget: 45k€
-
Summary:
Alcohol consumption is responsible for 3 million annual deaths worldwide (5.1 percent of the global burden of disease). It causes disease (liver cirrhosis, cancers, etc.) and other social costs (injuries, road accidents, alcohol dependence, etc.). Excessive alcohol consumption grows through adolescence. This type of behavior has also been shown to have subtle but significant deleterious effects on cognitive function in adolescents. Advances in the field of neuroimaging make it possible to characterize anatomical changes and the evolution of neuropsychological deficits. Besides, focusing on the societal causes of alcohol abuse, a large body of studies show that exposure to alcohol advertising through media bootstraps early consumption initiation, greater desire to drink, increased alcohol use and binge drinking patterns among young people, especially minors. We aim to combine the analysis of the locally acquired IMAJ dataset (PI Karine Gallopel-Morvan, INCA Funding) and data from the european consortium IMAGEN datasets to determine whether there are functional characteristics and external factors that can explain behavior towards alcohol and to extract biomarkers capable of predicting excessive behavior. Relying on the IMAJ dataset, we will analyze whether, depending on warning formats displayed on ads (small and text-only vs. larger, shock-inducing and pictorial), health messages can influence brain activity by decreasing the effect of attractive alcohol content ads on the reward system area and on behavioral responses. Relying on the already effective collaboration of Dr Robert with Prof Schumann, we will explore the longitudinal anatomical and functional data from the IMAGEN cohort to extract biomarkers of alcohol consumption evolution and complement the analysis with the results obtained from the IMAJ dataset.
10.2.11 PHRC EMISEP: Evaluation of early spinal cord injury and late physical disability in Relapsing Remitting Multiple Sclerosis
Participants: Elise Bannier, Emmanuel Caruyer, Benoit Combès, Gilles Edan, Jean-Christophe Ferré, Anne Kerbrat.
-
Funding:
PHRC - Duration: 2016-2023 - Budget: 200k€
-
Summary:
Multiple Sclerosis (MS) is the most frequent acquired neurological disease affecting young adults (1 over 1000 inhabitants in France) and leading to impairment. Early and well adapted treatment is essential for patients presenting aggressive forms of MS. This PHRC (Programme hospitalier de recherche clinique) project focuses on physical impairment and especially on the ability to walk. Several studies, whether epidemiologic or based on brain MRI, have shown that several factors are likely to announce aggressive development of the disease, such as age, number of focal lesions on baseline MRI, clinical activity. However, these factors only partially explain physical impairment progression, preventing their use at the individual level. Spinal cord is often affected in MS, as demonstrated in postmortem or imaging studies. Yet, early radiological depiction of spinal cord lesions is not always correlated with clinical symptoms. Preliminary data, on reduced number of patients, and only investigating the cervical spinal cord, have shown that diffuse spinal cord injury, observed via diffusion or magnetisation transfer imaging, would be correlated with physical impairment as evaluated by the (EDSS) Expanded Disability Status Scale score. Besides, the role of early spinal cord affection (first two years) in the evolution of physical impairment remains unknown. In this project, we propose to address these different issues and perform a longitudinal study on Relapsing Remitting Multiple Sclerosis (RRMS) patients, recruited in the first year of the disease. Our goal is to show that diffuse and focal lesions detected spinal cord MRI in the first two years can be used to predict disease evolution and physical impairment at 5 years. Twelve centers are involved in the study to include 80 patients. To date, all subjects have been included and the last visit of the last patient is scheduled early 2023. The EMISEP data consists of brain and spinal cord structural and quantitative MR images of early MS patients followed over 5 years. Four papers have been published so far on data acquired at baseline on healthy controls and patients. Three papers were co-authored in the context of international collaborations. Additional papers are in preparation.
10.2.12 Estimating the impact of multiple sclerosis lesions in motor and proprioceptive tracts, from the brain to the thoracic spinal cord, on their functions, assessed from clinical tests (MS-TRACTS and MAP-MS)
Participants: Elise Bannier, Benoit Combès, Malo Gaubert, Anne Kerbrat.
-
Funding:
ARSEP, COREC and INCR - Duration: 2020-2023 - Budget: 200k
-
Summary:
Previous studies, whether epidemiologic or based on brain MRI, have shown that several factors were likely to announce aggressive development of the disease, such as age, clinical relapses, number of focal lesions on baseline MRI. However, these factors only partially explain physical disability progression, preventing their use at the individual level. We hypothesize that a fine assessment of damage on specific networks, from the brain to the thoracic cord, offers a relevant biomarker of disability progression in MS. Such damage assessments must take into account both lesion location, assessed on structural brain and cord MR images and lesion severity, assessed using advanced brain and cord imaging through quantitative MRI.We propose to test this hypothesis by combining assessments of lesion location and severity on corticospinal and proprioceptive tracts from the brain to the thoracic cord with clinical and () electrophysiological measurements. The MS-TRACTS study involves two French centers (Rennes,Marseille) and includes a total of 60 relapsing remitting MS patients. The expected outcome is to obtain early biomarkers of physical impairment evolution in RRMS patients, first treated with immunomodulatory treatment. The long-term goal is to provide the clinician with biomarkers able to anticipate therapeutic decisions and support the switch to alternative more aggressive treatment. Inclusions are ongoing. The MAP-MS study involves the same two French centers and will nclude 40 progressive MS patients. The investigation will focus on motor asymetry in these more advanced patients. This study includes two French centers (Rennes, Marseille) and includes a total of 60 patients. The expected outcome is to obtain early biomarkers of physical impairment evolution in RRMS patients, first treated with immunomodulatory treatment. The long-term goal is to provide the clinician with biomarkers able to anticipate therapeutic decisions and support the switch to alternative more aggressive treatment. Inclusions are ongoing.
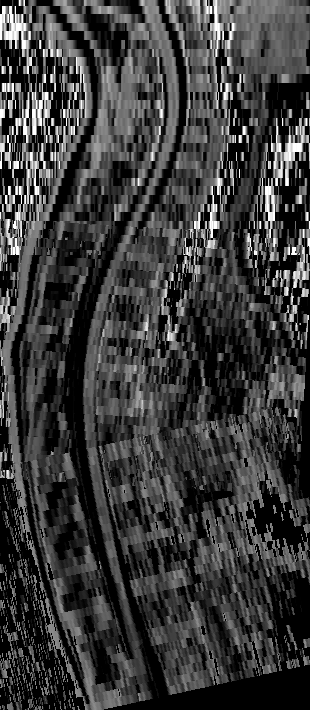

Estimating the impact of multiple sclerosis lesions in motor and proprioceptive tracts, from the brain to the thoracic spinal cord, on their functions, assessed from clinical tests (MS-TRACTS and MAP-MS): An example of Magnetization Transfer Ratio (MTR) mapping of the whole spinal cord acquired from the MS-TRACTS imaging protocol.
10.2.13 France Life Imaging (FLI)
Participants: Michael Kain, Camille Maumet, Jean-Christophe Ferré.
-
Funding:
Funding: FLI - Duration: 2012-2023 - Total budget: 2000k€ (phase 1) + 1200k€ (phase 2) + 800k€ (phase 3)
-
Summary:
France Life Imaging (FLI) is a large-scale research infrastructure project to establish a coordinated and harmonized network of biomedical imaging in France. This project was selected by the call “Investissements d’Avenir - Infrastructure en Biologie et Santé”. One node of this project is the node Information Analysis and Management (IAM), a transversal node built by a consortium of teams that contribute to the construction of a network for data storage and information processing. Instead of building yet other dedicated facilities, the IAM node use already existing data storage and information processing facilities (LaTIM Brest; CREATIS Lyon; CIC-IT Nancy; Empenn U1228 Inria Rennes; CATI CEA Saclay; ICube Strasbourg) that increase their capacities for the FLI infrastructure. Inter-connections and access to services are achieved through a dedicated software platform that is developed based on the expertise gained through successful existing developments. The IAM node has several goals. It is building a versatile facility for data management that inter-connects the data production sites and data processing for which state-of-the-art solutions, hardware and software, are available to infrastructure users. Modular solutions are preferred to accommodate the large variety of modalities acquisitions, scientific problems, data size, and to be adapted for future challenges. Second, it offers the latest development that are made available to image processing research teams. The team Empenn fulfills multiple roles in this nation-wide project. Michael Kain is the technical manager, Camille Maumet is part of the steering committee. Apart from the team members, software solutions like MedInria and Shanoir are part of the software platform.
10.2.14 OFSEP: French Multiple Sclerosis Observatory
Participants: Elise Bannier, Gilles Edan, Jean-Christophe Ferré, Francesca Galassi, Arthur Masson, Benoît Combès, Anne Kerbrat.
-
Funding:
ANR-PIA - Duration: since 2017 - Budget: 175k€
-
Summary:
The French Observatory of Multiple Sclerosis (OFSEP) is one of ten projects selected in January 2011 in response to the call for proposal in the "Investissements d’Avenir - Cohorts 2010" program aunched by the French Government. It allows support from the National Agency for Research (ANR) of approximately 10 million € for 10 years. It is coordinated by the Department of Neurology at the Neurological Hospital Pierre Wertheimer in Lyon (Professor Christian Confavreux), and it is supported by the EDMUS Foundation against multiple sclerosis, the University Claude Bernard Lyon 1 and the Hospices Civils de Lyon. OFSEP is based on a network of neurologists and radiologists distributed throughout the French territory and linked to 61 centers. OFSEP national cohort includes more than 50,000 people with Multiple Sclerosis, approximately half of the patients residing in France. The generalization of longitudinal monitoring and systematic association of clinical data and neuroimaging data is one of the objectives of OFSEP in order to improve the quality, efficiency and safety of care and promote clinical, basic and translational research in MS. For the concern of data management, the Shanoir platform of Inria has been retained to manage the imaging data of the National OFSEP cohort in multiple sclerosis. One long term objective of the OFSEP project is to identify prognostic factors of the evolution of Multiple Sclerosis. The HD Cohort is an enhanced cohort specifically designed for this purpose in which some patients are followed-up on a yearly basis. Additional clinical, quality of life and other patient-reported data is also collected. This study aims at developing personalized predictive tools to improve patient care management, and help in making decision to start, maintain or adapt medical care. Collected data will be processed to extract valuable information enabling to determine specific biomarkers of the evolution of the disease. Multiple Sclerosis brain lesions are of particular interest, hence the need for a careful comparison of lesion segmentation methods. A litterature review enabled to gather most promising cross-sectionnal methods, designed to identify and localize lesions with precise measurement of the lesion load at one particular point in time ; and longitudinal methods which gives more insight on the evolution of those lesions over the different time points. Those later methods are particularly interesting for clinicians for whom the type of lesion evolution is of foremost importance. A cross-sectionnal method and a longitudinal method were trained and evaluated to select the ones which will be used to analyze the entire HD Cohort dataset. Moreover, an experimental and a statistical design to compare the accuracy, sensitivity and specificity of the active/inactive classification of MS patients based on brain MRI as assessed using the analysis of brain was proposed. These designs will allow to assess the interest of re-analyzing the MRI data to improve the quality of the standardized reports used in most epidemiologic studies from the OFSEP cohorts. The collaboration has recently been extended until end of 2025 with a particular focus on spinal cord imaging and slowly evolving lesions.
10.2.15 QSM-SPICO: Quantitative Susceptibility Mapping for Spinal Cord
Participants: Elise Bannier, Benjamin Streichenberger, Anne Kerbrat, Benoit Combès.
-
Funding:
FLI-RE4 - 20k€
-
Summary:
Quantitative Susceptibility Mapping (QSM) is a promissing quantative imaging technique for the characterization of lesions in Multiple Sclerosis (MS). QSM provides a novel type of contrast linked to the tissue magnetic susceptibility. The latter is sensitive to iron accumulation and myelin content, which are both important metrics when studying MS lesions. As part as a research expertise transfer sponsored by France Life Imaging, in collaboratin with Mathieu Santin, at ICM/CENIR and Ludovic de Rochefort and Stéphane Roche from the Ventio Startup in Marseille, we are exploring the possibility to perform QSM in the spinal cord. This is challenging because of size of the cord and the presence of fat in the spine. To tackle this challenge, we use the IDEAL algorithm - iterative decomposition of water and fat with echo asymmetry and least-squares estimation. The aim is be able to characterize spinal MS lesions using QSM. The first results are encouraging.
10.2.16 PEPR ShareFAIR
Participants: Camille Maumet, Elise Bannier.
-
Funding:
PEPR Santé numérique.
-
Summary:
Access to a wide variety of complementary, multi-scale and massive data collections offers unprecedented opportunities for healthcare research. A large number of analyses can be performed on these datasets, for scientific advances and discoveries to emerge. The national 'Digital Health'Acceleration Strategy ambitions to boost digital health innovation which includes designing innovative health data analysis approaches.
Importantly, such data analyses are complex, they rely on various computational tools that have to be parametrized and chained together. There is now compelling evidence that many scientific discoveries will not stand the test of time: increasing the reproducibility of computed results is of paramount importance, especially in the healthcare domain.
Sharing of health data is often hampered by personal data protection requirements and comes up against technical constraints (security, volume). These constraints can however be limited when the protocols and the workflows implementing analyses are sufficiently reusable to reproduce analyses in situ.
Additionally, when designed to be reusable, protocols and their implementations - workflows - provide the provenance traces of the analyzed data, describing how data results have been obtained and thus increasing scientists’ confidence in the results produced.
This calls for innovative solutions for the annotation of biomedical and clinical datasets and extraction of provenance. Protocols and their implementation as workflows using and generating datasets should be elevated to first-class objects and the inherent dual relationship between datasets and protocols/workflows should be better exploited.
Challenges thus include standardization and annotation for datasets and protocols, extracting protocols and workflows from text and other datasets, and synthesizing them into interoperable, yet shareable protocols.
The originality of ShareFAIR lies in tackling both the reliability of datasets and analysis protocols and in harnessing the dual relationship between datasets and protocols. Specifically, ShareFAIR will provide:
(i) standards to uniformly represent datasets, ontologies/common vocabularies to annotate datasets and protocols/workflows, and provenance to trace the origin of datasets,
(ii) an interoperable framework for the design, annotation and reuse of reliable and shareable protocols,
(iii) approaches to extract protocols from textual data to enrich the set of protocols and workflows and better document the provenance of datasets, and approaches to learn protocols from biomedical and clinical datasets.
This project is led by Sarah Boulakia-Cohen from Univ Paris Saclay.
10.3 Regional initiatives
10.3.1 PEPERONI : Portable and Personalized Neurofeedback for Stroke Rehabilitation
Participants: Elise Bannier, Isabelle Bonan, Julie Coloigner, Isabelle Corouge, Claire Cury, Pierre Maurel, Camille Muller, Caroline Pinte.
-
Funding:
Labex CominLabs : from Sept. 2022 to end of 2024 - Budget: 290k€
-
Summary:
Neurofeedback (NF) consists in presenting a person with a stimulus directly related to his or her ongoing brain activity. NF can be used to teach subjects how to regulate their own brain functions by providing real-time sensory feedback of the brain “in action”. Recent studies showed that NF is promising for the treatment of various neuronal pathologies. Electroencephalography (EEG), which has historically been the preferred modality for NF, suffers from a lack of specificity, preventing the transfer of this treatment to clinical use. On the other hand functional Magnetic Resonance Imaging (fMRI) has a good specificity, but it is a cumbersome and expensive modality, making it difficult to develop personalized protocols. In this project, we aim to develop a methodological and experimental framework opening the door to a more portable and personalized NF, for easier and effective clinical use, with a focus on post-stroke motor rehabilitation. We propose to organize the project in four work packages, grouped in two axes. The following figure summarizes the organization of the project.
10.3.2 Région Bretagne: SAD
Participants: Burhan Rashid Hussein, Francesca Galassi, Benoit Combès.
-
Funding:
SAD 2022 - 2024 : RHU PRIMUS - Duration: 48 months (start: December 2022) - Budget: 75k€
-
Summary:
Complementary funding for a post-doctoral position (Burhan Rashid Hussein) within the RHU PRIMUS project.
10.3.3 Région Bretagne: ARED
Participants: Nolwenn Jégou, Anne Kerbrat, Benoit Combès.
-
Funding:
ARED 2023 - 2026 : RHU PRIMUS - Duration: 36 months (start: December 2023)
-
Summary:
Complementary funding for a PhD position (Nolwenn Jégou) within the RHU PRIMUS project.
10.3.4 Région Bretagne: ARED
Participants: Elodie Germani, Camille Maumet.
-
Funding:
ARED 2022 - 2024 : ARED MAPPIS - Duration: 36 months
-
Summary:
Complementary funding for a PhD position (Elodie Germani).
10.3.5 Région Bretagne: ARED
Participants: Sebastien Dam, Julie Coloigner, Pierre Maurel.
-
Funding:
ARED 2022 - 2024 : ARED CONGRATS - Duration: 36 months
-
Summary:
Complementary funding for a PhD position (Sébastien Dam).
11 Dissemination
11.1 Promoting scientific activities
Participants: Elise Bannier, Emmanuel Caruyer, Julie Coloigner, Benoit Combès, Claire Cury, Fanny Dégeilh, Quentin Duche, Agustina Fragueiro, Francesca Galassi, Burhan Rashid Hussein, Anne Kerbrat, Jérémy Lefort-Besnard, Pierre Maurel, Cédric Meurée, Caroline Pinte, Ricky Walsh.
11.1.1 Scientific events: organisation
General chair, scientific chair
- Julie Coloigner: scientific chair at the IEEE International Symposium on Biomedical Imaging (ISBI) 2023
Member of the organizing committees
- Julie Coloigner: communication chair and and session chair in IEEE International Symposium on Biomedical Imaging (ISBI) 2023
- Camille Maumet: session chair in IEEE International Symposium on Biomedical Imaging (ISBI) 2023
- Camille Maumet: session chair in OHBM 2023 for the oral sessions "Analysis and methods"
- Elise Bannier, Julie Coloigner and Quentin Duche are part of the local organization committe of the SFRMBM2025, to be held in March 2025 in Saint Malo.
- Elise Bannier is part of the organization commmittee of the MR Physics school to be held in April 2024 near Bordeaux.
Participation in organizing scientific events
- Caroline Pinte, participated in the organization of the D6 "Signal, Image, langage" department seminar at IRISA
- Jérémy Lefort-Besnard, contributed to the working group 2 of GliMR EU COST action.
- Elise Bannier, co-organized 3 meetings of the REMI network in January (virtual), in March (following the SFRMRM2023) and in November (in Paris) 2023. Jérémy Lefort-Besnard, participated to the virtual meeting of the Working Group 2nd level statistical analysis of the REMI network REMI (Réseau d'Entraide Multicentrique en IRM)
11.1.2 Scientific events: selection
Member of the conference program committees
- Francesca Galassi, Claire Cury: special track on "Artificial Intelligence in Medical Imaging: from the research lab to the clinical practice", IEEE International Symposium on Computer-Based Medical Systems (CBMS) 2023.
- Julie Coloigner, Claire Cury: special session IEEE International Symposium on Biomedical Imaging (ISBI) 2023
- Pierre Maurel, Claire Cury: IABM 2023
- Elise Bannier, Anne Kerbrat: Workshop IRM ARSEP in February 2023
- Camille Maumet: special session IEEE International Symposium on Biomedical Imaging (ISBI) 2023
Reviewer
- Camille Maumet: Reviewer for OHBM 2023.
- Elise Bannier: Reviewer for SFRMBM 2023.
11.1.3 Journal
Member of the editorial boards
- Camille Maumet, editorial board member, Scientific Data (Nature).
- Camille Maumet, editorial board member, Communications Biology (Nature).
- Camille Maumet, editorial board member, Neuroinformatics (Springer).
Reviewer - reviewing activities
- Francesca Galassi, Burhan Rashid Hussein, Cédric Meurée, Julie Coloigner: International Symposium on Biomedical Imaging (ISBI)
- Francesca Galassi, Cédric Meurée: International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI)
- Francesca Galassi, Cédric Meurée, Burhan Rashid Hussein, Ricky Walsh: IEEE 36th International Symposium on Computer-Based Medical Systems (CBMS)
- Francesca Galassi: Frontiers
- Burhan Rashid Hussein: Computers in Biology and Medicine
- Burhan Rashid Hussein: Expert Systems With Applications
- Burhan Rashid Hussein: MDPI
- Burhan Rashid Hussein: Engineering Applications of Artificial Intelligence
- Burhan Rashid Hussein: Applied Soft Computing
- Burhan Rashid Hussein: Pattern Recognition
- Burhan Rashid Hussein: PlosOne
- Burhan Rashid Hussein: Computer and Electronics in Agriculture
- Burhan Rashid Hussein: Computers in biology and medicine
- Burhan Rashid Hussein: Heliyon
- Fanny Dégeilh: Brain
- Fanny Dégeilh: Biological Psychiatry
- Julie Coloigner: Imaging Neuroscience, Nature, Journal of Neuroradiology
- Emmanuel Caruyer: Scientific Data.
- Camille Maumet: Aperture Neuro
- Jérémy Lefort-Besnard: Journal of Open Source Software (review in open access here)
11.1.4 Invited talks
- Agustina Fragueiro, “Pilot study: eye-tracking and skin conductance to monitor task engagement during bimodal neurofeedback” in the special session “Bimodal functional neuroimaging data fusion: methods and applications”, 20th IEEE-International Symposium on Biomedical Imaging (ISBI), Cartagena de Indias, Colombia, April 2023
- Caroline Pinte, "EMPENN projects presentation & Improving portability of bi-modal neurofeedback", Advanced Telecommunications Research Institute International (ATR) Kyoto Japan, September 2023.
- Agustina Fragueiro, “Shift in hippocampal medial position and increased fissure volumes in individuals affected by Developmental Topographical Disorientation”, 8th Scientific Meeting of the Federation of European Societies of Neuropsychology (FESN) and 2nd Panhellenic Conference on Neuropsychology, Thessaloniki, Greece, September 2023
- Francesca Galassi, "Deep Learning in Medical Imaging: What's Needed for Training Data?", Biogenouest days 2023, Oct 2023, Rennes (Campus Villejean), France 82.
- Fanny Dégeilh "Imaging the uniqueness of young child's brain". AI in Pervasive Well-Being and Healthy Ageing Workshop, University of Glasgow, Glasgow, Scotland, June 2023.
- Jérémy Lefort-Besnard, "Data sharing in Europe: reviewing current practices". Conférence GliMR Imaging 2.0 à Porto, Portugal. Mai 2023.
- Jérémy Lefort-Besnard, "The next generation of tools for meta-analysis". Panéliste pour la table ronde de la conférence Cogbases à l'Institut Pasteur, Paris. Octobre 2023.
- Emmanuel Caruyer, "Towards feature-specific diffusion acquisitions?". Microstructure by the Lake, EPFL, Lausanne, Switzerland, September 2023.
- Elise Bannier, Anne Hespel, Camille Maumet "Information, consentement au traitement des données et au recours à l'IA", Journée Les données et l'intelligence artificielle en santé, Rennes, Juin 2023.73.
- Elise Bannier, Claire Cury and Julie Coloigner, "fMRI & EEG bimodal neurofeedback for brain rehabilitation : methodological and clinical aspects", Genève, Switzerland, September 2023.
- Elise Bannier, Anne Hespel, "Partage des données de neuroimagerie : considérations règlementaires, techniques et enjeux de science ouverte", Journée ARDoISE, Atelier rennais de la Donnée, Rennes, Décembre 2023.
- Julie Coloigner, "Marqueurs IRM de l'inflammation : Nouvelles approches et applications en neuropsychiatrie", Journées Neurosciences Psychiatrie Neurologie, Rennes, Juin 2023.
- Camille Maumet "Reproductibilité en neuroimagerie état des lieux", Journées Recherche reproductible: état des lieux, Mars 2023, Paris, France.
- Camille Maumet "BIDS-Prov: Recording neuroimaging provenance", Brain Imaging Data Structure (BIDS) derivatives meeting, Jun 2023, Copenhagen, Denmark.
- Camille Maumet "Open science practices for keeping inventions alive after project ends", ESMRMB 2023 - 39th Annual Scientific Meeting on European Society for Magnetic Resonance in Medicine and Biology, Oct 2023, Basel (CH), Switzerland. 2023. 75.
- Camille Maumet "Towards reproducible neuroimaging across different analysis pipelines" CogBases 2023, Oct 2023, Paris, France. 2023
- Camille Maumet "Towards reusable derived data in neuroimaging", MRI Together, Dec 2023, Online.
11.1.5 Leadership within the scientific community
- Elise Bannier is elected member of the Board of the French Society for MRI Physics (SFRMBM, )
- Elise Bannier is founding member of the Board of the REMI francophone Network (https://remi.network/) for multicentric mutual aid in MRI Clinical research - until December 2023.
- Camille Maumet is member (elected) of the steering committee of the Brain Imaging Data Structure (BIDS)
- Camille Maumet is member (by selection) of the national committee on Open Science, Working group "open software" led by Roberto Di Cosmo and François Pellegrini.
11.1.6 Research administration
- Anne Kerbrat, Francesca Galassi, and Benoit Combès are part of the Scientific Committee of the RHU Primus project.
11.2 Teaching - Supervision - Juries
Participants: Elise Bannier, Isabelle Bonan, Emmanuel Caruyer, Julie Coloigner, Benoit Combès, Isabelle Corouge, Claire Cury, Sébastien Dam, Fanny Dégeilh, Quentin Duche, Jean-Christophe Ferré, Francesca Galassi, Malo Gaubert, Elodie Germani, Burhan Rashid Hussein, Piere-Yves Jonin, Carla Joud, Anne Kerbrat, Stephanie Leplaideur, Jérémy Lefort-Besnard, Camille Maumet, Pierre Maurel, Cédric Meurée, Caroline Pinte, Gabriel Robert, Ricky Walsh.
11.2.1 Teaching
- ESIR, École Supérieure d'Ingénieur de Rennes:
- Pierre Maurel is co-head of the Master program "imagerie numérique" (two last year of the Engineering School)
- Pierre Maurel, "General image processing" (30h).
- Pierre Maurel, "Algorithmique et complexité" (30h).
- Pierre Maurel, "Imagerie médicale" (30h).
- Francesca Galassi is co-head of the Master program "Systemes d'Information" (two last year of the Engineering School)
- Francesca Galassi, "Apprentissage Automatique" (Plenary: 12h, TP: 24h).
- Francesca Galassi, "Algoritmique des graphes" (Plenary: 12h, TD: 6h, TP: 6h).
- Francesca Galassi, "Base de données" (Plenary: 8h).
- Francesca Galassi, "Imagerie médicale" (TP: 15h).
- Ricky Walsh, "Apprentissage Automatique" (TP: 24h).
- Julie Coloigner, "Analyse avancé de Signaux et images" (35h).
- Julie Coloigner, "Mathématiques appliqués" (25h).
- Claire Cury, "Traitement avancé des images" (Plenary : 12h).
- Cédric Meurée, "Algorithmique et complexité" (TD: 14h)
- Caroline Pinte, "Mathématiques appliquées au traitement d'images" (TP: 18h)
- Caroline Pinte, "Projets d'imagerie médicale" (TP: 12h)
- Caroline Pinte, "Neurofeedback (NF) et Brain Computer Interface (BCI) : Une Introduction" (Plenary: 3h)
- Sébastien Dam, "Algorithmique et complexité" (TP: 28h).
- Sébastien Dam, "Algoritmique des graphes" (TD: 6h, TP: 12h).
- Carla Joud, ESIR 1, Maths-S5/S4, Rennes Engineering school (TD: 10h, TP:10h)
- Jérémy Lefort-Besnard, "Algorithmique et complexité" (Master ESIR, TD:14h et TP:14h)
- Master SIBM, M2, University of Angers-Brest-Rennes:
- Jean-Christophe Ferré is head of the master.
- Benoît Combès is co-head of the UE “Modélisation et Apprentissage Automatique pour le Traitement des Images Médicales”.
- Camille Maumet is co-head of the UE “Gestion de données massives et complexes”.
- Emmanuel Caruyer, "Méthodes d'analyse d'IRM de diffusion" (Plenary: 3h).
- Julie Coloigner, "Méthode d'analyse de la connectivité cérébrale" (Plenary: 3h).
- Benoit Combès, "Méthodes de segmentation pour l'imagerie médicale" (Plenary: 3h).
- Benoit Combès, “Méthodes de recalage linéaire et non-linéaires des images médicales” (Plenary: 6h).
- Benoit Combès, “Applications des méthodes de traitement des images médicales” (Plenary: 3h).
- Benoit Combès, “Eléments de statistiques pour l’induction scientifique” (Plenary: 4.5h).
- Benoit Combès, Camille Maumet, “Soutenance de présentations critiques d'articles scientifiques” (TD: 3h).
- Isabelle Corouge, "IRM de perfusion par Arterial Spin Labeling (ASL)" (Plenary: 3h).
- Elise Bannier, "Imagerie fonctionnelle cérébrale" (Plenary: 1h).
- Quentin Duché, "Traitement des données d'IRM fonctionnelle" (Plenary: 1h).
- Elise Bannier, “Utilisation et réutilisation des données d'imagerie” (Plenary: 1h).
- Elodie Germani, "Workflows de traitement d'images" (Plenary: 3h)
- ENS Rennes/Univ. Rennes:
- Emmanuel Caruyer, "Méthodes numériques pour le traitement d'images", L3 SIF (Plenary: 20h).
- Francesca Galassi, "Traitement d'Images", M2 Informatique (Plenary: 10h, TP: 10h).
- Master EIT Data Science, M2, Univ. Rennes:
- Francesca Galassi, "Deep Learning" (Plenary: 12h, TP: 12h).
- Ricky Walsh, "Machine Learning II" (TP: 12h).
- Ricky Walsh, "Case Study" (TD: 12h).
- Master Informatique, ISTIC, Univ. Rennes:
- Julie Coloigner, "Computer Vision" (Plenary: 10h)parcours Science informatique (SIF), M2.
- Elodie Germani, "Option Machine Learning" (TP: 10h), specialités IL (Ingenierie Logiciel).
- Licence 1 ISTN, ISTIC, Université de Rennes 1:
- Elodie Germani, "Principes des Systèmes Informatiques" (TP: 24h)
- Elodie Germani, "Projet professionnel et communication" (TP: 8h)
- Licence BECV (Biologie, Environnement, Chimie du Vivant), University of Rennes 1: Elodie Germani, "Mathématiques pour la biologie" (TD: 27h).
- Bachelor for speech therapy, L3, University of Rennes: Elise Bannier, "Imagerie fonctionnelle cérébrale du langage" (Plenary: 2h).
- Diplome Universitaire MERC (Manipulateur en Recherche Clinique), University of Montpellier: Elise Bannier, "Spécificités de la recherche clinique en imagerie" (Plenary: 7h).
- Master Physique Médicale, M2, University of Rennes:
- Elise Bannier, "Imagerie par Résonance Magnétique " (TD: 4h).
- L2, IFMEM, MR Technologists, University Hospital of Rennes:
- Elise Bannier, "Imagerie par Résonance Magnétique " (Plenary: 10h).
- Master Neuropsychologie, M2, University of Savoie: Pierre-Yves Jonin, "Limites méthodologiques du bilan neuropsychologique à visée diagnostique" (Plenary: 3h30).
- Master Psychologie et neuropsychologie de l’enfant et de l’adulte : langage, cognition et apprentissage, M2, University of Poitiers: Pierre-Yves Jonin, "Méthodologie de l'étude de cas" (Plenary: 3h).
- Master Biologie et Santé, M1, University of Bretagne Occidentale: Pierre-Yves Jonin, "Explorations neuropsychologiques des maladies neurologiques et psychiatriques" (Plenary: 4h).
- Master Psychologie Clinique, Psychopathologie et Psychologie de la Santé, M2, University of Rennes 2:
- Pierre-Yves Jonin, "Neuropsychologie clinique des pathologies neurodégénératives" (Plenary: 4h).
- Pierre-Yves Jonin, "Méthodologie de l'étude de cas" (Plenary: 4h).
- Licence Psychologie, L3, University of Rennes 2:
- Pierre-Yves Jonin, "Les syndromes neuropsychologiques" (TD: 16h).
- Pierre-Yves Jonin, "Approche neuropsychologique du handicap" (TD: 4h).
- Master Neurosciences Cliniques, M2, University of Rennes 1:
- Pierre-Yves Jonin, "Neurosciences cognitives et cliniques de la mémoire humaine" (Plenary: 3h).
- STAPS, DEUST Métiers de la forme, Université de Rennes 2 :
- Camille Muller, "Neurosciences" (TD: 12h).
11.2.2 Supervision
M2 Internship
- M2 SIBM, Univ. Rennes : Alice Dufey, "Quantification of motor tract lesions load and severity and consequences on motor function in patients with relapsing-remitting multiple sclerosis", supervised by Anne Kerbrat and Malo Gaubert.
- M2 Physique, Univ. Rennes : Gaëlle Baudron, "EMISEP : Evaluation de l’impact du changement de machine IRM sur la qualité des images acquises", supervised by Benoit Combès, Malo Gaubert and Anne Kerbrat.
- M2 SIBM, Univ. Rennes : Vivien Caron, "Spinal cord microstructural damage measured in recently diagnosed Relapsing Remitting MS patients: prognostic value at 5-year", supervised by Anne Kerbrat and Benoit Combès.
- M2 IMaGHE Sciences du vivant, EPHE-PSL: Mathilde Liffran "Etude du lien entre l'atteinte structurelle des voies motrices en IRM et l'asymétrie clinique chez les patients atteints de sclérose en plaques progressive", supervised by Anne Kerbrat, Malo Gaubert and Benoit Combès.
- M1, ESIR: Louis-Gabriel Capliez, "Intégration d'un prototype de segmentation de lésions de sclérose en plaques dans la moëlle épinière - Implémentation d'une batterie de tests unitaires, d'intégration et de fonctionnement", supervised by Burhan Rashid Hussein and Cédric Meurée.
- M1, ENS/Univ. Rennes: Lounes Meddhai, "Segmentation of chronique stroke lesions", supervised by Stephanie Leplaideur, Elise Bannier and Francesca Galassi.
- L3, Univ. Rennes: Emma Redor, "Identifying and comparing multiple brain image segmentation algorithms in young children", supervised by Fanny Dégeilh
- M2 Univ. Rennes: Mathys Bizière, "Association between pediatric mild traumatic brain injury and brain structure development in a large population-based study" supervised by Fanny Dégeilh and Claire Cury
- M1, ENS: Emma Redor, "Analyse de la variabilité logicielle dans l'harmonisation inter-scanner des imageries pédiatriques en IRM", supervised by Camille Maumet and Fanny Dégeilh
- M. Sc. Artificial Intelligence and Data Science, M2, University of Düsseldorf: Maiwenn Fleig, "Image-to-image transition between pipelines for functional MRI statistic maps", supervised by Elodie Germani and Elisa Fromont.
- Master Informatique, M1, ENS Rennes: Thibault Chanus, "Image-to-image transition between fMRI pipelines statistic maps using diffusion models", supervised by Elodie Germani and Elisa Fromont.
- Valentin Septiers, ESIR 3, "Évaluation de tâches d'IRM fonctionnelle d'activation pour cartographier les fonctions primaires avant une opération" supervised by Elise Bannier, Quentin Duché, Pierre-Yves Jonin and Nicolas Lassalle.
- Benjamin Prigent, M2 Physique, "Spatio-temporal exploration of fMRI and fNIRS signals" supervised by Elise Bannier, Nicolas Coquery and Yann Serrand.
PhD
- PhD: Ambre Godet, "Intervention par neurofeedback-fNIRS contre l’hyperphagie émotionnelle : caractérisation cérébrale et comportementale", Univ. Rennes, from Oct 2020 to Novembre 2023, David Val-Laillet, Nicolas Coquery, Elise Bannier - Defended in November 2023.
- PhD: Jean-Charles Roy, "Apathy in Late Life Depression: New Biomarkers Using Actimetry and Magnetic Resonance Imaging (ACTIDEP)", Univ. Rennes / INCR, from Oct 2020 to December 2023, Julie Coloigner and Gabriel Robert - Defended in November 2023.
- PhD; Thomas Durantel, "Anatomy and microstructure informed tractography for connectivity evaluation in neurological pathologies", Univ. Rennes, from Nov 2020 to December 2023, Olivier Commowick and Julie Coloigner - Defended in December 2023.
- PhD in progress; Ricky Walsh, "Accurate and robust automated segmentation of multiple sclerosis lesions in spinal cord MRI", Univ. Rennes, from Nov 2022 to December 2025, Francesca Galassi, Benoit Combès and Anne Kerbrat.
- PhD in progress; Nolwenn Jégou, "Apports de nouveaux biomarqueurs de démyélinisation issus de l’IRM quantitative pour le suivi des patients ayant une sclérose en plaques", Inria, from Decembre 2023 to Janvier 2027, Anne Kerbrat, Elise Bannier and Benoit Combès
- PhD in progress; Alix Lamouroux, "Connectivity and Neurofeedback", at IMT Atlantique Brest, from Oct. 2022, co-supervised by Julie Coloigner and Pierre Maurel, with Giulia Lioi (Brain team, IMT Atlantique) and Nicolas Farrugia (Brain team, IMT Atlantique).
- PhD in progress: Carla Joud, "Analyse conjointe de données multimodales en épilepsie", Univ. Rennes, from Nov 2022, Julie Coloigner.
- PhD in progress: Sébastien Dam, "Structural Brain Connectivity and Treatment Response in Mood Depressive Disorder", Inria, from Oct 2022, Julie Coloigner and Pierre Maurel.
- PhD in progress: Caroline Pinte, “Methodology for enhanced and adapted Neurofeedback training”, Univ. Rennes, from Oct 2021, Claire Cury and Pierre Maurel.
- PhD in progress: Lisa Hemforth, "Methodology for automatic scoring of Incomplete Hippocampal Inversion", Sorbonne University, from Oct. 2021, Claire Cury, Baptiste Couvy-Duchesne and Olivier Colliot.
- PhD in progress: Carlo Ferritto "Modeling brain structural and functional connectivity in neurodegenerative diseases", IRISA, CNRS, from Oct 2023, Julie Coloigner.
- PhD in progress: Constance Bocquillon, "Optimizing diffusion MRI acquisition for quantitative connectivity mapping", Univ Rennes, from Oct. 2022, Emmanuel Caruyer and Isabelle Corouge.
- PhD in progress: Marie Poirier, "Robust, patient-specific statistics for the evaluation of brain pathologies using diffusion MRI", Univ. Rennes, from Dec. 2022, Emmanuel Caruyer and Aymeric Stamm (IR CNRS, Laboratoire de Mathématiques Jean Leray, Nantes).
- PhD in progress; Maud Guillen, "Etude longitudinale de la neuroplasticité cérébrale pour la récupération motrice du membre supérieur post-AVC", CHU/Univ. Rennes from December 2023 to Janvier 2027, Isabelle Bonan, Pierre Maurel, Elise Bannier
- PhD in progress: Elodie Germani, “Mapping the fMRI pipeline-space towards more robust pipelines”, Univ. Rennes, from Oct 2021, Camille Maumet and Elisa Fromont (Lacodam).
11.2.3 Juries
- Fanny Dégeilh: Member - Jury d'Examen général de synthèse (doctorat) : Polytechnique Montréal (Canada)
- Francesca Galassi: Member of Selection Committee (COS) for the position of Associate Professor (Maître de conférence, section 61) Toulouse NeuroImaging Center, Université de Toulouse.
- Emmanuel Caruyer: Reviewer of the PhD thesis of Zheyi Yang, Institut polytechnique de Paris/Inria Saclay.
- Elise Bannier: Reviewer of the PhD Thesis of Giulia Rocco, Université de Nice.
- Elise Bannier: Examiner of the PhD Thesis of Emile Kadalie, Université de Bordeaux.
- Francesca Galassi: Member of Selection Committee (COS) for the position of Associate Professor (Maître de conférence, section 61) Toulouse NeuroImaging Center, Université de Toulouse.
- Emmanuel Caruyer: reviewer of the PhD thesis of Zheyi Yang, Institut polytechnique de Paris/Inria Saclay.
- Pierre Maurel : President of the doctoral defence Jury of Jean-Baptiste de Saint-Aubert, Institut de la Vision, Sorbonne University.
- Pierre Maurel : President of the doctoral defense jury of Thomas Durantel, IRISA, Rennes University.
11.3 Popularization
Participants: Claire Cury, Elodie Germani, Jérémy Lefort-Besnard.
11.3.1 Internal or external Inria responsibilities
- Claire Cury, mediation Officer of the Inria Rennes Scientific mediation team.
- Camille Maumet is a co-organizer of the local version of the program "L codent L créent", an outreach program to send PhD students to teach Python to middle school students in 8 sessions of 45 minutes. It was initiated in Lille, with Anne-Cécile Orgerie and Tassadit Bouadi. The program is currently supported by: Fondation Blaise Pascal, ED MathSTIC, Inria and Fondation Rennes 1.
11.3.2 Education
- Claire Cury, setting-up a general audience writing workshop for 2nd year PhD students.
11.3.3 Interventions
- Elodie Germani, participating in the LCLC (Elles Codent Elles Créent) program for middle schoolers at Collège Le Landry, Rennes, 8 * 45min of workshops between January and March 2023.
- Elodie Germani, intervention as part of the training on scientific mediation, in the TISSAGE project (TrIptyque Science Société pour AGir Ensemble), May 2023.
- Jérémy Lefort-Besnard, organisateur d’un atelier de conception, câblage et programmation d’un robot avec Arduino
12 Scientific production
12.1 Major publications
- 1 articleUnsupervised Domain Adaptation With Optimal Transport in Multi-Site Segmentation of Multiple Sclerosis Lesions From MRI Data.Frontiers in Computational Neuroscience14March 2020, 1-13HALDOI
- 2 articleIsolating the Sources of Pipeline-Variability in Group-Level Task-fMRI results.Human Brain MappingNovember 2021HALDOI
- 3 articleFocal and diffuse cervical spinal cord damage in patients with early relapsing--remitting MS: A multicentre magnetisation transfer ratio study.Multiple Sclerosis Journal258February 2019, 1113-1123HALDOI
- 4 articleObjective Evaluation of Multiple Sclerosis Lesion Segmentation using a Data Management and Processing Infrastructure.Scientific Reports81December 2018, 13650HALDOI
- 5 articleA sparse EEG-informed fMRI model for hybrid EEG-fMRI neurofeedback prediction.Frontiers in Neuroscience13January 2020HALDOI
- 6 articleMultiple sclerosis lesions in motor tracts from brain to cervical cord: spatial distribution and correlation with disability.Brain - A Journal of Neurology 1437July 2020, 2089-2105HALDOI
- 7 articleAn iterative centroid approach for diffeomorphic online atlasing.IEEE Transactions on Medical Imaging4192022, 2521-2531HALDOI
- 8 articleThe impact of Neurofeedback on effective connectivity networks in chronic stroke patients: an exploratory study.Journal of Neural Engineering185September 2021, 056052HALDOI
- 9 articlePatch-Based Super-Resolution of Arterial Spin Labeling Magnetic Resonance Images.NeuroImage189January 2019, 85-94HALDOI
- 10 articleAssociation of Gray Matter and Personality Development With Increased Drunkenness Frequency During Adolescence.JAMA Psychiatry774April 2020, 409-419HALDOI
- 11 articleConnectivity patterns of the core resting-state networks associated with apathy in late-life depression.Journal of Psychiatry and Neuroscience486November 2023, E404 - E413HALDOI
12.2 Publications of the year
International journals
International peer-reviewed conferences
Conferences without proceedings
Reports & preprints
Other scientific publications
12.3 Other
Scientific popularization
12.4 Cited publications
- 83 articleA sparse EEG-informed fMRI model for hybrid EEG-fMRI neurofeedback prediction.Frontiers in Neuroscience13January 2020HALDOIback to text
- 84 articleUnimodal Versus Bimodal EEG-fMRI Neurofeedback of a Motor Imagery Task.Frontiers in Human Neuroscience11April 2017HALDOIback to text
